-
Dear Editor,
Hepacivirus C virus (HCV) is a major cause of liver disease in human, which is estimated to infect 2%–3% of the world's population and has resulted in a high global health burden. HCV was first discovered in 1989, and was the sole member of the Hepacivirus genus before 2011. Recently, genetically-diverse HCV-like viruses have been independently determined in a number of animal species, expending the members of the genus Hepacivirus to 14 species (Hepacivirus A–N) as recommended by International Committee on Taxonomy of Viruses (ICTV) (Hartlage et al. 2016; Smith et al. 2016).
In 2019, a novel HCV-like virus (duck hepacivirus-like virus, DuHV) was first identified in ducks in five cities/provinces in China by random amplification and reverse transcription (RT)-PCR (Chu et al. 2019). As a newly determined HCV-like virus, no further study on DuHV was reported in other regions in China or other countries. Many HCV recombination events have been reported worldwide (Gonzalez-Candelas et al. 2011). An intra-subtype recombination event in HCV-like virus in equines (equine hepacivirus) was previously determined by us (Lu et al. 2019). However, no recombination events in other HCV-like viruses have been reported.
There are increasing evidences supporting the crossspecies transmission of HCV-like viruses. Origins of HCV may involve one or many zoonoses of HCV-like viruses (Pybus and Theze 2016). Like HCV, HCV-like viruses presented hepatotropic characteristic (Pfaender et al. 2015; Trivedi et al. 2018; Baechlein et al. 2019). Consumption of raw or undercooked duck livers from the infected animals pose a potential human health threat. In addition, it is still unclear whether DuHV could infect other avian species. The present study was performed to (1) investigate whether novel HCV-like virus was present in avian livers in meat markets in China and (2) detect the potential recombination events in the viruses.
During September to November in 2019, a total of 296 avian liver samples from individual animals (116 chicken, 72 duck, 108 goose) sold in five meat markets in Guangdong Province were carefully collected to avoid contamination and then processed for further study. Briefly, total RNA of each sample was extracted and then reverse transcribed into cDNA using a HiScript II 1st Strand cDNA Synthesis Kit (Vazyme, China). Then, standard PCR targeting the DuHV NS2–NS3 partial genomic region was performed using GenStar Taq Polymerase Premix (Kangrun Chengye, China) based on primer pairs of F/R-TONG (Chu et al. 2019).
After separated on 1% agarose gel electrophoresis, PCR products from three duck liver samples (obtained from two independent markets) presented the expected bands of 272 bp. The sequencing results and subsequent BLAST hits in the NCBI database (https://www.ncbi.nlm.nih.gov/) indicated that all three duck liver samples were DuHV-positive. The NS2–NS3 partial genomic of the three DuHV strains had a genetic similarity of 91.5%–98.5% between each other, and had a genetic similarity of 91.5%–100% with the previously published Chinese strains (HCL-1, HCL-2, and HCL-3). According to the meat sellers, all the investigated livers were obtained from domestic animals in Guangdong Province, thus providing the first evidence supporting DuHV's prevalence in field ducks in Southern China.
To further understand the genomic characteristics of DuHV in Southern China, one field DuHV strain was obtained by gap-filling PCR and sequenced in the present study, and was named GD-61. Briefly, a total of eight primer pairs covering the genome of published DuHV strains were designed using Oligo 7.0 (Supplementary Table S1). After PCR amplification, the DNA fragments were cloned into a pCloneEZ-blunt plasmid vector (CloneSmarter, USA), and then transformed into E. coli DH5α competent cells (Weidi, China). The genome of GD-61 was assembled by SeqMan 7.1.0 based on the sequencing raw data. The polyprotein gene of GD-61 has been submitted in the GenBank database with the accession number MT135177.
The polyprotein gene of GD-61 obtained in the present study was 10, 824 nucleotides in length. Compared with HCL-1, HCL-2, and HCL-3, no nucleotide insert/deletion was found in the genome of GD-61. The G + C content of the polyprotein gene of GD-61 (49.69%) was similar with HCL-1 (49.84%), HCL-2 (49.46%), and HCL-3 (49.57%). The polyprotein of GD-61 was predicted to be cleaved into 3 structural proteins (Core, E1, and E2) and 7 nonstructural proteins (p7, NS2, NS3, NS4A, NS4B, NS5A, and NS5B) with bioinformatics analysis, demonstrating the same genome organization of GD-61 with other HCV/HCV-like viruses.
As recombination events can mislead the genetic similarity and phylogenetic analysis, we investigated whether potential natural recombination events occurred within DuHV or between DuHV and HCV/HCV-like virus. Recombination events were detected by RDP 4.95 based on seven methods (RDP, GENECONV, Chimaera, MaxChi, BootScan, SiSican and 3Seq) and a P value of < 0.01. The signal was considered to be recombination if it was detected by at least four of the seven methods. Two recombination events within the DuHV strain HCL-2 were finally determined, which occurred in the Core gene, 5' terminal of E1 and 3' terminal of NS5B (event 1) and 3' terminal of NS2 and 5' terminal of NS3 (event 2), respectively (Supplementary Table S2). The result demonstrated HCL-2 was a recombinant strain. Its major and minor parent strains were HCL-1 and GD-61 respectively. The break points were determined to be at positions 1029, 3050, 3297, and 8281 (Fig. 1). As revealed by Bootscanning and Simplot analysis, the Core gene, 5' terminal portion of E1 and NS3, and 3' terminal portion of NS2 and NS5B of HCL-2 was provided by GD-61, and the remaining genomic region of HCL-2 was provided by HCL-1 (Fig. 1A, 1B). The two recombination events were further supported by phylogenetic and genetic similarity analysis based on five recombinant and non-recombinant regions (Fig. 1C, 1D). Therefore, HCL-2 was not included in further genetic similarity and phylogenetic analysis based on complete polyprotein gene.
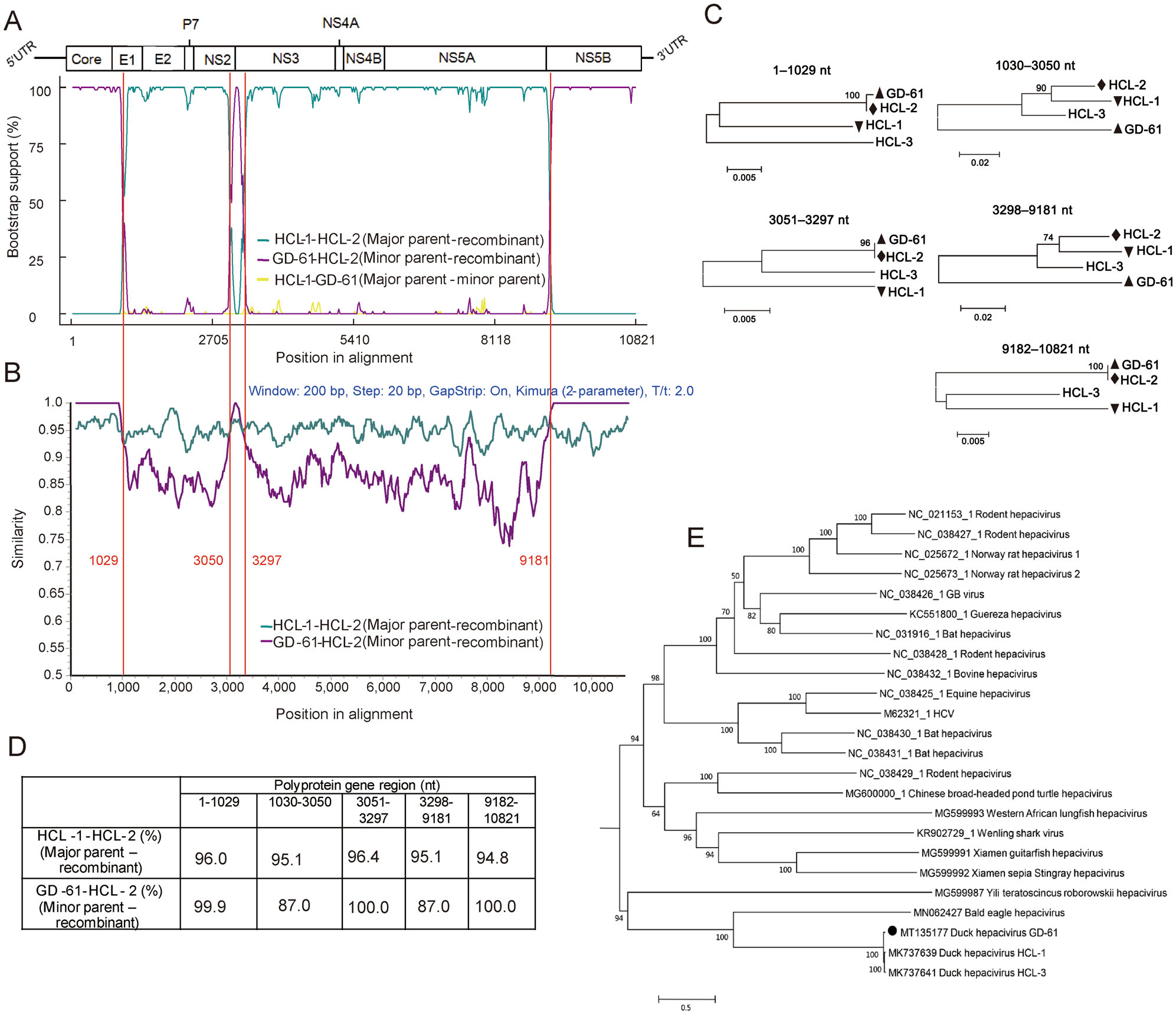
Figure 1. Recombination events occurred in DuHV strain HCL-2 but not GD-61 or HCL-1 (A–D), and phylogenetic analysis of novel isolated DuHV strain GD-61 (E). A Bootscanning analysis of HCL-2. B Simplot analysis of HCL-2. C Phylogenetic analysis of five recombinant and non-recombinant regions. GD-61, HCL-1, and HCL-2 were indicated by an upright triangle, an upside-down triangle, and a diamond, respectively. D Genetic similarity analysis of HCL-1, HCL-2, and GD-61 based on five recombinant and non-recombinant regions of polyprotein gene. E Phylogenetic analysis of GD-61 based on polyprotein sequences. To understand the evolutionary relationship between GD-61 and other HCV/HCV-like viruses, their polyprotein sequences were aligned and used for phylogenetic analysis. After estimated by a "Find Best Protein Models" program, rtREV + G+I + F model was chosen for further phylogenetic analysis using MEGA 5.05. The maximum-likelihood phylogenetic tree was constructed based on 1000 bootstrap replicates. GD-61 was indicated by a circle.
The polyprotein gene of GD-61 had a genetic similarity of 89.0% and 89.3% with HCL-1 and HCL-3 respectively, indicating GD-61 was a genetically divergent strain. However, DuHV strains were classified into one genotype and one subtype according to genotyping and subtyping reference values of HCV/HCV-viruses (Lu et al. 2019). Accordingly, HCL-2 was an intra-subtype recombinant strain. In addition, a total of 73 unique amino acid substitutions were observed in the GD-61 polyprotein, and 63 substitutions were located in NS5A (Supplementary Table S3). When compared with other HCV/HCV-like viruses, the polyprotein gene of GD-61 had a highest genetic similarity of 46.1% with Wenling shark virus, and a lowest genetic similarity of 24.5% with Yili teratoscincus roborowskii hepacivirus. Therefore, GD-61 is a highly genetically divergent HCV-like virus and should be classified as a novel hepacivirus-related, unassigned flavivirus. To understand the evolutionary relationship between GD-61 and other HCV/HCV-like viruses, their polyprotein sequences were aligned and used for phylogenetic analysis (Fig. 1E). It demonstrated that DuHV strains were clustered together in a group. In addition, DuHV and Bald eagle hepacivirus had a close evolutionary relationship, indicating a similar evolutionary pattern.
In summary, DuHV RNA was detected in duck livers in meat markets in China for the first time. Further study still needs to fully understand the prevalence and zoonotic potential of DuHV. Recombinant events were determined in multiple genes of DuHV, providing a new insight into the evolutional pattern of the virus.
HTML
-
This work was supported by the Guangdong Provincial Natural Science Foundation under Grant [Number 2017A030310367].
-
The authors declare that they have no conflict of interest.
-
All institutional and national guidelines for the care and use of laboratory animals were followed.
















 DownLoad:
DownLoad: