-
Dear Editor,
Recombination contributes greatly to the diversity of human immunodeficiency virus type 1 (HIV-1). A large number of recombinant strains have been found in China, particularly in Yunnan, which is considered the HIV-1 epicenter of China. Surveillance of unique recombinant forms is helpful for prediction of new circulating recombinant forms. In this study, we identified two unique recombinant forms of HIV-1 in Yunnan Province. The near full-length genomes of HIV-1 strains were amplified in three parts, and the genomic structures of two strains (12YN10159 and 12YN10192) were analyzed. Strain 12YN10159 was comprised of subtypes C and CRF01_AE, while strain 12YN10192 was comprised of subtypes B and C. Genomic breakpoints were different from all previously reported strains. The data emphasized that more surveillance of unique strains of HIV is needed in Yunnan.
HIV-1 is characterized by a high level of genetic variation and diversity, contributing to its high mutation rate and short generation time (Ho et al., 1995). Recombination is also a major mechanism contributing to sequence diversity. Within the last decade, many circulating recombinant forms (CRFs) have been identified; these recombinant HIV strains have been shown to account for more than 20% of all HIV infections worldwide. Currently, 73 CRFs (http://www.hiv.lanl.gov/content/sequence/HIV/CRFs/CRFs.html) and several hundred unique recombinant forms (URFs) have also been reported.
Yunnan Province has been the location of most HIV-1 epidemic outbreaks in China. Subtypes B and C were responsible for the initial epidemic in the early 1990s (Graf et al., 1998). Subsequently, more HIV-1 subtypes, including B, Thai B, C, CRF07_BC, CRF08_BC, and CRF01_AE, have been detected (Tu et al., 2009). The presence of many subtype strains circulating in the same population will always lead to the occurrence of recombinant strains. In this study, we identified two HIV-1 URFs in Yunnan Province, China.
Plasma samples were obtained from two HIV-1-infected individuals living in Yunnan Province in 2012. In one patient, strain 12YN10159 was acquired through heterosexual transmission, and the patient was diagnosed as HIV-1 positive in 2012. HAART therapy was administered beginning in 2012. In the other patient, strain 12YN10192 was acquired through intravenous drug use, and the patient was diagnosed as HIV-1 positive in 2004. HAART therapy was administered beginning in 2008. Virions in 500 μL plasma were concentrated by centrifugation at 23, 000 × g for 1 h, and viral RNA was extracted using a High Pure Viral RNA Kit (Roche, South San Francisco, USA). The near full-length genome (NFLG) of HIV-1 was amplified in three parts, including the gag-pol region and 3′ halves using nested reverse transcriptional polymerase chain reaction (PCR), as previously described (Li et al., 2010).
The positive PCR products were sequenced by Huada Genomics Company (Beijing, China) with a variety of internal specific primers (available on request) after being purified. A BLAST search against the HIV-1 sequence database was used to exclude potential contamination.
All sequences segments were assembled using ContigExpress software, a component of Vector NTI Suite 6.0. The NFLGs of strains 12YN10159 (KT321211, 8957 bp) and 12YN10192 (KT960983, 8876 bp) were obtained. Gene cutter (http://www.hiv.lanl.gov/content/sequence/GENE_CUTTER/cutter.html) was used to analyze the assembled NFLGs of strains 12YN10159 and 12YN10192, which revealed nine proper open reading frames (ORFs). Phylogenetic analysis of the NFLGs demonstrated that strain 12YN10159 clustered with subtype C, whereas strain 12YN10192 clustered with the subtype B reference, suggesting that they originated from subtype C or B strains separately (Figure 1A). Both sequences were located between reference strains belonging to the different subtypes, suggesting that they were recombinant forms. Strain 12YN10159 clustered with subtype C strains from India, and strain 12YN10192 clustered with subtype B references from Thail and and Japan. The recombination was further confirmed using the Recombination Identification Program (RIP, version 3.0), available at the HIV sequence database (http://www.hiv.lanl.gov/content/sequence/RIP/RIP.html) (data not shown). To determine the recombinant forms of both strains, bootscanning analysis was performed by using the neighbor-joining method with the representative reference strains. In the 12YN10159 genome, three CRF01_AE segments were inserted into the genome of the subtype C backbone (Figure 1B). In the 12YN10192 genome, several small segments of subtype C were inserted into the subtype B backbone (Figure 1B). These strains were distinct from the other CRFs and URFs reported in Yunnan. Neighbor-joining trees were further built based on the segments belonging to the different subtypes to explore the origin of each segment. Bootstrap analysis (1000 replicates) was used to estimate the reliability of topologies.
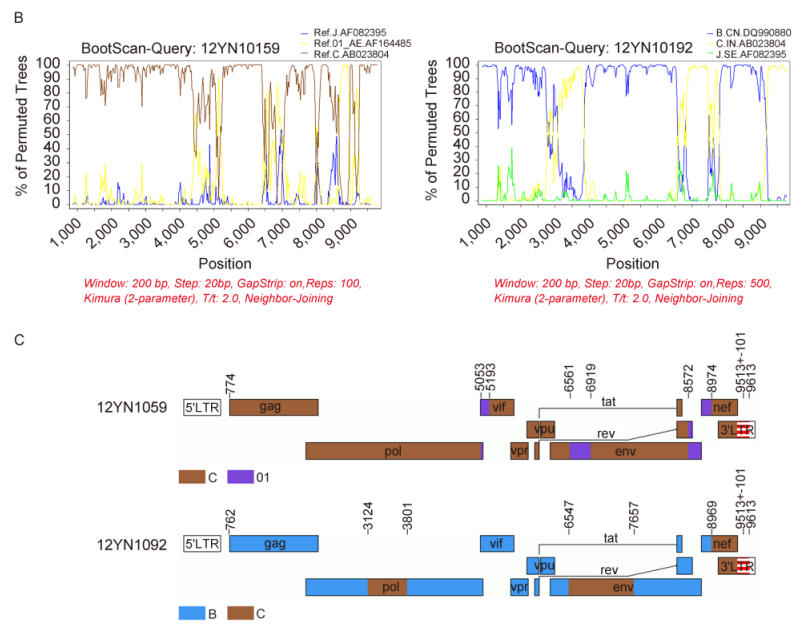
Figure 1. (A) Neighbor-joining tree based on alignment of near full-length genomes of 12YN10159, 12YN10192, reference sequences representing different HIV-1 group M subtypes (http://hiv-web.lanl.gov/), and some geographically important CRFs (CRF01_AE, CRF07_BC, and CRF08_BC). The bootstrap values are indicated at the corresponding nodes of the tree. The scale bar represents 1% genetic distance (0.01 substitution per site). (B) Bootscanning analysis of the near full-length nucleotide sequences of strains 12YN10159 and 12YN10192 using Similarity software (version 3.5.1; S. Ray, Johns Hopkins University, Baltimore, MD; http://sray.med.som.jhmi.edu/RaySoft/SimPlot/). Subtypes C (93IN101.AB023804) and CRF01_AE (93TH9021.AF164485) were used as references for 12YN10159 analysis, while subtypes C (93IN101.AB023804) and B (905CNHB_hp3.DQ990880) were used as references for 12YN10192 analysis. Subtype J (SE9173_7022.AF082395) was used as an outgroup. The x-axis is the nucleotide position in the multiple alignments of the near full-length HIV-1 sequences, and the y-axis is the bootstrap value (percent). (C) Schematic representation of the subtype structures of 12YN10159 and 12YN10192. All breakpoints are listed above the genomic map separately.
To clarify the exact genomic structure and breakpoints in both newly identified HIV-1 unique recombinant strains, genetic maps were constructed using jpHMM software (Georg-August University of Göttingen, Germany; http://jphmm.Gobics.de/submission_hiv). In the 12YN10159 genome, three CRF01_AE segments were separately inserted into the subtype C genome in env and nef regions at the location of nucleotide positions 5053–5193, 6561–6919, and 8572–8974 according to the HXB2 calibrator (Figure 1C). In the 12YN10192 genome, three small subtype C segments were located in the pol (3124–3801), env (6547–7657), and nef (8969–9412) gene regions separately (Figure 1C).
The subtype C strain was first detected in intravenous drug users (IDUs) in Yunnan in 1997 (Guan et al., 1997). With the introduction of other HIV-1 subtypes into Yunnan province, increasing recombination between subtype C strains and other subtypes has been observed. A recombinant strain involving subtypes C and CRF01_AE was reported in Yunnan in 2010 (Li et al., 2010), which contained the genome of subtype CRF01_AE as a backbone and several subtype C segments inserted at different locations within this genome. In contrast, strain 12YN10159 contained the subtype C strain as the backbone with CRF01_ AE segments inserted. Very few pure subtype C strains are currently found in China, most of which have been found in IDUs. Phylogenetic analysis of strain 12YN10159 demonstrated that the parental strain subtype C backbone may be from India, i.e., one of the strains that initially spread into China. The identification of strain 12YN10159 in the heterosexual population indicated that subtype C strains may be still prevalent in some specific populations.
In contrast to strain 12YN10159, strain 12YN10192 contained a genome with subtype B as the backbone. Phylogenetic analysis of this strain showed that this backbone may have a common origin with reference strains from Thail and and Japan; moreover, the subtype B backbone in Yunnan was from Thail and, as reported in earlier studies (Chen et al., 2013). In China, many B/C CRFs, including CRF07_BC, CRF08_BC, CRF57_BC, CRF61_BC, CRF62_BC, and CRF64_BC, have been identified. Among these CRFs, CRF07_BC and CRF08_ BC are thought to have originated in Yunnan province (McClutchan et al., 2002; Su et al., 2000; Yang et al., 2002) and subsequently spread to other areas. No genomic structure similar to that of strain 12YN0192 has been found. Moreover, in contrast to most B/C recombinations, strain 12YN10192 contained the subtype B sequence as the skeleton with C fragments inserted. Because the subtype C backbone was common in B/C recombination, the identification of strain 12YN10192 suggested that HIV prevalent in IDUs in Yunnan provin-ce may have complex origins.
To date, many URFs have been reported in Yunnan Province, particularly in Dehong Prefecture, which is close to Myanmar border (Yang et al., 2002). However, there are still very few NFLGs available for analysis. Most sequences obtained in this area have been analyzed basing on partial fragments of the HIV-1 genome. The identification of the two URFs identified in this study will provide important information regarding the HIV genome. Moreover, although only a few recombinant strains have spread outside Yunnan province, surveillance of these URFs will be helpful for the control and prevention of HIV in this area.
HTML
-
This work was supported by the Applied Basic Research Programs of the Science and Technology Department of Yunnan Province (Grant No. 2013FZ217) and National Key S & T Special Projects on Major Infectious Diseases (Grant No. 2012ZX10001-002). The authors declare that they have no competing interests. The study was approved by the ethics committees of Health Department of Yunnan Province. Written consents were obtained from these two patients involved in the study.














 DownLoad:
DownLoad: