-
Progress in cell and molecular biology has furthered our understanding of eukaryotic gene transcriptional regulation and expanded the study of chromosomal and nuclear structure. Researchers of correlative gene transcriptional regulation need to grasp nuclear fine structure and formation in more detail. It is now appreciated that many DNA replication factors, transcription factors and splicing factors are not uniformly scattered in the nuclei but rather exist in condensed form aggregating into uneven "foci" within nuclear chromosome territories[4, 20]. The non-random distribution of these "foci" [7] and their functional role in cellular transcriptional regulation merit further investigation. Cell and molecular biology research has acknowledged the importance of understanding the formation of different functional domains although there is no consensus on the actual function of these nuclear "foci."
A number of studies have indicated that several important transcriptional regulatory factors in eukaryotes, such as E2F-1, GATA-1, SP1, and OCT1, could disperse or aggregate at specified regions in the nuclei [6, 10, 16, 18], and this location distinction might contribute to determining which phase a cell is in during the cell cycle. This deduction is mainly based on the observation that nuclear dispersed DNA replication factors only concentrate to about 150 replication focuses in S phase[1] and nuclear dissociation may also lead to disaggregation of the nuclear protein complex as parental cells divide into daughter cells. However, this deduction has not been fully confirmed by experiments. Therefore, research into viral transcriptional regulation factors expressed in host cells is warranted. Studies of the molecular biology of herpes simplex virus 1 (HSV-1) infection have identified many regulatory proteins generated after HSV-1 infected host cells could form typical punctiform structures in the nucleus[13, 14]. For example, as a multifunctional regulatory protein, HSV-1 infected-cell protein 22 (ICP22) not only regulates expression levels of viral and host genes but also affects mRNA splicing and changes the phosphorylated form of RNA polymerase LS[2, 8, 15, 17]. During the process of viral infection, the localization of ICP22 changes from aggregation to dispersion and back to aggregation[12]. ICP22 expressed alone is localized to small dense nuclear bodies and is paired with the SC-35 domain in the nucleus. Its biological characteristics are still unknown although these dense bodies have been found to correlate with its function by several experiments. Therefore, exploring the biological characteristics of this protein's cellular localization not only strengthens our knowledge of HSV-1 infection but also provides more information to understand cellular transcriptional regulation. Forthcoming experiments have confirmed that the structure of ICP22 provides two nuclear localization signals responsible for getting proteins into the nucleus. The N-terminus of ICP22 determines the formation of small dense nuclear bodies[5, 19]. Thus, a fusion protein (ICP22-EGFP), in which the C-terminus of ICP22 is combined with green fluorescence, was constructed to observe cellular localization of ICP22. Then we investigated ICP22's cellular biological characteristics by cell synchronization and a Tet-induced expression regulated system induced by vibramycin(Dox). Results indicated that cellular localization of ICP22 is not affected by cell cycle. However, it is worth noting that expression intensity of ICP22 correlated with its cellular localization. Results suggest that the cellular biological characteristics of proteins with transcriptional regulatory functions depend on the extent of protein self-expression. In addition, we developed a new non-protease expression reporter system, which could evaluate transcriptional control of internal ribosome entry sites (IRES), based on the observation that the localization of ICP22 reflects its own expression level.
HTML
-
Chinese hamster ovary (CHO) cells were grown in complete Ham's F12 media containing 5% fetal calf serum with 100 U/mL penicillin and 20 μg/mL streptomycin under 5% CO2 at 37℃. Vero cells were grown in DMEM containing 10% fetal calf serum with 100 U/mL penicillin and 20 μg/mL streptomycin under 5% CO2 at 37℃. Both cells were maintained in this laboratory. Plasmids pEGFPN2 (to express GFP) and pEGFPN-us1 (to express ICP22-GFP fusion protein) were maintained and constructed in this laboratory (Fig. 1) [5]. Tet regulatory system plasmids, pRevTRE and pRevTet-On, and the retrovirus package cell line, Retropack PT67, were obtained from Clontech.
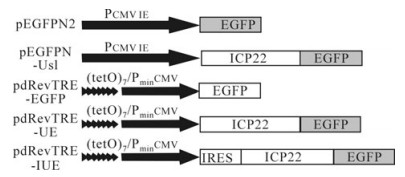
Figure 1. Recombinant plasmid. pEGFPN2 and pEGFPN-Us1, driven by the Pcmv promoter, express EGFP and ICP22-EGFP, respectively. pdRevTRE-EGFP and pdRevTRE-UE, which consist of 7 repeated Tet operons (tetO) and are under the control of CMV promoters lacking the enhancer, following induction express EGFP and ICP22-EGFP, respectively. Recombinant plasmid pdRevTRE-IUE was constructed by inserting an IRES element of EMCV into the pdRevTRE-UE plasmid between its promoter and the protein of interest.
-
The double-stranded DNA adaptor fragment 5 -AGCGTCGACCTCGAGAAGCTTGCT-3 was digested with SalⅠ and Hind Ⅲ and ligated into identically treated pRevTRE vectors to remove the ATG between SalⅠ and Hind Ⅲ sites and construct the plasmid pdRevTRE without a translation initiation codon. The GFP gene was amplified from plasmid pEGFPN2 by primers 5 -AGCATCGATCCGGTCGCCACCATGGTG-3 and 5 -AGCATCGATCGTCGCGGCCGCTTTACTTG-3 and then ligated into the pdRevTRE vector through a ClaⅠsite to construct the recombinant plasmid pdRevTRE-EGFP. The non-terminator Us1 gene codon of ICP22 was amplified from plasmid pEGFPN-Us1 by primers 5 -AGCAAGCTTATGGCCGACATTTCCCCA GG-3 and 5 -AGCAAGCTTCGGCCGGAGAAACGTGTCG-3 and then ligated into the pdRevTRE-EGFP vector through a Hind Ⅲ site to construct pdRevTRE-UE. An IRES element of the Encephalomyocarditis virus was amplified from plasmid pMSCV-IRES-GFP by primers 5 -AGCGTCGACCCCTCTCCCTCCCCCCCCCCTAACG-3 and 5 -AGCGTCGACGGTTGT GGCGTTATTATCATCGTG-3 and ligated into the pdRevTRE-UE vector through a SalⅠsite to construct recombinant plasmid pdRevTRE-IUE. All recombinant plasmids were identified by DNA sequencing. Expressed ORFs from each constructed plasmid are shown in Fig. 1.
-
CHO cells, grown in 6-well plates to 70% confluence, were transfected with Lipofectamine 2000 and the related plasmid (1 mg/well) according to the manufacturer's instructions.
-
Transient transfected CHO-k1 cells were diluted in Pancebrin after 12 h cultivation, then synchronized at the beginning of S phase by a double thymidine block[11] and release protocol (20 h incubation with 7.5 mmol/L double thymidine followed by an interval of thymidine-free incubation for 8 h and then a second thymidine incubation for 14 h).
-
Tetracycline-controlled transactivator (rtTA) plasmid pRevTet-On, recombinant plasmids (pdRevTRE-EGFP, pdRevTRE-UE, and pdRevTRE-IUE), and control plasmid pMSCV-IRES-GFP were transfected into the retrovirus package cell line Retropack PT67. Each culture medium supernatant was collected at 48 h post-transfection and filtered with a 0.45 μm membrane to acquire the recombinant retroviruses Vtet-on, Ve, Vue, and Viue. These recombinant retroviruses were used to directly infect cells or were stored at -80℃. The recombinant retrovirus MSCV-IRES-GFP with synchronization was evaluated by detecting virus titer (CHO-K1 cells were infected with virus diluted 10-times continuously and then cells expressing EGFP were counted).
-
CHO-k1 cells were cultivated and selected with 450 μg/mL G418 following infection with Vue or Viue virus in the presence of 4 μg/mL hexadimethrine bromide. Several positive clones were then selected and cultivated to establish a stable CHO-k1 Tet-on cell line.
-
Cells from the stable CHO-k1 Tet-on cell line were cultivated and selected with 300 μg/mL hygromycin B following infection with the Vue or Viue virus in the presence of 4 μg/mL hexadimerethrine bromide. Induced expression cell lines CHO-TRE-E, CHO-TRE-UE and CHO-TRE-IUE were established, selected with 450 μg/mL G418 and 300 μg/mL hygromycin B, and examined by fluorescence microscopy 48 h after treatment with different concentrations of vibramycin (Dox).
Cells and plasmids
Plasmid construction
Cell transfection
Cell synchronization
Recombinant retrovirus and virus titer detection
Establishment of stable CHO-k1 Tet-on cell line
Establishment of Tet-induced expression cell line and examination of induced expression
-
Observation of CHO-k1 (hamster), Vero (monkey) and KMB17 (human) cells transfected with pEGFPN2 and pEGFPN-us1 independently demonstrated that ICP22-EGFP fusion proteins localize in the nucleus, in contrast to EGFP (Fig. 2). Briefly, ICP22-EGFP tended to form small dense nuclear bodies in CHO-k1 cells and to disperse in KMB17 cells. In Vero cells, ICP22-EGFP underwent moderate aggregation.
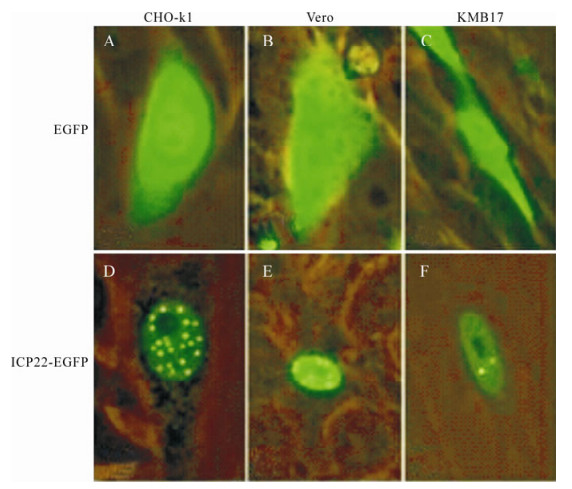
Figure 2. Analysis of localization of ICP22 in CHO-k1, Vero and KMB17 cells. A-C: pEGFPN2 was transfected into CHO-k1, Vero and KMB17 cells, respectively. D-F: pEGFPN-Us1 was transfected into CHO-k1, Vero and KMB17 cells, respectively. Expression was examined under a fluorescence microscope 48 h post-transfection. ICP22-EGFP localized into nucleus and formed small dense nuclear bodies; in contrast, EGFP dispersed throughout each cell.
-
In order to investigate the aggregation of ICP22-EGFP in CHO-k1 nuclei, observations were carried out consecutively at different time points. The time at which Pancebrin was added after transient transfection was considered the initiation point. Results demonstrated that ICP22-EGFP localizes to the nucleus at 3 h post-transfection and begins to form dense fluorescent spots at 12 h post-transfection. At 24 h post-transfection, most CHO-k1 cells transfected with pEGFPN-Us1 showed dense fluorescence spots in their nuclei (Fig. 3).
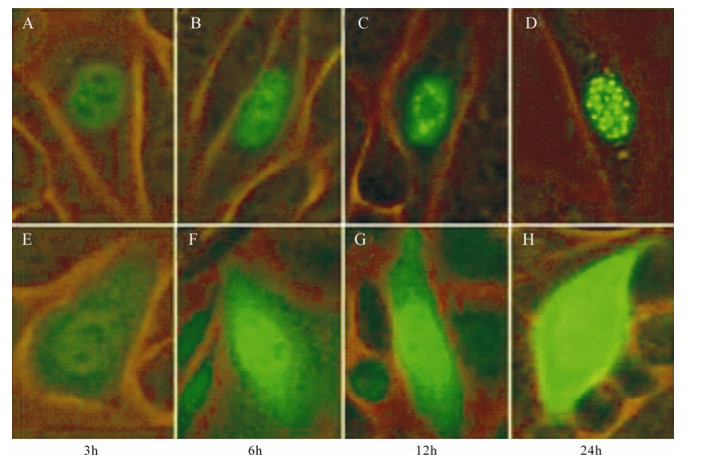
Figure 3. Cellular aggregation of ICP22. A-D: Expression of ICP22-EGFP by CHO-k1 cells transfected with pEGFPN-Us1 at different time. E-H: Expression of EGFP by CHO-k1 cells transfected with pEGFP-N2 at different times. ICP22-EGFP gradually formed punctiform structures in nuclei, whereas control EGFP was distributed throughout the cells at all time points.
-
Observation of ICP22-EGFP localization was performed after synchronization of transfected CHO-k1 cells in order to investigate the correlation between ICP22 location and cell cycle. Unexpectedly, our results indicated that ICP22-EGFP concentrated in spots in the nuclei of cells arrested in G1/S phase, S phase (3 h post-blockade), G2/M phase (7 h post-blockade) and G2 phase (20 h post-blockade), in contrast to control EGFP, which did not localize to the nuclei (Fig. 4). Notably, the small dense nuclear bodies formed by ICP22-EGFP persisted while most nuclear structures changed in metaphase.
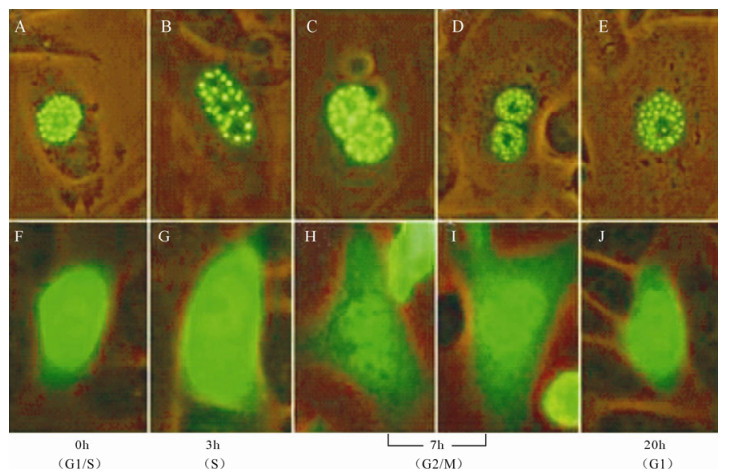
Figure 4. ICP22 forms small dense nuclear bodies throughout the cell cycle. A-E: Observation of ICP22-EGFP localization was performed after cell synchronization of transfected CHO-k1 cells in G1/S phase, S phase, G2/M phase, and G1 phase. EGFP diffused throughout whole CHO-k1 cells while ICP22-EGFP always formed punctiform structures in nuclei. F-J: Observation of EGFP localization was performed after synchronization of transfected CHO-k1 cells in G1/S phase, S phase, G2/M phase, and G1 phase.
-
It is known that the expression level of a given protein increases with the concentration of Tet or Dox in Tet-induced expression cell lines. Fig. 5 shows aggregation levels of EGFP increasing along with the concentration of Dox in the CHO-TRE-E cell line, and the localization of ICP22-EGFP was markedly different. With 0.1 ng/mL Dox, ICP22-EGFP dispersed even after 48 h. When concentration of Dox was increased to 1 ng/mL, ICP22-EGFP aggregated in nucleus and sporadically formed small dense nuclear bodies. The aggregation of ICP22-EGFP was increasingly distinct with increasing Dox concentration. ICP22-EGFP formed uniform stable small dense nuclear bodies when the concentration of Dox reached 100 ng/mL.
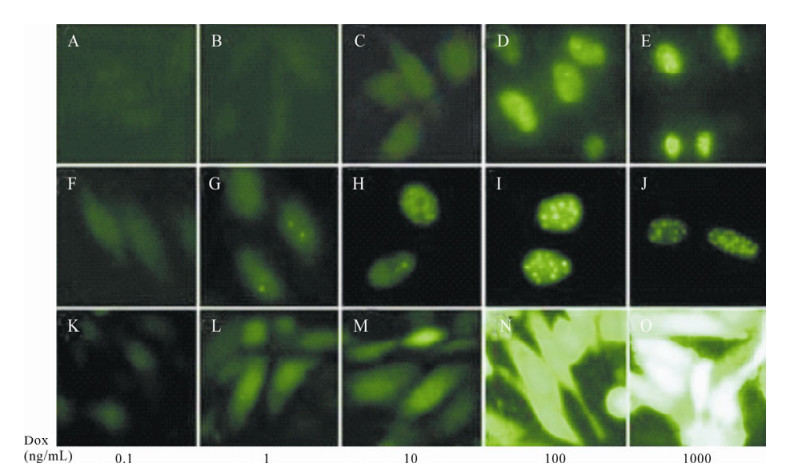
Figure 5. Cellular localization of ICP22 is regulated by its expression intensity. A-E: Expression of fluorescent protein in CHO-TRE-IUE cells under 0.1 ng/mL, 1 ng/mL, 10 ng/mL, 100 ng/mL, and 1000 ng/mL Dox. F-J: Expression of fluorescent protein in CHO-TRE-UE cells under 0.1 ng/mL, 1 ng/mL, 10 ng/mL, 100 ng/mL, and 1000 ng/mL Dox. K-O: Expression of fluorescent protein in CHO-TRE-E cells under 0.1 ng/mL, 1 ng/mL, 10 ng/mL, 100 ng/mL, and 1000 ng/mL Dox.
-
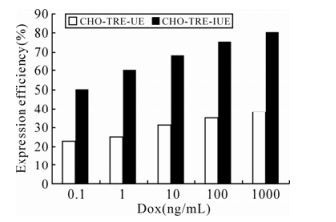
Since the expression intensity of ICP22-EGFP could determine its nuclear localization, the nuclear localization of ICP22-EGFP could reflect its protein expression level directly. In order to examine the effects of IRES on the expression of our protein of interest, an IRES element was inserted into the pdRevTRE-UE vector between the promoter and the protein of interest to construct pdRevTRE-IUE, which was used to create the Tet-induced expression cell line CHO-TRE-IUE. Results are shown in Fig. 5. ICP22-EGFP entered nuclei when the Dox concentration reached 10 ng/mL. Sporadic small dense nuclear bodies formed in nuclei when the concentration of Dox reached 100 ng/ml. ICP22-EGFP formed uniform stable small dense nuclear bodies when the Dox concentration reached 1000 ng/mL. Compared to the CHO-TRE-UE cell line, the presence of an IRES element reduced expression efficiency of the protein interest in an individual cell but the presence of an IRES element increased the percentage of cells that expressed fluorescence protein (Fig. 6).
Formation of small dense nuclear bodies by ICP22 is not cell-specific
ICP22-EGFP forms stable small dense nuclear bodies at 24 h post-transfection
Cell division does not induce disaggregation of the small dense nuclear bodies
The aggregation of ICP22-EGFP correlated with its expression intensity
Effects of IRES element on expression of ICP22-EGFP
-
An EGFP expression system was applied in this research to investigate the localization of ICP22-EGFP in mammalian cells. Previous data demonstrated that EGFP can diffuse passively through cell nuclear membranes, passing back and forth between the cytoplasm and nucleus. But when EGFP is expressed as a fusion protein and the molecular weight of this fusion protein exceeds 60 kDa, it can only enter nuclei via the nuclear pore complex with the aid of a nuclear localization signal, which is not possessed by EGFP[3, 21]. Therefore, the entrance of ICP22-EGFP fusion protein (effective molecular weight 73 kDa; apparent molecular weight 106 kDa) into nuclei completely depends on ICP22. Furthermore, the nuclear aggregation of ICP22 is determined by its N-terminal sequence, and the EGFP fusion tail of ICP22 C-terminal does not affect its aggregation in nuclei[5]. Localization of ICP22 in nuclei could be observed directly by measuring the fluorescence emitted by ICP22-EGFP and removing background interference.
Our experiments demonstrated that cell specificity is not a major determinant of protein aggregation or diffusion in nuclei (Fig. 2). Furthermore, the time between nuclear entry and ICP22-EGFP aggregation in nuclei was usually less than 24 h in CHO-k1 cells (Fig. 3). Given the two points discussed above, we decided to further examine ICP22-EGFP distribution during the cell cycle. Out results indicated that the small dense nuclear bodies formed by ICP22-EGFP do not disaggregate during the cell cycle. Contrary to our expectations, ICP22-EGFP persists in an aggregated state even in the phase of cell division, in which nuclear structures undergo great changes. Aggregated ICP22-EGFP seemed to be allocated equally between two daughter cells. This finding suggests that the aggregated region of ICP22 may represent an undetermined nuclear functional domain; alternatively, this phenomenon may be related to specific mechanisms of viral infection.
Transcription of different pEGFPN vectors utilized in our experiments was promoted by the cytomegalovirus immediate-early promoter (Pcmv) (Fig. 1). To our knowledge, Pcmv is a strong transcriptional promoter, and protein expression controlled by Pcmv should be abnormally excessive[22]. In order to analyze the effects of different expression levels on protein localization, a Tet-induced regulatory system was adopted to investigate the expression of our protein of interest as influenced by Dox concentration[9]. Experimental results indicated that sufficient time was available for ICP22-EGFP to aggregate in nuclei even at 48 h post-induction but, at low transcriptional levels, ICP22-EGFP could only exist in a dispersed form. ICP22-EGFP could form small dense nuclear bodies only at high transcriptional levels. This observation may be attributed to simple aggregation of excessive proteins or may be the result of specific biological mechanisms. If the former statement is correct, we need to re-evaluate interactions between many proteins. If the latter is correct, this suggests a new transcriptional regulation mechanism. We prefer the latter hypothesis for the following three reasons. First, if this aggregation is related to denaturation of excessive protein, then the aggregation should trigger the formation of inclusion bodies at the site of protein synthesis (cytoplasm) instead of in nuclei. Second, small dense nuclear bodies formed by ICP22 seem to be stably located. Moreover, they are equally distributed to nuclei of daughter cells during cell division. Hence, this is not random aggregation. Third, if this were simple aggregation of excess protein, this aggregation would not be related to the structure of ICP22. Our previous experiments demonstrated that amino acid substitutions in ICP22 lead to dispersion of small dense nuclear bodies even with high transcription levels under control by the Pcmv promoter[5]. Therefore, it can be presumed that, with regard to the ICP22-EGFP fusion protein, variation of its transcriptional level could regulate cellular localization of ICP22. The biological significance of this location variation requires further investigation. Changes in ICP22 localization in viral replication suggest that such a change might be essential to this process. Therefore, the effects of ICP22 protein transcriptional level on protein localization and function discussed in this article might provide new clues to better understand gene transcriptional control. However, since transcriptional levels could determine cellular localization of ICP22-EGFP, cellular localization of ICP22-EGFP may conversely reflect expression level. With regard to this point, it is possible for us to construct a new reporter system for the purpose of examining the effects of certain factors on protein expression. In our research, IRES reduced expression levels of a reporter gene in individual cells but increased the cell population in which the reporter gene was expressed. This system overcomes defects of protease reporter genes (such as CAT, β-gal), which are difficult to examine on a single-cell level. We will elaborate on this point in detail in subsequent research.













 DownLoad:
DownLoad: