HTML
-
For decades, contaminated and misidentified cell lines in biological research remains a serious problem which the researchers call it "identity crisis" (Masters J R, 2009). As reported in March 15, 2010 by U.S. Food and Drug Administration (FDA), DNA from porcine circovirus type 1 (PCV1) is present in the GlaxoSmithKline (GSK)Rotarix® vaccine (Victoria J G, et al., 2010). The vaccine produced in the Vero cell line which differentiated from African green monkey kidney cell is an orally administered live and attenuated rotavirus vaccine indicating for the prevention of rotavirus gastroenteritis in infants. It is likely that the vaccine was contaminated by PCV1 through porcine-derived trypsin used for digestion of Vero cells during the process of cell culture. In view of the potential risk of infection of PCV1 to humans, the FDA recommended on March 22, 2010 that public health professionals and clinicians in the United States temporarily suspend the use of Rotarix®. Such viral contamination has a negative impact on the safety and clinical use of biological drugs.
With the current rapid advances in biotechnology, a large number of biologics are being produced from cell lines in vitro. For example the anti-cancer monoclonal antibody 806 (ch806), produced from mouse fibroblast cells (Scott A M, et al., 2007); Xyntha, a recombinant anti-hemophilic factor Ⅷ, produced from Chinese hamster ovary (CHO)cells (Eriksson R K, et al., 2001); and rabies vaccines from vero cells (Li R, et al., 2013). These kinds of biologics finally lead to clinical use. However, murine-derived viruses usually gain entry through infection of animals (mice) or murine-derived products such as sera, cells, or transplantable tumors, and these viruses then indirectly become potential pathogens to humans (Macy J D, et al., 2011). Thus, it's an essentially important task to screen the contamination of viruses in working cells and the related-biological productions, and further guarantee the safety of biological drugs and pharmaceuticals.
In consideration of emerging contamination problem, the 2010 edition of the Chinese Pharmacopoeia has been revised and added items to detect viral contamination in murine monoclonal antibody products. Chinese Pharmacopoeia has demanded eight of murine viruses including hemorrhagic fever virus (HFV), lymphocytic choriomeningitis virus (LCMV), mammalian orthoreovirus 3 (MRV-3), ectromelia virus (ECTV), sendai virus (SeV), pneumonia virus of mice (PVM), retrovirus and murine adenovirus 1 (MAdV-1).
The classic detection methods suggested by the Chinese Pharmacopoeia include three kinds of assays: cytotoxicity test, animal antibody test, and ch ick embryos infected assay. These classic detection methods are widely accepted because they are truly reliable and seldom show false-positive results. However, the protocols are complicated and the assays are time-consuming (Jacoby R O, et al., 1996). We developed a series of real-time quantitative PCR assays for the rapid detection of five mouse viruses, which are based on SYBR Green I, with conserved virus gene sequence used as the target sequence. These real-time PCR assays are obviously superior to classical methods, as they are less time-consuming, easier to handle and have good reproducibility.
In this study, we developed new assays for detection of five murine viruses: ECTV, SeV, PVM, MAdV-1 and mouse parvovirus 1 (MPV-1).
In order to achieve the aforementioned advantages, we designed a batch of primers and finally screened out five pairs of ideal primers (Table 1). The design of the primers is the key to this real-time PCR assay. The primers were designed according to the highly conserved region of non-structural genes such as viral polymerase gene. The primers generate short PCR products, ranging from 128 to 176 bp, so the amplification time can be limited to 40 min or less. The real-time PCR can be rapidly finished.
Table 1. Sequences of the primers.
For an example of detection of MPV-1, our testing assay based on real-time quantitative PCR reaches three advantages. The similar results were obtained for the other four viruses.
First, the assay is fast. The primers were designed to generate a short PCR product (only 132bp in length. Thus, the assay took only 39 min to perform. The thermal cycling parameters used were: 1 cycle at 95℃ for 2 min, followed by 40 cycles at 95℃ for 10 s and 60℃ for 45 s.
Second, specificity is high. The melting curve (Figure 1A)shows that Tm reaches a peak at 83℃, and the single sharp peak means high specificity as well. To further assess the specificity of the assay, positive and negative samples were used. The positive MPV-1 genome DNA resulted in a product with correct size while the negative control (deionized water) did not generate any products regardless of the number of PCR cycles. We also tried to detect the normal cell genome DNA, and did not show any amplification of PCR-targeted gene, indicating that the PCR products are simply specific and the assay is highly reliable.
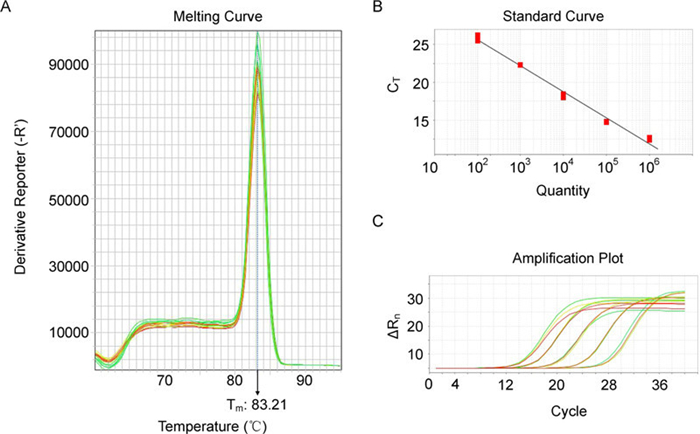
Figure 1. Tenfold serial dilutions of purified template DNA (100 to 106 copies/μL) were tested with SYBR Green I on the StepOneTM Real-Time PCR System. A: Melting curve shows that melting temperature is about 83℃. B: Regression analysis was performed on the copy number vs. threshold cycle (CT) values against log copy numbers. Because 1 μL of plasmids was used for each concentration in each run, the numbers of copies are represented as 102, 103, 104, 105, and 106. Tenfold serial dilutions of purified template DNA from 0.65 fg/μL (1×102 copies) to 6.5 ng/μL (1×106) were tested. The values are shown with the linearity coefficient (R2=0.989). C: Three duplicating amplification of MPV-1 template DNA extracted from plasmid dilution.
Third, this quantitative method is accurate. Tenfold serial dilutions of purified standard template DNA (plasmid pTA-MPV DNA constructed by inserting a 132bp PCR product of MPV-1 into pTA2 vector) from 0.65 fg/μL (1×102 copies) to 6.5 ng/μL (1×106) were tested. The values are shown with a good linearity coefficient (R2=0.989) (Figure 1B). The detection limit was evaluated with purified plasmid pTA-MPV DNA. The detection limit with the plasmid DNA was 0.065 fg, approximately equal to 10 viral gene copies. Thus, our assay is highly sensitive, and can detect MPV-1 at very low virus copies. In addition, the assay has good reproducibility. Each diluted concentration was repeated three times as PCR template and the amplification efficiency was consistent across the different concentrations and different assay repeats (Figure 1C, five diluted concentrations).
We then put this verified testing system into practical application. Still for an example of MPV-1, we collaborated with a biotechnology company in Wuhan to improve its manufacturing process. This company was carrying out research and development work on a neural growth factor expressed in a mouse cell line. Two batches of samples from two different manufacturing techniques, A and B, were randomly examined, and we tested MPV-1 with viral copies of 6.6×105 and 5.8×105, respectively. Our feedback to the manufacturer resulted in the development of a third manufacturing technique (C). We repeated the assay, and at this time, no MPV-1 was detected in the samples.
Therefore, this assay is proved to be very useful for detection of viral contamination in biological products, and further for maintenance of safety record in biological industry. Compared with real-time PCR using TaqMan probe (Mackay I M, et al., 2002), our assay, based on SYBR Green I chemistry, is cheaper, faster, easier to design and perform, and of higher quality, which make it suitable for widespread use in the pharmaceutical industry.
In conclusion, this successful case of testing for MPV-1 contamination in murine biologicals based on a SYBR Green 1 PCR method indicates that this testing assay, which detects five murine viruses, could be very useful for early monitoring of viral contamination in murine-derived biologicals. Furthermore, it could be expanded to other animal-derived biologics in the pharmaceutical industry.
-
We thank Professor Congyi Zheng of China Center for Type Culture Collection (CCTCC) for help with the five mouse viruses, as well as the Wuhan Key Laboratory on Emerging Infectious Diseases and Biosafety for its support. This work was supported by The Fifth Batch of "China Optics Valley 3551 Talent Program". All the authors declare that they have no competing interests. This article does not contain any studies with human participants or animals performed by any of the authors.













 DownLoad:
DownLoad: