HTML
-
Influenza, commonly known as "the flu", is a widespread infection caused by the influenza virus that is highly contagious and has a high incidence rate. According to the World Health Organization, the annual global attack rate of influenza is estimated at 5%-10% of adults and 20%-30% of children, of which 30-50 million cases result in severe infections, and approximately 3-5 million in death. Recently, it has been revealed that the influenza virus mainly attacks respiratory epithelial cells and alveolar cells, which can lead to lung infection, oedema, shock, multiple organ failure, and other types of injury (Desdouits et al. 2013; Hendrickson et al. 2013; Guedj et al. 2012; Mammas et al. 2011). In general, central nervous system complications, including febrile seizures, acute disseminated encephalitis (encephalopathy), and Reye syndrome, as well as other types of nervous system abnormalities, can occur following influenza virus infection. These complications with neurologic sequelae have a poor prognosis and high mortality rate (Blackmore et al. 2017; Ishida et al. 2015; Wilking et al. 2014). However, the induction and mechanisms of these serious adverse reactions caused by influenza virus infection are not fully understood.
Heterophilic antigens, first described by Forssman (1911), are a group of common antigens that exist in humans, animals, and microorganisms. Prediction, identification, and structural analysis of heterophilic epitopes of antigens can improve our understanding of viral antigenic variation, epidemics, and pathogenesis of nervous system diseases. For example, heterophilic epitopes common to both Escherichia coli O14 lipopolysaccharides and human colon mucosa antigens may lead to the occurrence of ulcerative colitis (Wang et al. 2014). Levin et al. (2002) reported the existence of heterophilic epitopes between human T-lymphotropic virus type 1 (HTLV-1) and heterogeneous nuclear riboprotein A1 (hnRNPA1) in the central nervous system, and hypothesised that cross-reactive epitopes can lead to HTLV-1-associated myelopathy/tropical spastic paraparesis. The similar response of antibodies against the coxsackie enterovirus VP1 protein and mitochondrial protein of islet β-cells may be associated with infection-induced diabetes (Coppieters et al. 2013). Association between the AS03-adjuvanted pandemic H1N1 vaccine Pandemrix and narcolepsy is due to cross-reaction of antibodies to the influenza virus nucleoprotein with human hypocretin receptor 2 (Ahmed et al. 2015; Sarkanen et al. 2018). Srinivasappa et al. (1986) hypothesised that infection with influenza virus might stimulate the autoimmune response in tissues from other animals, and heterophilic antigens may trigger this cross-reaction.
The aforementioned diseases and complications are mostly the result of cross-reaction between antibodies that recognise the influenza virus and antigens in the host; common antigens between microorganism and the host may form the basis of the aetiology of these diseases. Hence, determination of heterophilic epitopes on the influenza virus is of great significance for the prevention and control of diseases caused by infection with this virus.
In our previous study, 84 monoclonal antibodies (mAbs) against H1N1 influenza virus hemagglutinin (HA) were produced and two mAbs against HA (A1-10 and H1-84) bound to linear epitopes of the HA antigen and cross-reacted with human brain tissue. It was hypothesised that heterophilic epitopes exist in HA of H1N1 influenza virus as well as human brain tissue (Guo et al. 2015). In the present study, the two mAbs that reacted with human brain tissue were used to confirm the presence of an epitope. The epitope sequence and key amino acids were investigated to understand the pathogenic mechanism of heterophilic epitopes in the nervous system following infection with the influenza virus.
-
The H1N1 influenza virus split vaccine (2009; SFDA approval no. S20090015) was obtained from Hualan Biological Bacterin Co., Ltd. Seasonal H1N1 and H3N2 influenza vaccines (veterinary drug production approval no. [2009] 150132145), and H5N1 and H9N2 influenza vaccines were produced by Yebio Engineering Co., Ltd.
-
Two mAbs of the anti-H1N1 influenza virus HA protein (A1-10 and H1-84) were prepared in our laboratory. The mouse myeloma cell SP2/0 was provided by the Department of Immunology, Air Force Military Medical University, and the culture supernatant of SP2/0 cells was used as a negative control for the identification and localization of A1-10 and H1-84 recognition polypeptides. Horseradish peroxidase (HRP)-conjugated goat anti-mouse antibodies were obtained from Beijing Zhongshan Golden Bridge Co., Ltd.
-
DNAMAN software was used to analyse features of the H1N1 influenza virus HA protein sequence and continuous amino acid sequences were obtained from the NCBI database (https://www.ncbi.nlm.nih.gov/). Subsequently, nine peptide sequences were defined and synthesised by China Peptides Co., Ltd.
-
A total RNA extraction kit and cDNA first-strand synthesis kit were purchased from Tiangen Biotech (Beijing) Co., Ltd. Polymerase for polymerase chain reaction (PCR), the pMD19-T vector, and DNA markers were obtained from Takara Bio. Primer synthesis and sequencing were performed by the Beijing Genomics Institute.
-
An indirect enzyme-linked immunosorbent assay (ELISA) was used to identify cross-reactivity of hybridoma cell culture supernatants of the mAbs A1-10 and H1-84 with the following five HA subtypes: 2009 H1N1 influenza virus split vaccine, the seasonal influenza vaccines H1N1 and H3N2, and the bird flu vaccines H5N1 and H9N2. The culture supernatant of mouse myeloma cell SP2/0 was used as a negative control. Subsequently, the antibodies were categorised into different groups based on the cross-reactivity between the two antibodies and the five HA subtypes.
-
The sequences of the five subtypes of the influenza virus and related information were downloaded from the NCBI database using GenBank IDs (Table 1). Alignment of multiple sequences was performed for the five subtypes using DNAMAN software (version 5.2.2). Nine continuous amino acid sequences, each 5-7 residues in length, were designed as candidate epitopes from the conserved sequence between the five different influenza virus HA protein subtypes (Table 2). The nine peptides were synthesised with a purity of > 85%, as measured by high-performance liquid chromatography and mass spectrometry methods. The candidate epitopes were stored as freeze-dried powder at -20 ℃.

Table 1. Information about the amino acid sequences of subtype influenza virus.

Table 2. Sequence and position of candiate peptides of HA antigens.
-
PyMOL and Swiss-PdbViewer software were used to analyse the distribution of peptides on the HA crystal structure. Swiss-PdbViewer was used to generate a HA protein X-ray crystal texture model, and 3LZG (Protein Data Bank [PDB]), the crystal structure of HA from A/California/04/2009 H1N1 virus, which was similar (> 99%) to that of the antigen under study, was used as the reference. Subsequently, the distribution of the peptides was determined using PyMOL according to the manufacturer's instructions.
-
Using a blocking ELISA, with binding peptides of the two mAbs, the nine candidate epitopes were tested and screened. An ELISA plate was coated with the H1N1 influenza virus HA antigens at a concentration of 2 μg/mL. The synthesised polypeptides were mixed with the mAbs A1-10 and H1-84 separately, and incubated for 1 h at 37 ℃. Subsequently, 100 μL of the mixture was added to each well to coat the ELISA plate containing the HA antigens, and incubated for an additional 1 h at 37 ℃. SP2/0 cell culture supernatant was used as the negative control. After washing three times, 100 μL HRP-labelled goat anti-mouse secondary antibody (1:2500 dilution) was added to each well. The chromagen 3, 3', 5, 5'-tetramethylbenzidine (TMB) was used as the substrate for HRP, with detection of the oxidised product at OD450. The inhibition rate (IR) of binding peptides was calculated using the following formula: IR = (ODCTL - ODTEST)/ODCTL. Correlations between the HA antigen and binding epitopes of antibodies were defined as follows: IR ≤ 0.4, no correlation; 0.4 ≤ IR ≤ 0.8, correlation; and IR ≥ 0.8, strong correlation (Xu et al. 2011).
-
Polyglutamic acid was added to the carboxy terminus of positive peptides to facilitate their coating of ELISA plates (Li et al. 2018). Each well was coated with 2 μg/mL of the peptide and incubated at 4 ℃ overnight. Subsequently, the two mAbs were added at 37 ℃ and incubated for 1 h. SP2/0 cell culture supernatant was used as the negative control. After washing three times, 100 μL HRP-labelled goat anti-mouse secondary antibody (1:2500 dilution) was added to each well, and incubated at 37 ℃ for 1 h. Subsequently, 100 μL TMB-H2O2 chromogenic solution was added to each well and incubated for 10 min at 37 ℃ in the dark. The reaction was terminated using H2SO4 solution (2 mol/L; 50 μL/well). The proportion of bound antibodies, which was correlated with the colour intensity, was measured with an ELISA reader by absorbance at 450 nm. The ratio of each test sample (OD450 to negative control OD450) was calculated. Samples with a ratio of ≥ 2.1 were classified as exhibiting a positive reaction.
-
For the localised peptide, each amino acid of the sequence was individually replaced with an alanine residue, resulting in the synthesis of a collection of alanine replacement peptides (Table 3). A 96-well ELISA plate was coated with 100 μL of 2-5 μg/mL H1N1 influenza virus HA antigen. The two mAbs were incubated with the localised peptide and its family of alanine replacement peptides at 37 ℃ for 1 h. The mixtures were then added to the pre-coated ELISA plate and incubated at 37 ℃ for 1 h. After washing, HRP-labelled goat anti-mouse secondary antibody (1:2500 dilution) was added and incubated for 1 h at 37 ℃. After further washing, TMB colouring solution was added as substrate for HRP. The OD450 values were measured with an ELISA reader. IR was calculated according to the aforementioned formula based on the OD450 values of each well, as a reflection of the reactivities of the antibodies with the original and mutated peptides.

Table 3. Sequences of the alanine scan replacement peptides for HA protein.
-
RNA of the two mAbs was extracted and reverse transcribed to cDNA for cloning. In total, 27 primers were designed by comparing the heavy (H) and light (L) chains of published mouse HA variable regions (Supplementary Table S1). PCR was used to obtain the L-chain gene (approximately 320 bp) of the antibodies with eight primers (including seven forward primers and one reverse primer), and the H-chain gene (approximately 350 bp) of the antibodies with nine primers (including five forward primers and four reverse primers). The PCR-amplified products were subjected to further PCR identification (for the L chain, five primers were designed downstream of the amplification product, and for the H chain, five primers were designed upstream of the amplification product). The positive target L/H-chain sequence was identified (approximately 180 bp for the L chain, and 150 bp for the H chain) and purified by gel electrophoresis. Subsequently, the positive target sequences were ligated into pMD19-T and transformed into competent E. coli (DH5α). Cloned DNAs were confirmed by Sanger sequencing and the corresponding amino acid sequences were determined.
Antigens, Antibodies, and Related Reagents
Antigens
Antibodies
HA protein synthetic peptide
Reagents
Indirect ELISA and Antibody Classification
Screening Epitopes of the Influenza A virus HA Protein
Distribution of Epitopes on the HA Crystal Structure
Localization of Heterophilic Epitopes
Verification of indirect ELISA localization
Key Amino Acid Site Identification
Cloning of Heavy/light Chains of the Variable Region of the Two mAbs
-
Indirect ELISA was performed to characterise the reactivity of the mAbs A1-10 and H1-84 with different subtypes of the influenza virus. As shown in Table 1, both mAbs exhibited broad cross-reactivity with HAs from five subtypes of the influenza virus, including the 2009 influenza A H1N1 influenza virus vaccine, the seasonal influenza virus H1N1 and H3N2 vaccines, and the poultry influenza virus H5N1 and H9N2 vaccines. As a negative control, the supernatant of SP2/0 cell culture showed no reaction with any of the HAs. These results suggested that the epitopes recognised by the mAbs are conserved among the five strains of influenza viruses.
-
The GenBank accession numbers of the five strains of influenza viruses were used to obtain their HA amino acid sequences from the NCBI database. Multiple sequence alignment analysis was performed on the amino acid sequences of HA using DNAMAN software. The conserved sequences were examined as candidate heterophilic epitopes of the viruses that can cross-react with the mAbs A1-10 and H1-84. The nine synthesised peptides (P1-P9) that were estimated by DNAMAN software are listed in Table 2.
An ELISA blocking experiment was performed to identify heterophilic epitopes recognised by mAbs A1-10 and H1-84. P1-P9 candidate epitopes were tested and screened. As shown in Fig. 1A, the IR of the two mAbs binding to peptide 2 (P2) was > 0.8; however, the IR of the two mAbs binding to the other eight peptides (P1, P3-P9) was < 0.4. Indirect ELISAs were also performed to observe reactivity between the two mAbs and P2+; P2+ was synthesised with a 5-glutamic-acid tail (LVLWGIHHPEEEEE), as shown in Fig. 1B. Both of the mAbs reacted with P2+, while no reaction was observed between P2+ and the supernatant of SP2/0 cell culture. The results suggested that the mAbs can only react with P2 (LVLWGIHHP191-199).

Figure 1. Localization analysis of two mAbs against the HA antigen that recognize heterophilic epitopes. A P2 exhibited a higher inhibition rate with the two mAbs compared to the other 8 candidate polypeptides. B The reaction for the two mAbs was also significant for the P2+ polypeptide (synthesized with a 5-glutamic-acid tail). C P2 segment alignment with the HA amino acid sequences of the 5 flu strains. D Distribution of P2 on the HA crystal structure.
The amino acid sequences corresponding to peptide P2 within HAs of the five influenza strains were aligned, as shown in Fig. 1C; the same colour indicates that although the amino acids differed, they belonged to the same class, with similar structure and function. The results showed that peptide P2 was conserved among the five strains of influenza virus, which was the heterophilic epitope of HA recognised by the two mAbs. PyMOL and Swiss-PdbViewer were also used to analyse the distribution of peptide P2 on the HA crystal structure. As shown in Fig. 1D, 3LZG (PDB) was used as the reference structure, and the distribution of P2 on the HA trimer crystal structure is marked in red and located in the β sheets of the HA head region.
-
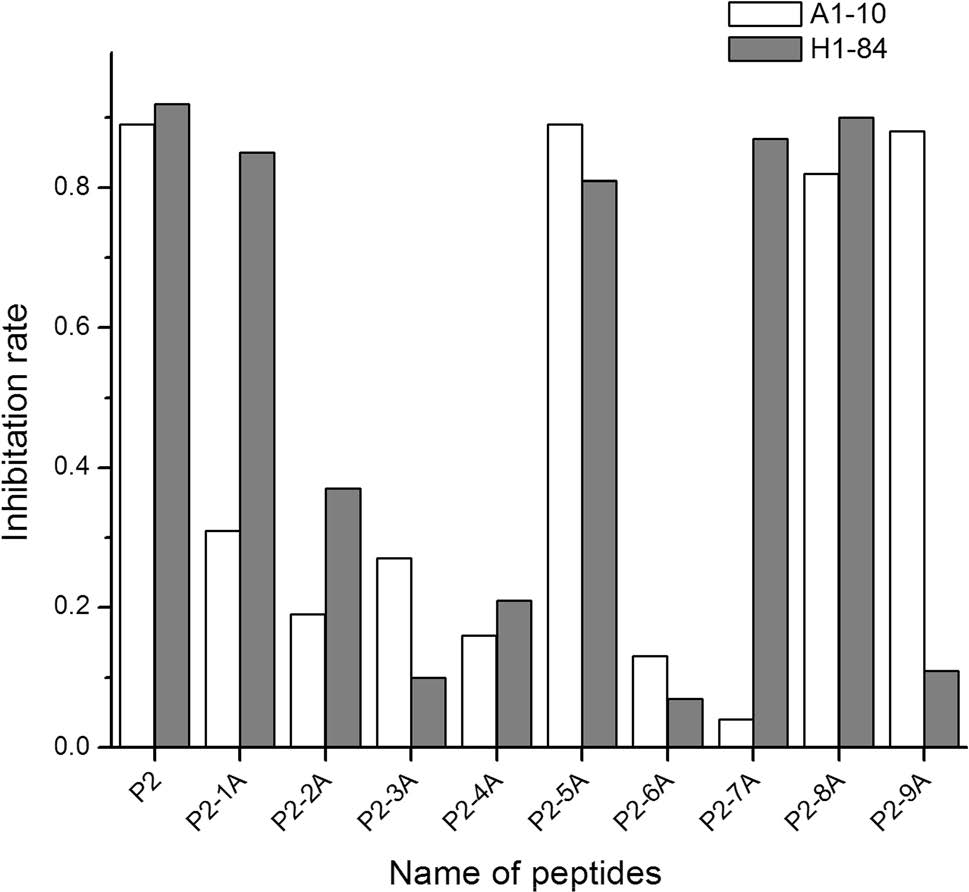
Each amino acid of the heterophilic epitope P2 sequence (LVLWGIHHP191-199) was individually replaced with alanine. As shown in Table 3, there were nine alanine replacement peptides: P2-1A and P2-2A to P2-9A. ELISA blocking experiments were used to identify key amino acid residues for the mAbs A1-10 and H1-84 mapped with different alanine replacement peptides. The IR was calculated by measuring the OD450.
As shown in Fig. 2, when the alanine replacement peptide could not block binding of the two mAbs to HA, the IR was < 0.4, suggesting that the alanine replacement site is a key amino acid site for mAb binding. Conversely, when the alanine replacement peptide could still block binding of the mAbs to HA, the IR was > 0.8, suggesting that the alanine replacement site was not a key amino acid site for mAb binding. Therefore, the amino acids critical for mAb A1-10 recognition were L191, V192, L193, W194, I196, and H197, while the amino acids critical for mAb H1-84 recognition were V192, L193, W194, I196, and P199. We clearly showed that the two mAbs recognise two different epitopes, although the two epitopes have overlapping amino acids.
-
Molecular biological methods were performed to obtain amino acid sequences of the L- and H-chain variable regions (VL and VH) of mAbs A1-10 and H1-84. The VL and VH sequences of the two mAbs are listed in Table 4. abYsis software was used to analyse the complementarity-determining regions (CDRs) and framework regions (FRs) of VL and VH for mAbs A1-10 and H1-84. As shown in Fig. 3, VL and VH contained three CDRs, namely CDR1, CDR2, and CDR3, and four FR regions, namely FR1, FR2, FR3, and FR4. Comparing the amino acid sequences of the CDRs and FRs of VL and VH for mAbs A1-10 and H1-84, it was found that the amino acids in the FRs did not change significantly; however, amino acid changes in the three CDRs of VL and VH were apparent. The three CDRs of VL and VH together constitute the antigen-binding site of the mAb, which recognises and binds the antigen. These results suggested that mAbs A1-10 and H1-84 are derived from different cell clones, and although both mAbs reacted with peptide P2, they recognised different key amino acid sites; i.e., different epitopes.
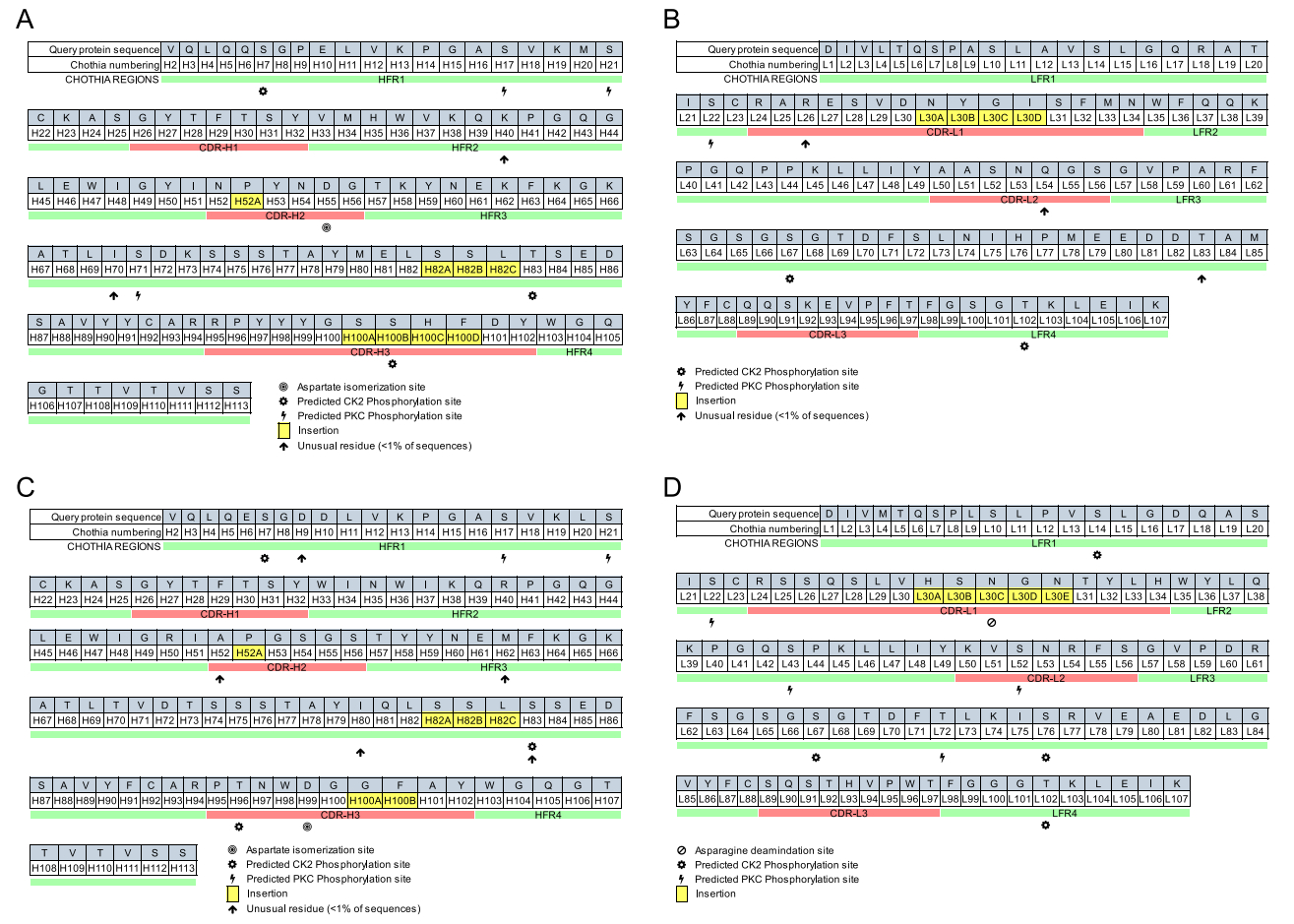
Figure 3. Sequence analysis of the L- and H-chain variable regions of the two mAbs. A A1-10VH, B A1-10VL, C H1-84VH, and D H1-84VL.

Table 4. Sequences of heavy chain and light chain variable region amino acid of two mAbs.
Reaction Characteristics of A1-10 and H1-84 with Different Subtypes of the Influenza Virus
Localization of the mAbs against Influenza Virus HA to Identify Heterophilic Epitopes
Identification of Key Amino Acids for the mAbs Recognising the Heterophilic Epitope
Sequence Analysis of L- and H-chain Variable Regions of the mAbs
-
A number of retrospective studies have demonstrated that patients with severe influenza virus infection often suffer neurological complications due to damage of the central nervous system and other neurological regions (de Wit et al. 2018; Hurwitz et al. 1982; Vellozzi et al. 2014). Furthermore, influenza virus vaccines can stimulate the production of cross-reacting antibodies, leading to central nervous system diseases (Sarkanen et al. 2018). In a previous study, of 84 mAbs against H1N1 influenza virus HA, cross-reactivity with human brain tissue was observed for two mAbs (A1-10 and H1-84) (Guo et al. 2015). Based on this observation, the present study verified heterophilic epitopes between H1N1 influenza virus HA and human brain tissue, and investigated their distribution on the HA crystal structure and the key amino acid residues.
In the present study, an indirect ELISA was used to confirm the presence of common antigens among five influenza viruses. Based on analysis of conserved regions, nine peptides were used to localize the epitopes for the mAbs on the influenza virus HA protein. The distribution of potential epitopes on the HA crystal structure was estimated using PyMOL software. The key amino acid sites of the two mAbs and the variable regions of their antibodies were also identified. The results indicated that H1N1 influenza virus HA and human brain tissues have heterophilic epitopes, which may be one of the causes of nervous system complications following influenza virus infection or adverse reactions following vaccination. The results of this study provide potential insights into the pathogenesis of influenza virus infection and nervous system diseases, and may enhance the sophistication of prevention and control measures for this virus.
Short peptides are good immunogens to study the immunogenicity and reactivity of virus heterophilic epitopes. For example, Li et al. (2018) directly immunised mice with short peptides of the influenza virus and successfully screened and prepared mAbs. Gong et al. (2016) coupled the short peptides P1-P6 of influenza virus H3N2 with keyhole limpet hemocyanin carrier proteins to increase the immunogenicity of peptides and induce a strong immune response. In the present study, determination of epitopes with linear peptides provided an experimental basis for subsequent studies of influenza virus heterotrophic epitopes and the mechanism of nervous system lesions, or adverse reactions following vaccination or influenza virus infection. In addition, using E. coli, different lengths of HA spheres, including amino acids 1-330 and 63-286 of HA, have been successfully expressed, which can induce higher levels of neutralising antibodies following immunisation of animals and protect against viral attacks in vivo (Khurana et al. 2010; Taylor et al. 2011). Therefore, the heterophilic polypeptide epitope LVLWGIHHP191-199, which was targeted in the present study, has the potential to be a neutralising epitope; however, further research is needed.
In the present study, a neutral alanine scanning library was used to identify the key amino acid residues of the mAbs by individually replacing target amino acid residues with alanine (Cong et al. 2010). Using the alanine scanning method, Engmark et al. (2016) identified linear epitopes recognised by antibodies and their key amino acid binding sites by detecting the binding of three different anti-venom serums to the venom toxin epitopes. Seow et al. (2017) also identified epitopes and the critical amino acid binding site of 13 polypeptides on 3D-MSO2, providing a new understanding of the conformation of the C-terminal region of merozoite surface protein 2 (MSP2) on Plasmodium falciparum and useful experimental data for the construction of potential candidate vaccines for recombinant MSP2. In the present study, the results of the alanine scanning confirmed the heterophilic locus recognised by the A1-10 and H1-84 mAbs, and identified their critical amino acid binding sites (L191, V192, L193, W194, I196, and H197 for A1-10; and V192, L193, W194, I196, and P199 for H1-84). These findings provide an important basis for a follow-up study of mutational heterophilic loci and the prevention and control of influenza.
B-cell receptors or antibodies may cross-recognise or exhibit multiple responses due to similar linear or spatial epitopes, which can even induce conformational changes. In addition, the same antibody can recognise different antigens, and the same epitope can be recognised by different antibodies (Hu et al. 2004). Nair et al. (2000) produced the crystal structure of PS1 (HQLDPAFGANSTNPD) from the hepatitis B virus surface antigen and confirmed their common interactive region with antibody CDR H3. Promiscuity of epitope recognition has also been identified in studies based on T-cell receptors and immunoglobulins as model antigens (Marchalonis et al. 2001; Nair et al. 2000; 2002; Robey et al. 2002). In the present study, the CDR sequences of the L- and H-chain variable regions were analysed and it was found that there were differences in the CDRs of A1-10 and H1-84 (Figure 3A, 3C). Although both of the mAbs were associated with P2 (LVLWGIHHP191-199), they bound different key amino acid sites; the underlying mechanism will require further investigation.
In summary, the present study confirmed that the anti-H1 subtype influenza virus mAbs that cross-reacted with the human brain recognised the heterophilic epitope of the HA head region (LVLWGIHHP191-199). The existence of this epitope provides a novel perspective for studying the pathogenesis of neurological diseases caused by influenza virus infection, and therefore may aid influenza prevention and control.
-
This work was supported by The National Key Research and Development Program of China (grant no. 2016YFD0500700), The Natural Science Basic Research Program of Shaanxi Province (grant no. 2016JM8065) and Shaanxi Provincial People's Hospital Incubation Fund Program (grant no. 2015YX-4).
-
CG, XX and LS designed the experiments. CG, JZ, HL, DL, QF, YL and YF conducted the experiments. CG and JH analyzed the data. CG and HZ wrote the paper. All authors approved the final manuscript.
-
The authors declare that they have no competing interests.
-
This article does not include any experiments that involve human or animal subjects.
Conflict of Interest
Animal and human rights statement
-

Table S1. Primers of the variable regions of the heavy and light chains of mouse mAb
















 DownLoad:
DownLoad: