HTML
-
Owing to the advent of new technology enabling effective culturing and antigenicity screening of single B cells (Wu et al. 2010; Scheid et al. 2011), many HIV broadly neutralizing antibodies (bnAbs) with high potency have been recently isolated (Scheid et al. 2009; Simek et al. 2009; Walker et al. 2009; Moore et al. 2011; Huang et al. 2012) By targeting vulnerable conserved sites of the native HIV-1 envelope glycoprotein (Env), a trimeric protein comprising two non-covalently associated subunits namely gp120 and gp41, bnAbs can effectively antagonize a broad range of circulating HIV-1 strains by blocking their cell entry (Burton and Mascola 2015). The fact that approximate 15%-20% of patients develop bnAb responses 2-4 years after HIV infection lend strong support to the notion that a bnAb-raising vaccine would be approachable (Pantaleo and Koup 2004; Doria-Rose et al. 2009).
In order to elicit nAb, an Env-based immunogen should be a trimer mimicking the prefusion form (Julien et al. 2013a, 2013b; Pancera et al. 2014; Lee et al. 2016). In retrospect, the failure of early-generation HIV vaccine can be largely attributed to the exclusive use of gp120 as the immunogen, which is a non-native monomer with an epitope profile distinct from that of a native-like trimer (Flynn et al. 2005; Pitisuttithum et al. 2006). Through extensive efforts focusing on engineering Env protein to restrain its metastability, several types of stabilized Env trimers with native-like epitope profile were identified, all of which were based on gp140, a truncation form of gp160 ending at position 664 with the removal of membrane attachment sequence to attain solubility (Ward and Wilson 2017). Gp145, which contains the entire membrane-proximal external region (MPER) and is thus membrane-bound, is another form of g160 that has also been explored in HIV vaccine development but has not attracted much attention (Barban et al. 2012; Emmer et al. 2016). However, the potential cooperativity between gp140 and gp145 for induction of protective HIV antibody response is highlighted by a previous study showing that a heterologous immunization combining DNA-based gp140 as prime and Tiantan vaccinia (TV)-based gp145 as boost was significantly more effective than homologous gp140 or gp145 immunization in the induction of neutralizing antibodies in mice and guinea pig (Liu et al. 2012). Since this study only tested one Env, which is derived from HIV-1 clade C strain CN54, the generality of gp140-gp145 combination strategy has yet been proven. More recently, Saunders et al. presented a vaccination approach using a combination of gp145 DNA priming and gp140 protein boost, where heterologous tier 2 bnAb response was attained in one out of four rhesus macaques that were vaccinated (Saunders et al. 2017).
A major milestone in the creation of gp140-based native-like trimers is represented by SOSIP design. This design is anchored on two metastability-reducing modifications, namely the introduction of an inter-subunit disulfide bond (termed SOS) to covalently connect gp120 to gp41 and an Ile-to-Pro substitution at position 559 of gp41 (termed IP) to promote the gp41-gp41 homointeraction (Binley et al. 2000; Sanders et al. 2000, 2002). The first product of SOSIP design, BG505 SOSIP.664, displayed hitherto unseen features of native-like trimers, effectively binding to a variety of bnAbs while showing little or no affinity to non-neutralizing antibodies (Julien et al. 2013b; Sanders et al. 2013). Immunization studies in rabbit and non-human primate (NHP) subsequently showed that BG505 SOSIP.664 trimer is able to effectively engage an autologous tier 2 nAb response, representing a huge leap from autologous tier 1 nAb response attained with gp120 monomers (Sanders et al. 2015; Klasse et al. 2016; Pauthner et al. 2017). However, only limited success has been achieved in terms of the induction of heterologous tier 2 nAb responses (Saunders et al. 2017, 2019).
As the SOSIP design has continued to evolve by incorporation of additional stabilizing mutations, advances were also made in creating cleavage-independent gp140 trimers. To date, three designs of cleavage-independent gp140 trimers have been developed, namely sc-gp140 (single chain-gp140) (Georgiev et al. 2015), NFL (native, flexibly linked) (Sharma et al. 2015), and UFO (Uncleaved prefusion-optimized) (Kong et al. 2016), on the common basis of replacement of the furin cleavage site by a flexible linker in a SOSIP gp140 template. Structural and biochemical analysis revealed that the overall antigenicity of the cleavage-independent trimers resembles that of SOSIP trimers, recognizing a wide range of bnAbs preferentially over non-neutralizing antibodies (Sanders and Moore 2017; Ward and Wilson 2017).
The cleavage-independent gp140 trimers are particularly valuable for the development of a particle-based vaccine (Sliepen et al. 2015; He et al. 2016). By displaying antigen in a regular array on the surface, a particulate vaccine allows multivalent ligation of B-cell receptor, thereby enhancing the engagement of cognate B-cells (He et al. 2016; Tokatlian et al. 2019). Particulate display of antigen has an additional bonus for conformation-dependent antigen like Env trimers, as antigen moieties in proximity may interact with each other, resulting in reciprocal stabilization (He et al. 2018). The most widely used nanoparticle platform is that based on ferritin, which naturally self-assemble into NP consisting of 24 monomers (Lin et al. 2005; He et al. 2016). Previous studies on BG505 SOSIP.664 trimers have shown that changing from soluble form to ferritin particle form could lead to a consistent, albeit moderate, increase in the immunogenicity (Sliepen et al. 2015). Sauders et al. further demonstrated that SOSIP Env trimers can be presented on ferritin NPs with the same antigenic profile, and the immunization of resulting nanoparticles is capable of elicitating tier 2 nAb featuring improbable somatic mutation critical for neutralization breath (Saunders et al. 2019). Compared to their cleavage-dependent counterparts, the cleavage-independent gp140 trimers are conceived to be more amenable to ferritin-particle design because their processing does not require furin-mediated cleavage, which is deemed to be less effective on compact repeats of trimers displayed on ferritin nanoparticle than on free trimers due to steric hindrance effect (Sliepen et al. 2015; He et al. 2016).
In this study, using both mouse and rhesus macaque model, we progressively optimize a vaccination regimen founded on both gp140 DNA prime-gp145 Tiantan vaccinia (TV) boost modality and utility of heterologous Env sequences. We found that this modality, although effective in mice for induction of nAb reponse, requires an additional NP-based protein boost to be effective in rhesus macaque. Consequently, we identified a novel short 8-week vaccination schedule capable of raising a tier 2 nAb response with substantial breath in rhesus macaque. A proof-of-principle study further revealed that viremia could be eliminated or effectively controlled in immunized animals upon SHIV challenge. Our study support gp140-gp145 sequential immunizations, aided by adoption of heterologous Env sequences, as a promising approach for HIV vaccine development.
-
DNA vaccines expressing RL42-gp140 and RL42-gp145 were constructed by cloning the target genes into pVRCSV1.0 vector. Recombinant vaccinia viruses encoding AE2F-gp140 and AE2F-gp145 were derived from China Tiantan virus strain (752-1) and were abbreviated as rTVs. Viruses were propagated in Vero cells, released by freeze-thaw, and purified using 36% (m/v) sucrose cushion. The viral titers were determined by plaque assay on thymidine kinase-negative (TK-) 143 cells. The immunogen expression by the generated DNA- and rTV-based vaccine vectors were respectively verified by western blotting analysis of transiently transfected HEK293 cell and infected Vero cells using sera from HIV-1 infected individuals.
-
A mammalian expression vector of AE2F SOSIP.664 was constructed by fusing a 6 × His-SpyTag sequence to the amino terminal end of AE2F SOSIP.664 sequence. The construct was transiently transfected into HEK293T cells, which were harvested 48 h later for purification of the soluble expressed 6 × His-SpyTag-AE2F SOSIP.664 proteins by nickel chelating chromatography (GE Health Care, Piscataway, NJ, USA) under native condition. VRC01 and 10E8 monoclonal antibodies were purified using Ab Spin Trap Protein G sepharose columns (GE Healthcare) following the manufacturer instructions. Protein purity was assessed by Coomassie-blue staining after separation denaturing gel electrophoresis followed by Coomassie blue staining, and the content of SpyTag-fused AE2F SOSIP.664 preparation was further analyzed by size exclusion chromatography. SpyCatcher-fused ferritin protein was kindly provided by Prof. Mingzhao Zhu (University of the Chinese Academy of Sciences, Beijing, China). The SOSIP.664-Ferritin nanoparticles were produced by mixing SOSIP.664-SpyTag with SpyCatcher-Ferritin at a molar ratio of 1:1, and then desalted on Amicon Ultra-30 centrifugal filter (Millipore, Billerica, MA, USA) into PBS.
-
Female C57BL/6 mice (6 weeks old, 18-22 g) were bought from the B & K Universal Group Ltd. (Shanghai, China). Three-four years old Female Chinese rhesus macaques, free of simian immunodeficiency virus, simian T lymphotropic virus type 1, B virus, simian retrovirus and tuberculosis, parasites (e.g. Entoameba), were purchased from a domestic source (Ningbo, China). Only the rhesus monkeys that did not express the Mamu-A∗01 allele, as determined by PCR screening, were selected for the vaccination study. Animals were maintained in accordance with national regulations on the Management of Laboratory Animals (GB14922- 2001).
-
The immunization schemes were individually specified in related figures. For mouse immunization, all the vaccines were diluted with appropriate amount of PBS for intramuscular delivery into the tibialis with 100 μL volumes. The amount of vaccine per administration was 100 μg, 1 × 107 pfu, and 10 μg respectively for DNA vaccine, rTV vaccine and NP protein vaccine. Animals were sacrificed 2 weeks after the last immunization, and sera were collected for antibody quantification.
For immunization of rhesus macaques, all the vaccines were intramuscularly injected into the leg with fixed volumes of 1 mL per shot. Each DNA and rTV vaccination used 1 mg of DNA and 1 × 108 pfu of virus respectively. The SOSIP.664-Ferritin nanoparticles were adjuvated with equal volume of alum shortly before application at a final protein concentration of 0.1 mg/mL, translated into 100 μg per immunization. Sera were harvested 2 weeks after each immunization to assess HIV-specific antibody response. At 2 weeks after the final immunization, animals were challenged intravenously with 1000 TCID50 (50% tissue culture infectious doses) of SHIV89.6 virus, followed by sera collection at various times points post infection to determine plasma SIV RNA viral load using real-time RT-PCR as described previously (Tian et al. 2015).
-
HIV-1 CN54 gp120, RL42 gp120, and AE2F gp120 proteins were expressed in HEK293T cells, purified, and dissolved in sodium bicarbonate buffer (pH 7.4) at a final concentration of 0.5 μg/mL. In the case of ferritin, AE2F SOSIP.664 and AE2F SOSIP.664-Ferritin proteins, preparations were made at 1 μg/mL. For ELISA assays, the target antigen(s) were added in 100 μL volumes to flat-bottom 96-well plates (Costar, New York, NY, USA). After overnight incubation at 4 ℃, the coated wells were washed with PBS, then blocked with PBS + 5% milk before adding 100 μL volumes of twofold serial dilutions of sera or monoclonal antibodies to be analyzed (sera starting at a 1:100 dilution and monoclonal antibodies starting at 10 μg/mL). The subsequent secondary antibody incubation and OPD-based color development was conducted following normal ELISA protocol, and the absorbance was then read at 492 nm on a plate reader (Multiscan Ascent, Thermo Corporation, Vantaa, Finland). The endpoint titers were defined as the highest dilution that yielded an OD twofold higher than control.
-
Env-pseudotyped viruses were produced from HEK293T cells by transient transfection of a mix of rev/env expression plasmid and an Env-deficient HIV-1 backbone vector (pSG3Δenv) using Turbofect (Thermo Scientific, Cat# R0532). The virus-containing supernatants were harvested 48-72 h post transfection and filtered through a 0.45 μm filter for cell debris removal. The filtered supernatants were aliquoted and stored at - 80 ℃ after the addition of FBS to a final concentration of 10%. Titering of Env-pseudotyped virus stocks were performed on TZM-bl cells (NIH AIDS Reagents Program) following published protocol (Montefiori 2005). SVA-MLV were kindly provided by Dr. Feng Gao (Jilin University, Jilin, China). VSV-G virus stocks were propagated and tittered on Vero E6 cells. The procedures for pseudovirus neutralization assay were the same as titration assay except that serially diluted sera was added to assay wells preceding the addition of a dose of Env-pseudotyped virus pre-determined for optimal infectivity. The 50% inhibitory dose (ID50) was defined as the reciprocal of the serum dilution that leads to 50% reduction of Relative Luminescence Units (RLU) relative to control wells with only virus addition after subtraction of background value (measured in mock-infected well).
-
All statistical analyses were conducted using GraphPad software (Prism 5, San Diego, CA, USA). The between group difference was determined by either unpaired Student's t test or Mann-Whitney tests. Significant difference was defined as *P < 0.05, **P < 0.01.
Design and Production of DNA- and Recombinant Vaccinia Virus-based Vaccines
Generation of AE2F SOSIP.664-Ferritin Nanoparticles Using SpyTag/SpyCatcher Ligation System
Animal Maintenance
Animal Immunization and Virus Challenge Studies
ELISA Assay for Determination of HIV-1 Envelope-specific Antibody
Env-pseudotyped Viruses and TZM-bl Cell-based Neutralization Assay
Statistical Analysis
-
The sequential combination of gp140 and gp145 within a DNA prime-rTV boost modality was previously reported to be more effective in inducing HIV neutralizing antibodies than gp140 or gp145 as the sole immunogen. That is, sv140 + rTV145 or sv145 + rTV140, raises significantly higher HIV nAb response than the homologous sv140 + rTV140 or sv145 + rTV145 combinations in mice and guinea pig (Liu et al. 2012) This finding was made on gp140 and gp145 both derived from HIV-1 CN54 strain. We first examined whether the same conclusion can be made on another Env sequence. To this end, we chose Env from RL42 strain-one of the earliest primary B′ clade HIV-1 strains identified in the Yunnan Province of China in the 1990s. To construct gp140- or gp145-encoding vaccines in either sv1.0-based DNA platform or recombinant Tiantan vaccinia (rTV) platform. The resulting two DNA vaccines and two rTV vaccines, denoted as sv140 and sv145, rTV140 and rTV145 respectively, were validated for their expression of encoded Env forms in transfected or infected cultured cells (Supplementary Fig. S1).
Following the scheme shown in Fig. 1A, we examined in mice three DNA prime-rTV boost regimens, including two immunizations with gp140/gp145 as the immunogen for both prime and boost (group 1 & 3) and one immunization with sv140 as the prime and rTV145 as the boost (group 2) (Fig. 1A). Mice administered empty DNA vector as prime and empty vaccinia vector as boost served as mock control. Sera were isolated at 3 weeks after the boost; all the samples from the same group were volume equally combined and the HIV neutralizing activities were subsequently evaluated by standard TZM-bl assay against a panel of five HIV strains including a B clade virus (SF33), a B' clade isolate (B020100419), and three B'/C clade isolates (XJDC 0793, XJDC 6371 and XJDC 6431). The sera from the sv140 + rTV145 group (group 2) neutralized all the five HIV-1 viruses at a titer of 1:180. In contrast, only moderate serum neutralizing activities were detected in the sv145 + rTV145 group (group 3) against four viruses (SF33, B020100419, XJDC 6431, XJDC 0793) with a titer significantly lower than that of the sv140 + rTV145 group, whereas the sv140 + rTV140 group (group 1) failed to display detectable neutralization activity against any of the five viruses (Fig. 1B). As the reference, the mouse serum neutralizing activities previously reported for CN54 Env-based sv140 + rTV145 immunization were 1:60 for XJDC 6371 and < 1:20 for the rest viruses. Thus, the RL42-version of sv140 + rTV145 immunization regimen was able to elicit an even greater HIV nAb response than the original CN54 version, thereby supporting the candidacy of sv140 prime-rTV145 boost as a promising HIV vaccine modality.
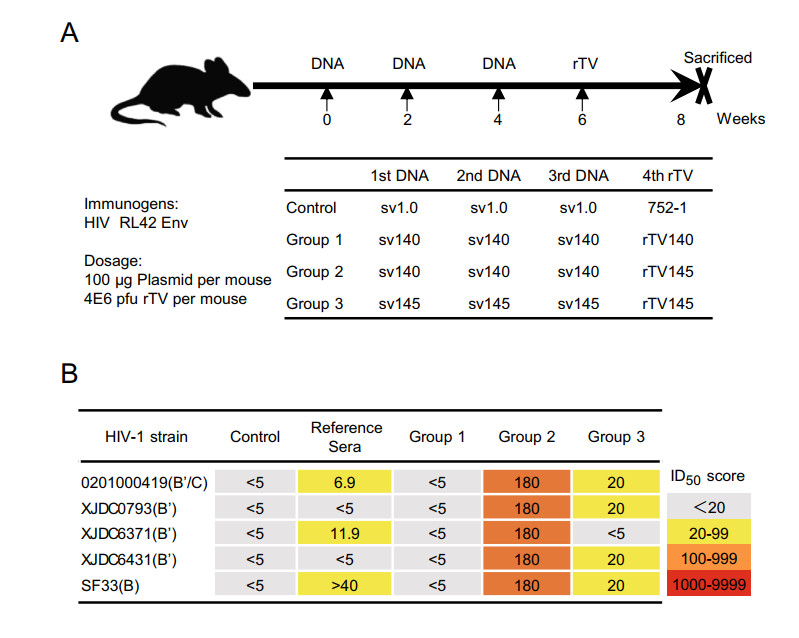
Figure 1. A regimen combining gp140 DNA prime and gp145 vaccinia-vectored boost elicited strong HIV neutralizing antibody responses in mice. (A) Schematic illustration of immunization protocol and animal groups. (B) End-point neutralizing titers. Sera were collected 3 weeks after the last immunization, and their end-point nAb titers were determined against a panel of five pseudoviruses from three clades using TZM-bl assay. The sera from a patient with HIV infection was used as reference sera.
-
Next, we evaluated the efficacy of sv140 + rTV145 immunization in non-human primate (NHP)-rhesus macaque setting. It is widely recognized that the induction of HIV nAb in NHP is more challenging than small animal model like mice (Sanders et al. 2015; Hessell et al. 2016). Thus, we turned to an enhanced version of sv140 + rTV145 modality wherein the Env sequences from three HIV strains of different subtypes, namely AE2F, CN54, and RL42, were used to construct three DNA vaccines and three rTV vaccines, together building a regimen comprising a triple heterologous rTV145s preceded by a heterologous triple sv140s. Following the scheme shown in Fig. 2A, we applied this regimen to China-origin rhesus macaques (Macaca mulatta) (group 1) in parallel with a homologous AE2F-derived sv145-gp145 regimen (group 2), and collected sera at various time points across the immunization course for analysis of gp120 binding activity and HIV neutralizing activity.
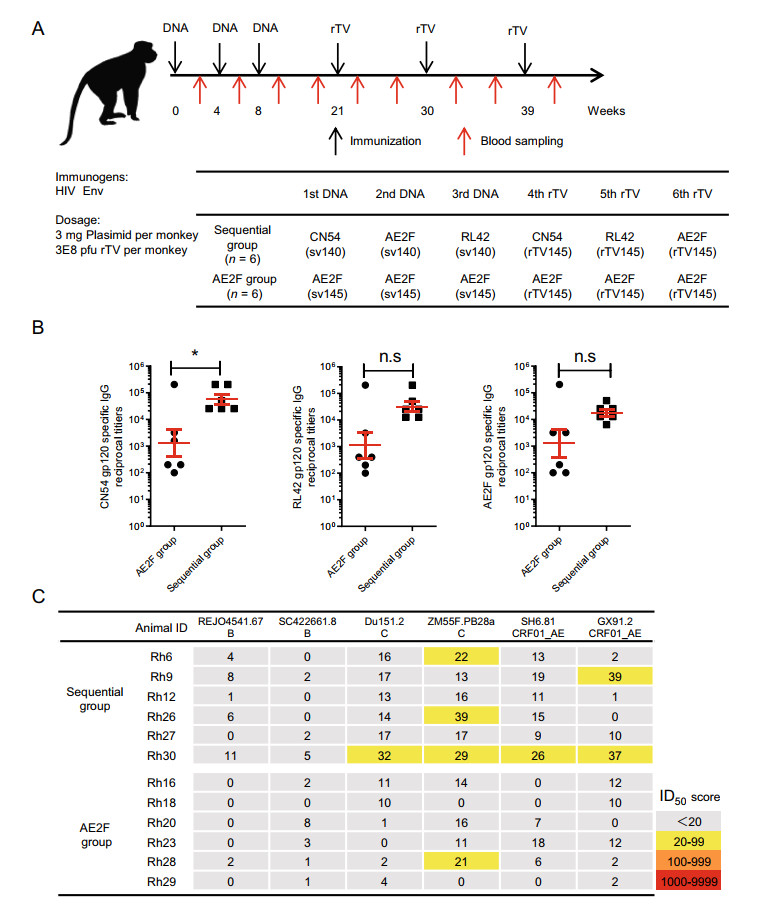
Figure 2. The sv140 + rTV145 sequential immunization was less effective in inducing HIV nAb responses in rhesus macaques. A Schematic illustration of immunization and blood sampling schedule, and animal group information. Sera were collected 2 weeks after the last immunization and analyzed for the presence of gp120 binding antibodies and HIV neutralization antibodies. B Endpoint titers of gp120 binding antibody determined by ELISA against gp120 proteins derived from CN54, AE2F or RL42 strains. The endpoint titers were expressed as the reciprocal of the highest dilution showing an absorbance two standard deviations above the control sera. C Endpoint neutralizing titers against a panel of six viruses representing B, C and CRF01_AE clades, determined by TZM-bl assay. Bars represent mean ± SEM. Statistical significance was evaluated by Student's t test and a P value < 0.05 was considered significant. *P < 0.05; n.s: not significant.
The serum binding antibody titers for autologous gp120s at the end of immunization were generally higher in group 1 than group 2. This difference is most evident for anti-CN54 gp120 antibody, with group 1 showing a titer 32-fold higher than group 2. As for antibody against RL42 or AE2F gp120, the titers of group 1 were also substantially higher than those of group 2, albeit statistically non-significant (Fig. 2B).
In contrast to autologous gp120 antibody response, both groups showed poor neutralizing activities against a panel of 6 heterologous HIV strains. A careful examination revealed a better performance of group 1 than group 2 (Fig. 2C). Among sera from group 1, one sample (Rh30) exhibited a relatively broad neutralizing activity against four viruses and there were also three samples (Rh6, Rh9, Rh26) capable of neutralizing a particular virus. In contrast, only one animal (Rh28) in group 2 displayed a moderate strain-specific neutralizing activity against ZM55F.PB28a virus (subtype C). Overall, despite attaining a strong antibody response to autologous gp120, sv140 + rTV145 immunization was ineffective in inducing HIV neutralizing antibodies in the NHP setting.
-
Given the limited efficacy of sv140 + rTV145 immunization in the NHP model, we sought to test whether improvement can be made by adding a potent second boost step, for which we opted to use SOSIP.664-Ferritin nanoparticles. SOSIP.664 design, was the first design capable of generating soluble and cleaved gp140 trimers with a native-like antigenic profile (Julien et al. 2013a; Sanders et al. 2013). Nanoparticle represents an effective strategy to increase the antigenicity of difficult antigens, and ferritin has been explored as a NP platform to boost the efficacy of SOSIP.664 vaccine with some success (Sliepen et al. 2015; He et al. 2016, 2018; Tokatlian et al. 2019).
We consider the newly developed cleavage-independent SOSIP a better choice for preparation of SOSIP ferritin NP as it bypass the furin requirement, that is, for the original cleavage-dependent version of SOSIP, furin protease is needed to be co-expressed at a high level to achieve efficient cleavage of SOSIP for preserving their native conformation(Sliepen et al. 2015; He et al. 2016). Specifically, we adopted a design of single-chain soluble SOSIP gp140 trimer in which the furin cleavage site is replaced with a flexible (GGGGS)3 (de Taeye et al. 2016) linker (Fig. 3A). We also switched to an in vitro platform for ferritin NP assembly inspired by the recently developed SpyTag/SpyCatcher chemistry where SpyTag and SpyCatcher, two reactive fragments derived from CnaB2 protein from Streptococcus pyogenes, can spontaneously bond to each other under mild conditions through iso-peptide formation. Accordingly, SpyTag-tagged SOSIP trimers and Spycatcher-fused ferritin can be individually expressed and purified, and subsequently mixed to form NP (Fig. 3B). The separation of SOSIP trimers from ferritin is envisioned to increase the flexibility of NP production platform.
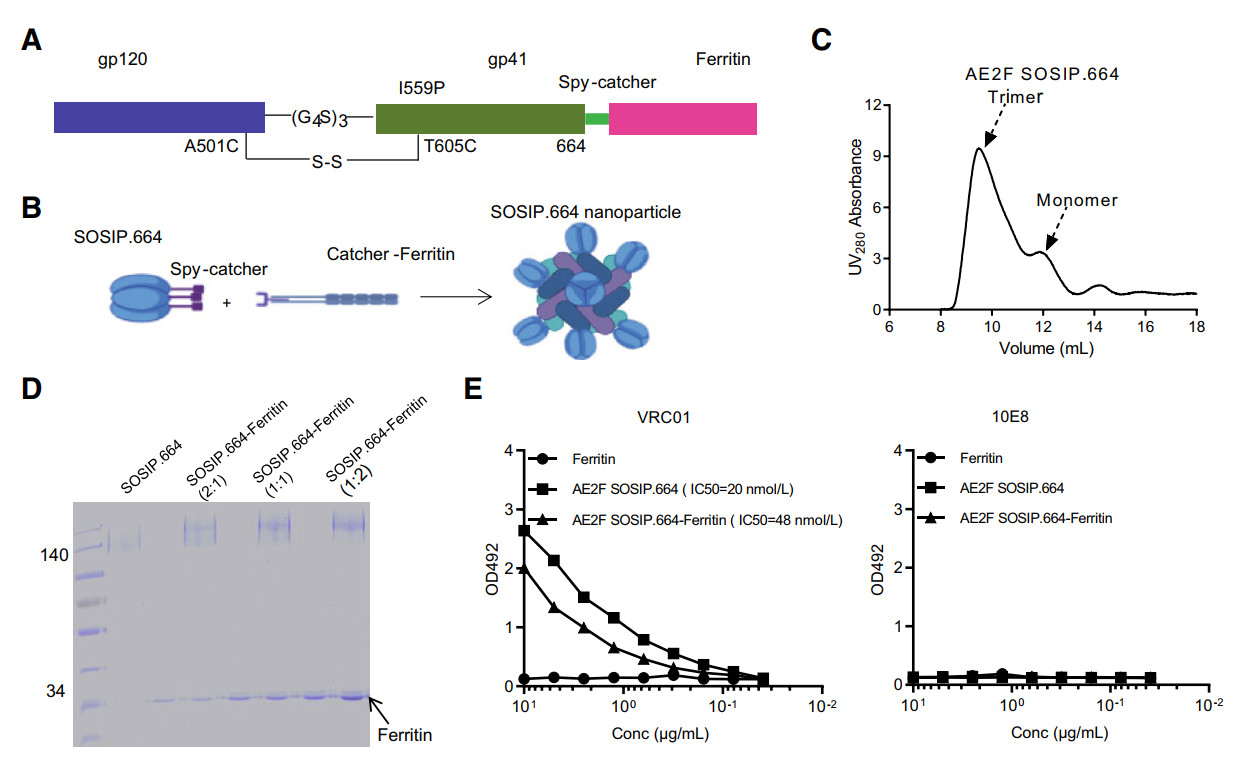
Figure 3. Design and production of SOSIP.664-ferritin nanoparticle using a two-component click approach. A Schematic representation of the basic unit of the SOSIP.664-ferritin nanoparticle used in this study. B Cartoon illustration of the SpyTag/SpyCatcher click system to assemble SOSIP.664-ferritin nanoparticle in vitro. C Size exclusion chromatography (SEC) profile of the purified AE2F SOSIP.664-SpyTag protein. D Optimization of reaction condition for generation of SOSIP.664-ferritin nanoparticles. The SOSIP.664-SpyTag and SpyCatcher-Ferritin proteins were mixed at three different molar ratios as indicated for overnight reaction at 4 ℃. The reaction products were subsequently separated by denaturing gel electrophoresis, followed by detection with Coomassie-blue staining. E Epitope recognition by SOSIP.664 protein and SOSIP.664-ferritin NP. ELISA assay was used to assess the binding of ferritin, SOSIP.664 protein, and SOSIP.664-ferritin NP to VRC01 bnAb and 10E8 bnAb, which respectively recognize the CD4 binding site and the MEPR region. Data are representative of three independent measurements.
Using this click approach, we prepared ferritin NPs displaying SOSIP.664 with gp140 sequence derived from AE2F (AE2F SOSIP.664). The SOSIP and ferritin components were respectively generated by transfecting his-tagged AE2F SOSIP.664- SpyTag and his-tagged SpyCatcher-Ferritin plasmid into 293 T cells, and purified with nickel-chelate chromatography. The resulting AE2F SOSIP.664-SpyTag preparation was analyzed by size exclusion chromatography (SEC), revealing a major peak in the fraction corresponding to SOSIP trimers. A minor peak corresponding to monomeric gp140 was also detected (Fig. 3C). To find the optimum condition for producing the AE2F SOSIP.664-Ferritin adduct, we performed small-scale reactions in which purified AE2F SOSIP.664-SpyTag and SpyCatcher Ferritin proteins were mixed at various molar ratios before incubation at 4 ℃ overnight. The reaction was separated by denaturing SDS-PAGE and then analyzed by Coomassie blue staining. The result indicated that approximately 80%-90% of the SOSIP.664-SpyTag input was conjugated to ferritin when the molar ratio of SOSIP.664 SpyTag/SpyCatcher-Ferritin was 1:1. Only a slight increase in conjugation efficiency was achieved by increasing the molar ratio of SOSIP.664 SpyTag/SpyCatcher-Ferritin to 1:2 (Fig. 3D). Consequently, we chose SOSIP.664 SpyTag/SpyCatcher-Ferritin molar ratio 1:1 as the reaction condition to perform a large-scale preparation of SOSIP.664-Ferritin NP. We also assessed the epitope presentation of the prepared NP by measuring their reactivity to HIV antibodies recognizing different sites on Env protein. As expected, SOSIP.664-Ferritin NP bound strongly to VRC01, a highly cross-reactive bnAb targeting CD4 binding site while showing little affinity for 10E8 bnAb, which recognizes an epitope in the MPER region (Fig. 3E). Thus, the SOSIP.664-Ferritin NP prepared by our approach likely preserved the antigenicity of the displayed SOSIP trimers.
-
According to the aforementioned sequential gp140-gp145 vaccination strategy, a SOSIP.664-Ferritin NP boost, which is based on gp140, needs to pair with gp145-encoded vaccine(s). We conceived a sequential administration of DNA-gp145 and rTV-gp145 as a modality to deliver a strong antibody response to HIV gp145; we also postulated that enriching the immunogenic heterogeneity could facilitate the ability of a sequential immunization to generate a broad nAb response. Consequently, we designed an immunization strategy involving three vaccines applied sequentially: first, a DNA vaccine carrying RL42-gp145 sequence; second, a rTV vaccine encoding AE2F-gp145; last, the AE2F SOSIP.664-Ferritin NP. We tested this DNA prime-rTV boost-NP boost regimen in a cohort of six rhesus macaques. Plasma and PBMCs were isolated before the immunization and 2 weeks after the first and second boost, and subsequently analyzed for the anti-HIV antibody response (Fig. 4A). Robust anti-gp120 antibody response was detected in all immunized animals after the rTV boost with an average titer of approximately 1:10, 000, which was further enhanced by tenfold after the NP boost (Fig. 4B). We also used the TZM-bl assay to assess the HIV neutralizing activity of post-NP boost sera relative to pre-immune sera. Among the six vaccinated animals, five (ID 1 & 3-6) displayed serum neutralizing activity against all the three autologous pseudoviruses examined (Fig. 4C). We next used a standardized panel of tier 2 HIV viruses to further assess the neutralization breadth. Importantly, the sera from three (ID 1, ID 3, ID 5) immunized animals showed cross-reactivity to 9 or 10 viruses, though the titers were generally low. Two other animals (ID 4, ID 6) displayed a narrower neutralizing spectrum restricted to 2-3 viruses (Fig. 4D). Overall, the breath of neutralization was positively correlated with the magnitude of autologous neutralization response. Measurements of pre-immune sera and immunized sera against murine leukemia virus (SVA-MLV) and VSVG virus demonstrated low background of TZM-bl neutralization assay, justifying the use of 20 as the cutoff for the presence of neutralization activities (Supplemantary Fig. S2). Collectively, by adding a potent heterologous protein boost in the NP form, we were able to significantly enhance the capability of the DNA-rTV modality to induce a broad tier 2 nAb response in rhesus macaques.
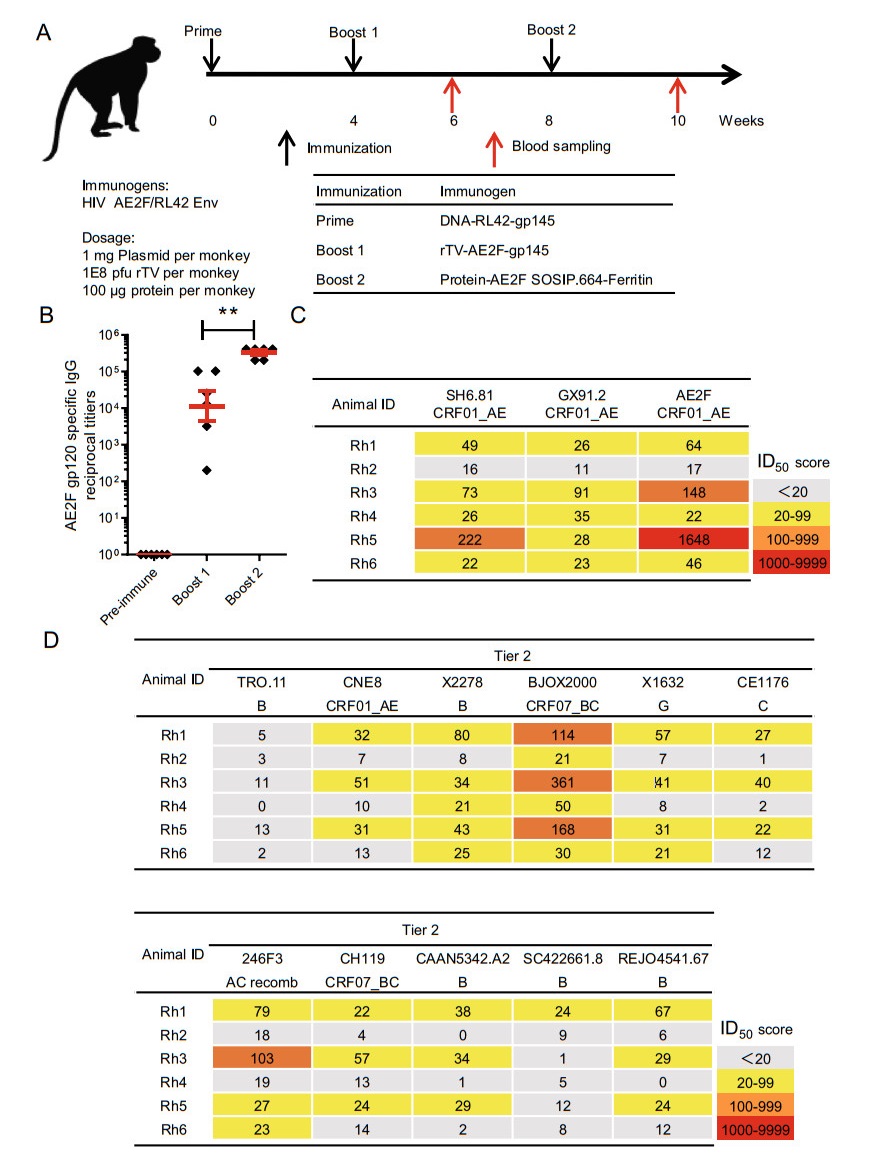
Figure 4. Addition of SOSIP.664-ferritin nanoparticle to the sv140 + rTV145 modality led to an improved HIV nAb response in the immunized rhesus macaques. A Schematic illustration of immunization and blood sampling schedule, and animal group information. B Effects of booster vaccinations on the gp120-binding IgG antibody titers. Serum samples were collected before immunization and 2 weeks after each booster vaccination, and assayed for gp120-binding IgG antibody titers by ELISA using AE2F gp120 protein. C, D Assessments of vaccine-induced serum neutralizing activity. Sera collected 2 weeks after the completion of vaccination were subjected to TZM-bl assay against either three CRF01_AE viruses (C) or a global reference panel consisting of 11 representative tier-2 viruses (D). (Yellow, ID50 > 20; orange, ID50 > 100; red, ID50 > 1000). Bars represent mean ± SEM. **P < 0.01, Student's t test.
-
Lastly, we sought to investigate whether the antibodies induced by the three-step sequential immunization strategy are protective against SHIV challenges in rhesus macaques. To this end, the six immunized rhesus macaques as described above were intravenously challenged with a single dose of 1000 TCID50 of heterologous SHIV89.6 2 weeks after the last immunization, with three unimmunized animals serving as the sham control group. Longitudinal monitoring of plasma viremia revealed that all the sham animals had a sustained viral load in the range of approximately 3 × 104-8 × 104 copies/mL. In contrast, among the immunized group, one animal (No. 6) displayed an undetectable viremia while the rest five animals showed a transient rise in viral loads, peaking at various time points after virus challenge, which were subsequently subdued over time. Importantly, even the highest observed peak viremia value among immunized animals was lower than the lowest viremia value displayed by sham group (Fig. 5). Thus, the sequential DNA-rTV-ferrtin NP regimen showed the potential to afford protection against SHIV challenge in rhesus macaques.
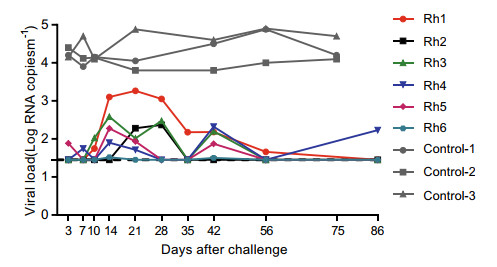
Figure 5. The sequential DNA gp145-rTV gp145-ferritin SOSIP.664 regimen with heterologous Env sequences afforded viremia control against SHIV challenge in rhesus macaques. The six immunized animals shown in Fig. 4 were subjected to a single intravenous injection of SHIV89.6 virus at 1000 TCID50 2 weeks after the completion of vaccination. The plasma viral loads were longitudinally monitored through the detection of gag RNA using quantitative RT-PCR.
Gp140 DNA Prime-gp145 Vaccinia Boost Provides a General Modality for Effective Induction of nAb Responses in Mice
The sv140 Prime-rTV145 Boost was Ineffective in Inducing HIV nAb Responses in Rhesus Macaques
Effective Production of SOSIP.664-Based Ferritin Nanoparticle by a Newly Developed Two-Component "Click" Approach
Addition of SOSIP.664-Ferritin NP to the sv140 + rTV145 Modality Boosted the Induction of a Broad HIV nAb Response
The Sequential DNA-rTV-ferrtin NP Regimen Afforded Viremia Control Against SHIV89.6 Challenge in Rhesus Macaques
-
Raising a heterologous tier 2 nAb response remains a daunting task for HIV vaccine development. In this study, we demonstrated the utility of diverse HIV-1 Env immunogens in a sequential immunization scheme toward this goal. Unlike most of current HIV vaccine studies which solely focus on gp140 construct(s), we explored the combination of gp140 and gp145 as immunogens in a sequential vaccination scheme. Our rationale is that gp145, which is larger than gp140 by the possession of the entire membrane-proximal external region (MPER) to be membrane bound, might more closely resemble the Env spikes on HIV virions. Accordingly, a sequential combination of gp145 and gp140 might provide a better chance for directing the B-cell response at the conserved neutralizing epitopes over non-neutralizing immunodominant epitopes. Such nAb-focusing effect is expected to be further enhanced by utilization of heterologous Env immunogens.
Using Env sequences derived from clade B'/C CN54 strain, we first generalized the notion that, for a DNA prime-rTV boost regimen, the combination of gp140 and gp145 as prime and boost succeeded in potently inducing autologous nAb antibodies while either gp140 or gp145 as the sole immunogen failed. The capability of DNA gp140 prime-rTV gp145 boost regimen to induce a nAb response in mice is remarkable, given previous documentation that mice were less responsive to immunization with BG505 SOSIP.664 trimers than other animal species for the generation of autologous tier-2 antibody response (Sanders and Moore 2017). However, the success of DNA prime-rTV boost regimen in mice did not translate into rhesus macaques. The vaccinate animals displayed at most sporadic induction of a narrow tier 2 nAb response despite high autologous binding antibody titers. This result is consistent with the general consensus within the field that raising a desirable antibody response in non-human primates is more challenging than small animal models.
We further tested whether the failure of DNA gp140 prime-rTV gp145 modality in rhesus macaque can be rescued by additional vaccination step, for which we tested a strong protein boost in the form of ferritin nanoparticle. The potency of NP-based vaccine lies in its presenting the loaded immunogen in multivalent form, conferring the avidity advantages over monovalent soluble vaccine for a successful B-cell activation. In this study, we explored a two-component click approach to produce SOSIP.664-presenting ferritin NP based on the SpyTag/SpyCatcher system (Wang et al. 2019). This newly developed click method is an attractive alternative to the conventional self-assembly method, especially in the case where the construct expressing the native-like trimers is already available. Besides flexibility, this method has a potential "hidden" advantage in trimer-presentation capacity. For the self-assembly approach, the maximal number of trimers displayed per ferritin NP is eight while that for the click method might extend to twenty-four. This increased trimer-presentation capacity could make a difference in the induction of nAb response.
The addition of ferritin-based SOSIP.664 NP to the DNA-rTV modality formulated the ultimate regimen showing encouraging effectiveness in rhesus macaques. With only three inoculations completed over 8 weeks, this regimen raised a HIV-specific antibody response featuring robust autologous binding antibody titers, neutralizing activities against autologous pseudoviruses, and more importantly consistent (4/6 animals) development of tier 2 nAb response of considerable breadth. However, we would like to point out that we adopted a cutoff ID50 of 20 in our study to fully capture the weak tier-2 antibody responses. Some of the previous studies used a higher cutoff, making it difficult to compare them with our studies. Using a heterologous SHIV challenge, our pilot study revealed that the new regimen led to an effective suppression of subsequent SHIV infection. Among six vaccinated animals, one animal exhibited essentially no development of viremia while the rest showed a consistent, effective virus containment despite a temporary virus aeeumnlation. Intriguingly, our data also points to that there might be other immune response(s) other than nAbs also contributing to the viremia control conferred by our new vaccination regimen, as the Rh6 monkey, despite showing the most potent inhibition of SHIV89.6 infection, did not exhibit a higher nAb activity. Along the same line, it has been previously reported for other vaccine approaches that protection against viral challenges can be achieved in the absence of nAb induction through the action of non-neutralizing antibodies (Barouch et al. 2013, 2015; Alter and Barouch 2018).
In summary, we explored the complementation between gp145 and gp140 as immunogens in raising Env-specific nAb response, leading to the identification of a short three-step immunization scheme capable of eliciting a tier-2 nAb response of substantial potency and breath in rhesus macaque. The efficacy of this scheme certainly awaits further validation in future study of a larger animal sample size and other SHIV infection model. It is also important to gain a better understanding of the mechanistic aspect of anti-HIV immunity raised by our vaccines, including determination of the immune correlates for the protection and the nature of elicited neutralizing antibodies: the B cell composition and the targeted site of vulnerability on Env protein. These explorations will provide instructive guidance on how to improve our vaccination strategy toward a vaccine capable of effective protection. Notwithstanding, based on the study presented here, we proposed that diversification of Env immunogens, in both types and formulations, under the framework of a sequential immunization scheme is a promising approach for the development of HIV vaccine.
-
This work was supported by the National Natural Science Foundation of China (81672018, 81561128008), the National Basic Research Program of China (973program#2014CB542502), the National 13th Five-Year Grand Program on Key Infectious Disease Control (2017ZX10202102), Shanghai Pujiang Program (19PJ1409100), and Intramural Funding from Shanghai Public Health Clinical Center.
-
JX and XZ conceived, designed and supervised the study. XD, KC, YW, and JW participated in study design, conducted the majority of the experiments, and analyzed the data. QC, YZ, MZ, RT, WW performed some experiments; CZ contributed to data analysis. XD, CZ, JX, and XZ wrote and finalized the manuscript. All authors reviewed and approved the manuscript.
-
No conflicts of interest to be declared.
-
Animal and Human Rights Statement Research was conducted in compliance with the Animal Welfare Act, adhering to the principles stated in the Guide for the Care and Use of Laboratory Animals, NRC Publication, 2011 edition. The protocols were reviewed and approved by the Ethics Committee of Shanghai Public Health Clinical Center (SPHCC).
















 DownLoad:
DownLoad: