-
Dear Editor,
Epstein-Barr virus (EBV, also termed human herpesvirus- 4) was the first identified human tumor virus. Since its discovery in 1964, studies have shown that EBV infects over 90% of all people by the time they are adults (Williams and Crawford 2006). EBV infection can result in mucocutaneous and systemic diseases, ranging from selflimited illnesses to aggressive malignancies, including B cell Hodgkin lymphoma and nasopharyngeal carcinoma. In vitro, EBV transforms resting B cells into proliferating blast cells (Pope et al. 1968). This transformation depends on the expression of nine viral latent proteins, including latent membrane protein 1 (LMP1). LMP1 expression in B lymphocytes can constitutively result in lymphoma. EBV-positive tumors express LMP1 on their surface (Deacon et al. 1993). These findings have implicated LMP1 as a suitable target for immunotherapy to treat EBV related Hodgkin lymphoma and nasopharyngeal carcinoma.
LMP1 is a membrane protein composed of a cytoplasmic amino-terminus (amino acids 1–24), six transmembrane domains (amino acids 25–186), and a cytoplasmic carboxy-terminal signaling domain (amino acids 187–386) (Fennewald et al. 1984). It plays an orchestrating role in viral infection, tumor transformation, immune modulation, and dissemination of tumor cells in the body (Young and Rickinson 2004). The transmembrane domain of wild-type LMP1 can produce a constitutive phenotype similar to the signaling effect of CD40 or the tumor necrosis factor-2 receptor (Gires et al. 1997). Virions lacking functional LMP1 lost the capability to efficiently transform B cells, which indicates the pivotal role of LMP1 as the primary oncogene of EBV (Kaye et al. 1996; Dirmeier et al. 2003). LMP1 is expressed in most EBV-associated lympho-proliferative diseases and malignancies, and it critically contributes to their pathogenesis and phenotypes. Approaches used to target LMP1 include inhibiting its expression by DNAzymes (Lu et al. 2008) and its downstream signaling molecule, c-Jun N-terminal kinase (Kutz et al. 2008). T-cells specifically recognizing EBV-encoded LMP1 can be isolated from HLA-A*0201 transgenic mice immunized with the minimal epitope LMP1166 (TLLVDLLWL). These T cells have potential applications in adoptive transfer therapy (Cho et al. 2018). One of the limitations in the development of a LMP1-targeting strategy for cancer immunotherapy is a lack of specific and potent antibodies against LMP1. The current commercially available monoclonal antibodies only recognize the intracellular domains of LMP1 and are not suitable for therapeutic targeting. So far, only one reports identified an antibody Fab (HLEAFab) against LMP1 extracellular domain to target nasopharyngeal carcinoma (Chen et al. 2012). Although innate and adaptive responses against EBV encoded proteins have been identified in EBV infected and EBV+ cancer patients, only very weakly detectable and occasional responses to LMP1 epitopes have been observed (mainly the CD4+ memory response) in a small minority of allele positive donors (Hislop et al. 2007). Thus, LMP1 might have intrinsic immune evasion functions (Hislop et al. 2007). In healthy carrier controls, circulatingCD4+ andCD8+ T memory cells against LMP1 are present at levels equivalent to those seen in nasopharyngeal carcinoma patients (Lin et al. 2008). Since no specific IgG response against LMP1 has been reported, here, we attempted to screen antibodies with low affinities to recognize LMP1 extracellular domains from a phagedisplay, naïve human, antigen-binding IgM fragment (hereafter, Fab) library. Naïve IgM libraries from healthy individuals display the true unbiased nature of the repertoire. Although the libraries are somewhat unfocused in targeting antigens, they are efficient for generating antibodies against disease-specific antigens (Omar and Lim 2018).
To screen the Fab library, we first constructed a vector which contained a DNA fragment encoding the LMP1 gene extracellular domain segments (residues 45–51, 97–106, and 161–163, separated by amino acid GGGS for each segment) fused with Fc fusion protein (LMP1-Fc) in a backbone vector pSegTag as described previously (Xiao et al. 2009). The plasmid encoding LMP1-Fc was transfected into 293T cells and the secreted LMP1-Fc protein was purified by Protein A agarose beads as bait for subsequent library panning.
The Fab library and its panning protocols were described previously (Ying et al. 2014). The antibody library was first panned with purified Fc protein to remove Fc binding clones, and then panned against LMP1-Fc fusion proteins for four more rounds. Significant enrichment against LMP1-Fc was observed after three rounds of panning (Fig. 1A). Four panels of 96 clones were picked from rounds 3 and 4, and monoclonal phage ELISA was performed. Fourteen clones were obtained (Fig. 1B) with >two-fold increases in the optical density at 405 nm when LMP1-Fc was used as antigen, as compared with Fc as the antigen. All clones were sequenced for the VH and VL regions and confirmed by the signature protein sequences (YYC…TFGGG for VH, and YYC…WGQGTMVYVS-S for VL). Sequencing revealed that the VH and VL insertions in clone 10-B2 and 12-C2 were identical, and clone 7-G9 and 9-F3 contained same DNA fragments for the VH and VL insertions. 11 Fabs was successfully purified by His tag followed the protocol described previously (Ying et al. 2014). The binding activity was further confirmed with serially diluted antigen (Fig. 1C). Among the 14 clones, clone 1-A11, 10-B2, and 15-H10 displayed strong binding activity against LMP1-Fc, but not to control protein Fc and bovine serum albumin at the highest concentration used for LMP1-Fc. The binding activity declined along with the antigen dilution.
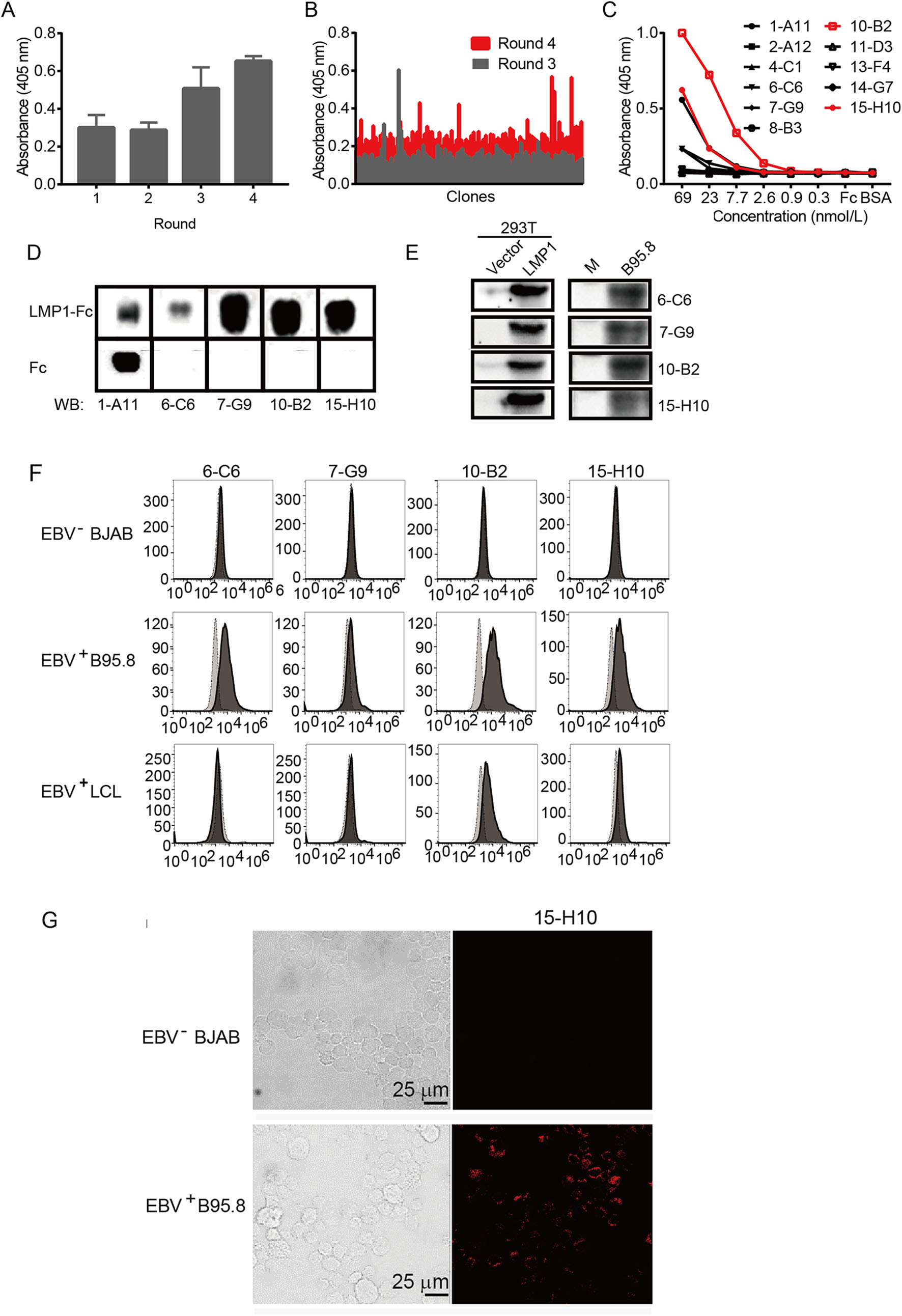
Figure 1. Isolation and characterization of LMP1 Fab from a naïve antibody phage display library. A Polyclonal phage ELISA showing the binding of the first to fourth rounds of phages to LMP1-Fc. Bound phages were detected with anti-M13-HRP conjugate. B Monoclonal phage ELISA showing the binding activity of single phage clones from rounds 3 and 4 to antigen LMP1-Fc. Bound phages were detected with anti-M13-HRP conjugate. C Binding of individual Fabs to LMP1-Fc as various concentrations. D Fabs, except for 1-A11, could recognize LMP1-Fc, but not Fc. E Fabs as indicated could detect LMP1 exogenous expressed in 293T and endogenous expressed in B95.8 cells. M stands for no cell lysate mock. F Flow cytometry data showing staining of EBV+ B95.8 cells and LCL cells by Fab 6-C6, 10-B2, and 15-H10 Fabs. G Confocal images of EBV+ B95.8 and EBV- BJAB cells stained with 15-H10. Scale bar 25 μm.
The recognition of the selected clone against LMP1-Fc was further confirmed by western blotting. The phage plasmid containing the Fab fragment with both His and Flag tags was transformed into Escherichia coli HB2151, and Fab was expressed and purified by His column chromatography (Qiagen, Germany). Western blotting data showed that Fabs from clones 6-C6, 7-G9, 10-B2, and 15-H10 recognized the antigen LMP1-Fc, but not Fc itself, which indicated that recognition involved Fab and the LMP1 extracellular domain (Fig. 1D). Fab from clone 1-A11 bound to both LMP1-Fc and Fc. To determine whether the Fab clones could bind to full length LMP1 in its natural form, we cloned the LMP1 gene from B95.8 EBV+ cells and ectopically expressed it in 293T cells. Western blotting using Fabs was performed to demonstrate their recognition of LMP1 expressed by 293T cells. A single band of approximately 70 kDa was detected from 293T cells transfected with LMP1, but not the empty vector control, by Western blotting with Fabs from clones 6-C6, 7-G9, 10-B2, and 15-H10 (Fig. 1E). The data confirmed that all Fabs from these four clones detected full length LMP1.
We wanted to determine whether selected Fabs could detect the endogenous level of LMP1 expressed at the cell surface of EBV infected B95.8 cells by Western blotting and flow cytometry. EBV+ B95.8 cell lysates were analyzed through Western blotting with Fabs from clones 1-A11, 6-C6, 10-B2, and 15-H10 (Fig. 1E). A band at the predicted size was detected. Flow cytometry analysis also indicated 1-A11, 6-C6, 10-B2, and 15-H10 bound specifically to EBV+ B95.8 cells, but not to EBV- BJAB cells (Fig. 1F). We also validated the effectiveness of the ability of the clones to recognize another EBV+ lymphoblastoid cell line (LCL). The Fab clones 10-B2 and 15-H10 displayed a distinctive shift compared to the control devoid of secondary antibody. To confirm the binding of Fab to the LMP1 cell surface protein, we also performed an immunofluorescence assay and visualized the binding of the antibodies using confocal microscopy. B95.8 and BJAB cells were fixed and incubated with the purified Fabs as primary antibodies and phycoerthyrin-anti-FLAG as secondary antibody for staining. Only 15-H10 showed a strong signal with B95.8 cells, but it did not show a signal with BJAB cells (Fig. 1G); the remaining three clones did not generate an appreciable recognition signal.
Here we isolated the Fab clones 1-A11, 6-C6, 10-B2, and 15-H10 and confirmed their recognition of LMP1 expressed on the surface of EBV+ B95.8 cells. Recognition of LMP1 on the cell membrane by clone 15-H10 was verified through immunofluorescent assay, while other clones failed the recognition. One of the possible reasons could be the stringent washing in this assay that ruptured the low affinity Fab-LMP1 binding. EBV infection is related with multiple diseases, and conventional approaches used in laboratory diagnostic tests for EBV infection and pathogenesis have their limitations [(generate false positive results or lack the ability to localize the expression of EBV within target cells) (Young and Rickinson 2004)]. The LMP1-specific Fab clones reported here need further evaluation in both affinity and specificity in testing EBV+ patient samples, and hopefully it could have potential applications in EBV diagnostics and directly targeting EBV-related tumors in adoptive T cell therapy.
HTML
-
This work was supported by the National Natural Science Foundation of China (Grant Numbers: 81402542 and 81772166) and the scholarship of Pujiang Talents in Shanghai to Fang Wei (Grant Number: 14PJ1405600).
-
The authors declare that they have no conflict of interest.
-
This article does not contain any studies with human or animal subjects performed by any of the authors.














 DownLoad:
DownLoad: