HTML
-
Enteric viruses are the most common cause of acute gastroenteritis (AGE) in infants and young children and a significant public health problem globally (Dennehy 2011). In countries that have not introduced universal rotavirus vaccination, hospital admissions of children under 5 years with diarrhea are primarily associated with group A rotavirus (RVA) infection (Dennehy 2015). In 2013, approximately 215, 000 rotavirus-associated deaths of children under 5 years of age occurred worldwide (Tate et al. 2016), accounting for approximately 4% of all child deaths. Rotavirus infection occurs worldwide, regardless of economic status or standards of hygiene. The World Health Organization (WHO) recommends that rotavirus vaccine is included in all national immunization programs for infants, particularly in countries with high infant mortality rates. Currently, rotavirus vaccines are part of the national immunization program in 95 countries worldwide (http://rotacouncil.org/vaccine-introduction/global-introductionstatus/). At present in the Russian Federation, infant immunization against RVA disease is only provided through private providers or in accordance with epidemic indications.
The efficacy of rotavirus vaccines as assessed by clinical trials is reported to range from 72% to 100% in high-and middle-income countries with low levels of mortality, to 46% to 72% in low-income countries with high child mortality rates (Dennehy 2015). Despite current data indicating that rotavirus vaccines provide cross-protection against the majority of serotypes of circulating RVA strains, including fully heterotypic strains (Leshem et al. 2014; Velasquez et al. 2014; Desselberger 2017; Parker et al. 2018), further research is required to determine the genetic and antigenic diversity of circulating RVA strains in order to facilitate optimization of vaccine composition.
The genotypes of co-circulating RVA strains vary geographically and seasonally (Patton 2012; Afrad et al. 2013a; Arana et al. 2016). The diversity of clinically relevant genetic variants of human rotaviruses is constantly increasing due to the rapid evolution of RVAs, which results from the accumulation of point mutations, gene reassortment, and their wide host range and consequent risk of interspecies virus transmission (Patton 2012; Desselberger 2014). Important factors influencing RVA genotype distribution in a specific area include the intensity of migration processes and possibly the influence of preventive vaccination programs (immunization coverage, vaccine composition).
The Moscow urban area is the most populated region in Russia and one of the most populous region in the world, with more than 16 million residents. According to the city authorities, by the end of 2016, Moscow had been visited by 17.5 million tourists, including 4.5 million people travelling from abroad. The high population density coupled with a high migration rate and tourist flow, necessitates enhanced epidemiological monitoring of infectious diseases, including AGE. Thus, implementation of an effective rotavirus vaccine program must account for the geographic variation of prevalent strains through continuous surveillance and monitoring of circulating genotypes.
The aim of this retrospective study was to evaluate the viral etiology of AGE in children under 5 years old hospitalized with symptoms of AGE in Moscow during 2009–2014. The G/[P] genotype distribution of RVA strains co-circulating in Moscow during the indicated period was also investigated.
-
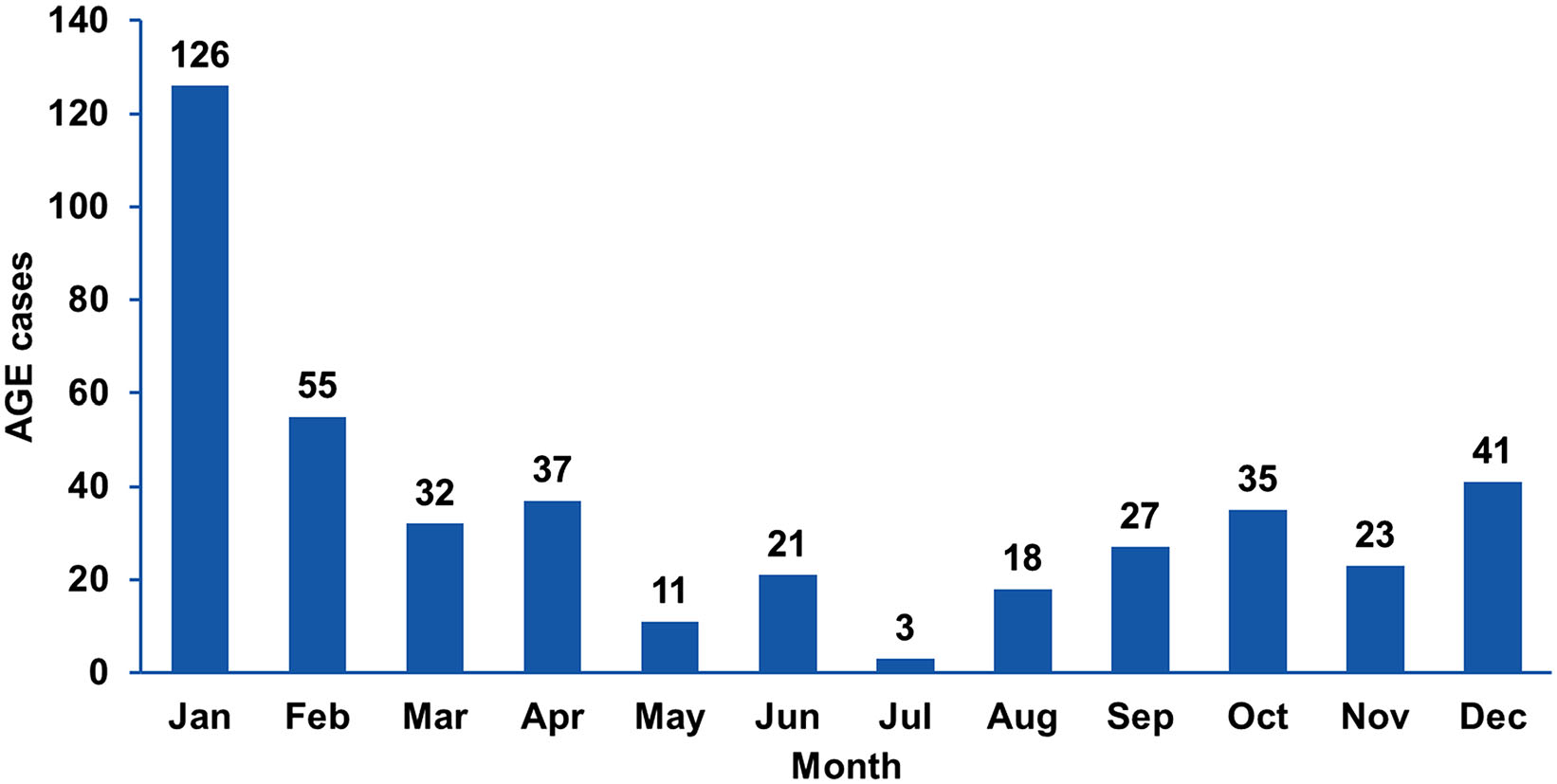
Fecal samples were collected from 469 children under 5 years old (median, 15 months) hospitalized within 72 h after AGE manifestation at the St. Vladimir Children's Municipal Clinical Hospital (Moscow Department of Health) and the Clinical Hospital for Infectious Diseases No. 1 (Moscow Department of Health) between January 2009 and January 2014. Among these 469 samples, 429 were from patients with all-cause AGE, and 40 additional samples were obtained from patients with rotaviral enteritis confirmed by the immunochromatographic test-system "RIDA® Quick Rotavirus/Adenovirus Combi" (Rbiopharm AG, Darmstadt, Germany). Stool samples were collected twice from 165 children: (1) within the first day of hospitalization (primary samples) and (2) on days 6–8 (median, day 7; secondary samples). Samples were collected throughout the year (Fig. 1). A control panel, comprising 42 fecal samples from healthy children under 5 years (median, 12 months), collected in summer 2010 from Mytishchi City Children's Polyclinic No. 2, was also used as a suitable control for studying the etiology of AGE during the summer. All samples were stored at -80 ℃.
-
Nucleic acids were isolated from 10% fecal extracts (100 μL) in physiological saline (0.9% NaCl) using a ZR Viral RNA KitTM (Zymo Research, Irvine, CA, USA) or an RNA/DNA Extraction Kit "MAGNO-sorb" (InterLabService Ltd., Moscow, Russia) according to the manufacturer's instructions. Extracted viral nucleic acids were suspended in 100 μL of nuclease-free water and stored at -80 ℃ until use.
-
All reverse transcription (RT) and PCR reactions were carried out using 2.5× reaction mixture for real-time PCR, supplied by Syntol (Moscow, Russia). Aliquots of extracted RNA (10 μL) were mixed with 3 pmol of primer for RT, incubated at 95 ℃ for 1 min, and cooled for 2–3 min at a room temperature. RT reactions were carried out in a total volume of 25 μL containing 3 pmol of RT primer, 25 units of MMLV reverse transcriptase (Syntol), and 10 units of RNase inhibitor (Syntol). The RT step involved incubation for cDNA synthesis at 45 ℃ for 10 min and MMLV reverse transcriptase inactivation at 95 ℃ for 5 min. Realtime TaqMan-based PCR was carried out in a total volume of 50 μL containing 25 μL of template cDNA, 6 pmol of each primer (forward and reverse), 5 pmol of TaqMan probe, and 2.5 units of SynTaq DNA-Polymerase (Syntol) per reaction. All primers and probes were synthesized by Syntol and their sequences are presented in Supplementary Table S1. Thermal cycling was carried out using a DT-96 Real-Time PCR Cycler (DNA-Technology, Moscow, Russia).
-
PCR for Differential Detection of Enteric Viruses Multiplex reverse transcription real-time PCR was performed as described previously (Marova et al. 2012) with samples obtained from children with AGE and from healthy children to detect nucleic acids from eight groups of human enteric viruses representing the following species: Human mastadenovirus A–G (AdV, adenoviruses), Enterovirus A–D (EnV, enteroviruses), Rotavirus A (RVA, group A rotaviruses), Norwalk virus (NoV, noroviruses), Mamastrovirus 1 (AstV, astroviruses), Sapporo virus (SaV, sapoviruses), Mammalian orthoreovirus (OrV, orthoreoviruses), and Rotavirus C (RVC, group C rotaviruses). A multiplex RT step was carried out using RT-AGE primer mix (a mixture of all primers for reverse transcription, Supplementary Table S1), comprising 3 pmol of each primer per reaction. A multiplex PCR step was carried out using primer mixes AGE-1, AGE-2, and AGE-3 (Supplementary Table S1) in separate tubes containing 6 pmol of each primer per reaction. Human parainfluenza virus type 2 was used as an internal positive control. Cycle parameters were 95 ℃ for 2 min, followed by 45 cycles of 95 ℃ for 20 s and 60 ℃ for 50 s.
-
All RVA RNA-positive samples were genotyped as described previously (Bakhtoiarov et al. 2014) by multiplex reverse transcription real-time PCR in five tubes to differentially detect dominant variants of the following RVA genes: VP7 (G1, G2, G3, G4, G9), VP4 ([P4], [P6], [P8]), and VP6 (I1, I2). A multiplex RT reaction was carried out using a mixture of all primers for reverse transcription RT-Gen (Supplementary Table S1). Multiplex real-time PCR assays were carried out using primer mixes Gen1, Gen2, Gen3, Gen4, and Gen5 (Supplementary Table S1). PCR cycle parameters were 95 ℃ for 2 min, followed by 45 cycles of 95 ℃ for 20 s and 55 ℃ for 50 s.
The multiplex reverse transcription PCR and gel electrophoresis method described in "Manual of rotavirus detection and characterization methods. WHO" (October 2009, Method 16) was used as a reference RVA genotyping method.
-
For PCR-amplification of the VP7 and VP4 cDNA fragments, primers VP7F, VP7R (Iturriza-Gomara et al. 2001) (positions 49–933 in GenBank accession, K02033) and VP4F, VP4R (Simmonds et al. 2008) (positions 142–805 in GenBank accession, KT694942) were used. Cycle parameters were 95 ℃ for 2 min, followed by 45 cycles of 95 ℃ for 1 min, 52 ℃ for 40 s, and 72 ℃ for 40 s, and a final 72 ℃ for 10 min. PCR products were purified using a "Cleanup Standard" kit (Evrogen, Moscow, Russia) and sequenced in both directions with the primers used for PCR. Sanger sequencing was performed by Syntol. Phylogenetic trees were constructed using MEGA6 (Tamura et al. 2013), based on the partial nucleotide sequences of the RVA genes (VP7 and VP4) sequenced in this study, and reference RVA strains with established genotypes, as recommended by the Rotavirus Classification Working Group (Matthijnssens et al. 2008).
-
Data were analyzed by χ2 test, or Fisher's exact test (twotailed, 95% confidence intervals) for statistical comparisons of prevalence rates. Since the data was not normality distributed, a non-parametric Mann–Whitney U test (M-W) was selected. The coefficient of determination (R2) was calculated to evaluate the degree of correlation between two data series.
Origin and Collection of Samples
Viral RNA Extraction
Reverse Transcription and Real-Time PCR
Multiplex Reverse Transcription and Real-Time
Multiplex Real-Time PCR for Genotyping of RVA
Sequencing of VP7 and VP4 Gene Fragments and Generation of Phylogenetic Trees
Statistical Methods
-
Among the 429 samples collected from children with allcause AGE on the day of hospitalization, 287 samples (67%) were positive for one or more enteric viruses, whereas 142 samples (33%) were negative for viral nucleic acids (Table 1). RVA was the most frequently detected pathogen, with a prevalence of 40.1% (172 cases), followed by norovirus and adenovirus infections, 11.4% (49 cases) and 4.7% (20 cases), respectively. Astroviruses, sapoviruses, enteroviruses, and orthoreoviruses were detected in 1.9%, 1.4%, 1.2%, and 0.2% of cases, respectively. Among the 287 positive samples, 261 were positive for one virus, while 26 were co-infected with two or more different enteric viruses, with a predominance of RVA, including co-infections with RVA and AdV or RVA and NoV, which were detected in 7 cases (1.6%), and RVA and AstV, which was identified in 4 cases (0.9%). In addition, single cases of co-infection with AdV/AstV, RVA/RVC, RVA/NoV/EnV/AstV, SaV/AstV, EnV/NoV, and RVA/EnV/NoV were detected.

Table 1. Distribution of enteric viruses detected in samples from children with AGE, according to their age (n = 429)
Next, the distribution of enteric viruses implicated in AGE etiology in different age groups (0–6, 7–12, 13–24, and 24–60 months) was estimated (Table 1). The lower levels of viral etiology in the age group 0–6 months is likely due to the presence of transplacentally transmitted maternal specific antibodies.
Furthermore, a substantial increase in RVA frequency (up to 66.7%) in children with AGE was observed during March and April, compared with other months (27.4%–42.9%) (Fig. 2).
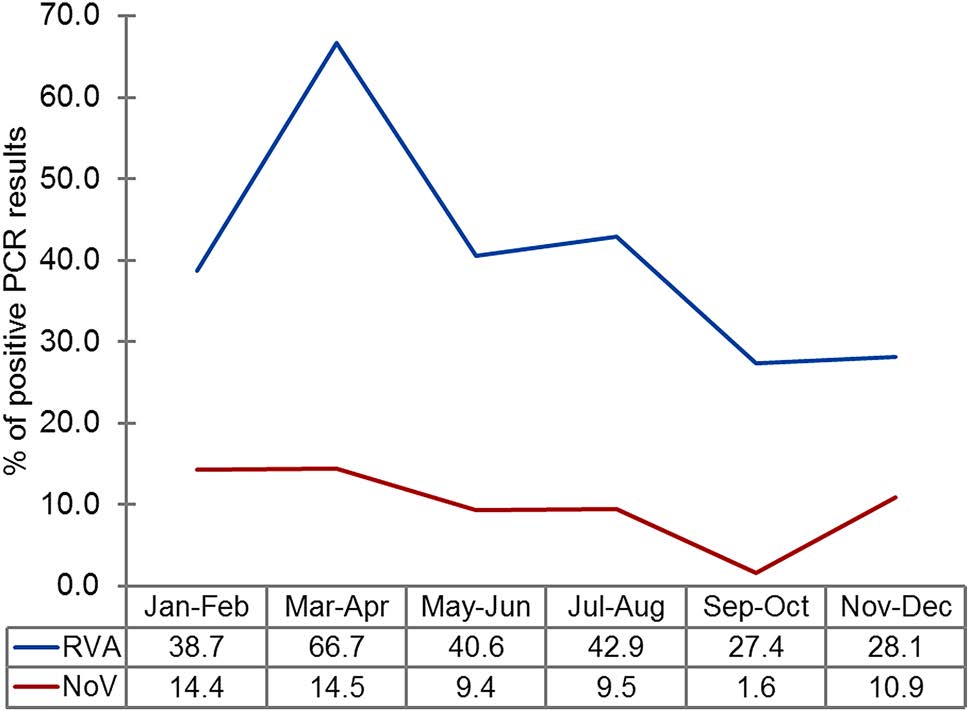
Figure 2. Distribution of rotavirus A and noroviruses detected in clinical samples according to month. The graph shows the change in the proportion of individuals positive for rotaviruses (RVA; blue line) and noroviruses (NoV; red line) during the year, determined by the percentage of samples with positive PCR results (n = 429).
-
No viral nucleic acids were detected in 38 of 42 specimens (90.5%) collected from healthy children (without AGE manifestation) during the summer of 2010. In four samples, adenoviral DNA (two cases with Ct values of 25.0 and 30.0) and sapoviral RNA (two cases with Ct values of 23.3 and 27.3) were detected. In contrast, among samples collected from children with AGE in summer 2010 (n = 42), enteric viruses were identified in 67.7% of cases with a predominance of RVAs (35.7%), followed by noroviruses (9.5%), adenoviruses, enteroviruses, and mixed infection (each 7.1%). Notably, a high proportion of RVA infection was detected in hospitalized children with AGE registered in summer 2010 (35.7%), which is not typical for the summer period in regions with a temperate continental climate.
-
In 42 of 165 secondary stool specimens collected twice, nucleic acid of a virus other than that detected in primary samples was identified, indicating that these were nosocomial infections (25.5%). Nosocomial infections were identified with rotaviruses, noroviruses, astroviruses, and adenovirus in 22 cases (52.4%), 12 cases (28.6%), 3 cases (7.1%), and 1 case (2.4%), respectively. In four cases (9.5%), nosocomial infection with two or three viruses was detected. Furthermore, a high linear correlation (R2 = 0.94) was determined between the viral population structure of nosocomial infections and that in children with AGE hospital admissions.
Nosocomial infections were significantly more prevalent in younger infants: median age, 5 (CI 3–9) months versus 13 (CI 6–23) months for children without nosocomial infections (P = 0.0007, M–W). Seventy-eight per cent of patients with nosocomial enteric viral infections (32 of 41) had clinical manifestations, such as reduced general condition, vomiting, and diarrhea. In 31 cases, severe clinical manifestations were associated with RVA (20 cases), NoV (8 cases), or both viruses (3 cases). AstV, AdV, RVC, and ReV were detected more frequently (4 among 9 cases) in children without clinical manifestations of nosocomial infection. Furthermore, among children with clinical manifestations of nosocomial viral infections, the viral load, determined by real-time PCR (according cycle threshold (Ct) values), was higher than in children without clinical manifestations of nosocomial infection [Ct = 19.5 (CI 16.1-22.5) vs. 26.9 (CI 22.0-30.6), P = 0.010, M–W]. Thus, nosocomial infection with enteric viruses was observed in 24.85% of hospitalized children, while accompanying clinical manifestations were present in only 19.4% of cases. The presence of clinical manifestations correlated with younger age and, probably, higher levels of virus replication activity.
-
RVA positive samples (n = 212) were analyzed by multiplex type-specific real-time PCR for G/[P]-genotyping. For the 170 RVA strains detected in clinical specimens, the G/[P]-genotype, or a variant of the VP7 or VP4 genes, was determined (Fig. 3). The predominant genotype was G4P[8], detected in 38.7% of cases, whereas genotypes G1P[8], G9P[8], G3P[8], G2P[4] varied in frequency from 11.8% to 3.3%. In almost 20% of cases, the RVA genotype was not determined, predominantly due to the low viral load in collected samples, and probably also because of insufficient sensitivity of the genotyping system. The 2012–2013 period was characterized by an unusually high proportion of the G9P[8] genotype (30%).
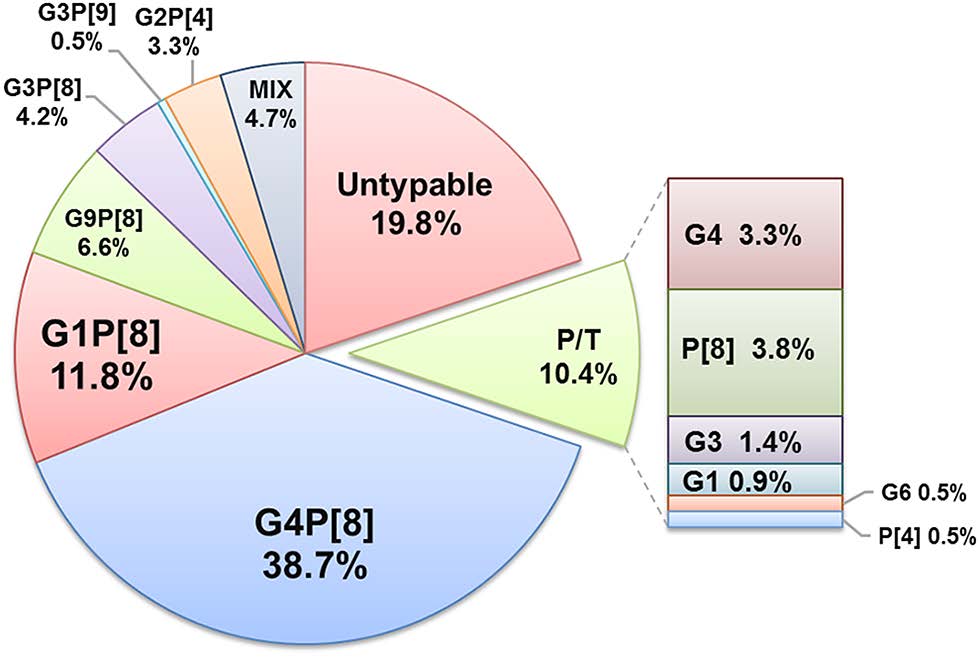
Figure 3. Distribution of RVA G/[P]-genotypes detected in clinical samples between 2009 and 2014 in Moscow. Each segment represents the relative distribution of an RVA G/[P] genotype based on data from type-specific multiplex real-time PCR analysis of fecal extracts collected from children with rotaviral enteritis (n = 212). The total figures for the whole period are presented. Mix mixed infection, P/T partially typed.
The reliability of the data obtained by type-specific PCR was confirmed by selective Sanger sequencing of the VP4 and VP7 genes fragments. Phylogenetic analysis was performed based on partial sequences of the VP4 (segment 4) and VP7 (segment 9) genes derived from 664 and 885 bp PCR amplicons, respectively. In total, 44 gene fragment sequences (11 VP4 and 33 VP7, GenBank accession numbers: KT000090–KT000133) were sequenced. To determine the taxonomy of the RVA strains, phylogenetic trees were generated using the Maximum Likelihood Method and the Kimura two-parameter evolution model, based on the partial VP4 and VP7 RVA gene nucleotide sequences (Fig. 4). The bootstrap values of tree nodes uniting particular variants of the genes ranged from 88% to 100%, indicating strong support for these nodes. Overall, in 100% of cases the studied strains could be grouped according to the genotype of the reference strains and VP4 and VP7 genes variants, previously determined by typespecific real-time RT-PCR, indicating the high accuracy of RVA genotyping by PCR and the reliability of the data obtained. Among RVA strains with genotype undetermined by type-specific real-time RT-PCR, two strains with rare gene variants were identified by sequencing: G6 and P[9] (GenBank accessions KT000122 and KT000133).
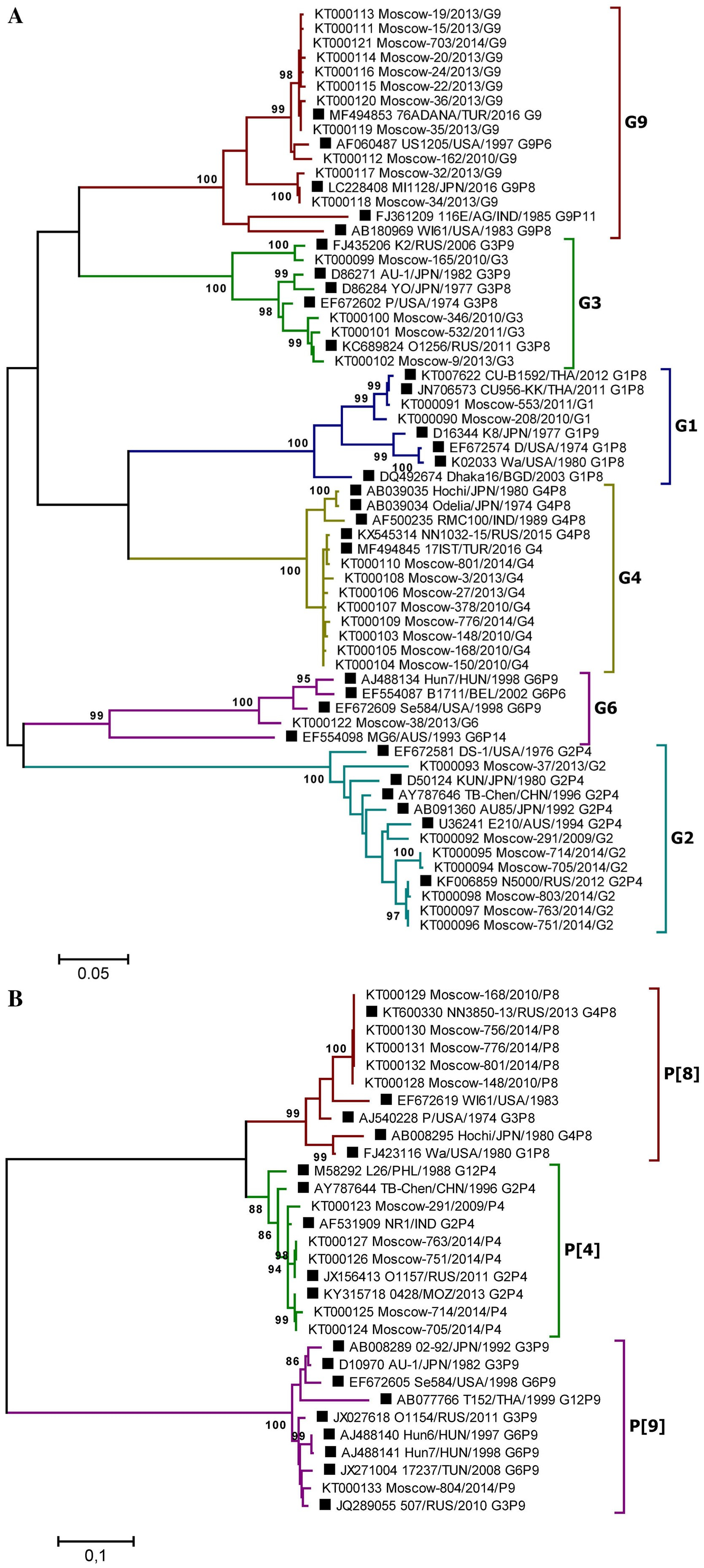
Figure 4. A Phylogenetic tree based on partial nucleotide sequences of the RVA VP7 (A) and VP4 (B) gene from sequenced clinical samples and reference strains. Phylogenetic trees were generated using MEGA6 (Tamura et al. 2013) with the maximum likelihood method and the Kimura two-parameter model, based on the partial nucleotide sequences of the RVA VP7 and VP4 genes. The names of clinical samples include the GenBank accession number, the sample isolation location (Moscow), a laboratory identification number, the year of isolation, and the gene variant determined by PCR. Reference strain names are indicated by GenBank accession number, strain (isolate) name, country and year of isolation, and genotype. Three-letter country codes were used according to ISO 3166-1. Numbers at the nodes are bootstrap values based on 1000 replications. Reference strains are labeled as filled square. Bootstrap cutoff values are 95% (A) and 85% (B).
It should be noted that the lengths of the sequenced PCR products determined in this study were less than 50% of the VP4 gene open reading frame sequence while, according to the recommendations for the RVA classification by Rotavirus Classification Working Group, to define the genotype of an RVA strain, at least 50% of the open reading frame sequence should be determined. Therefore, an additional reference method for G/P-genotyping of all sequenced specimens by multiplex RT-PCR and agarose gel electrophoresis was used, as recommended by the WHO "Manual of rotavirus detection and characterization methods". This analysis revealed a 100% agreement of the results obtained by the two methods.
The Prevalence of Enteric Viruses in AGE Etiology
The Frequency of Asymptomatic Carriage of Enteric Viruses During Summer
Nosocomial Enteric Viral Infections
Distribution of RVA Genotypes G and P
-
According to the WHO, the proportion of children with rotaviral enteritis among the total number of children hospitalized with AGE varied from 7% to 66% in various countries in 2012, while the global median was 38% (Global Rotavirus Information and Surveillance Bulletin. WHO. Volume 7. February 2013). In recent years, the viral etiology of severe AGE in children has changed in countries with a high rotavirus vaccine coverage (Dennehy 2011; Tam et al. 2012; Leshem et al. 2014, 2015; O'Ryan et al. 2015). Thus, in the USA by 2012, the proportion of children under 5 years with rotaviral enteritis among the total number of hospitalized patients with AGE decreased from 21.0% to 2.9%, compared with the pre-vaccination period (2002–2006) (Leshem et al. 2015). In 2012, when vaccine coverage was at its highest, the greatest reductions were observed for all-cause acute gastroenteritis (55%) and RVA-coded (94%) hospitalizations. Simultaneously, noroviruses have become the most prevalent cause of AGE in countries with high rotavirus vaccine coverage (Tam et al. 2012; Hemming et al. 2013; Payne et al. 2013; Bucardo et al. 2014).
The results obtained in our study confirm the leading role of rotaviruses (40.1%) in the etiology of AGE cases requiring hospitalization in Moscow during 2009–2014 (excluding 6.1% of cases with mixed infection), which is consistent with data for pre-vaccination periods in other countries. Other common pathogens leading to the hospitalizations for AGE included noroviruses and adenoviruses (11.4% and 4.7%, respectively), which corresponds with current understanding of the important role of these viruses in the etiology of AGE.
In our work, a high level of cases with laboratoryconfirmed nosocomial infection with enteric viruses was revealed (25.5% of all children hospitalized with AGE). In the population with nosocomial infections, RVA (52.4%) and norovirus (28.6%) infections were predominant, corresponding to previously published data (Gleizes et al. 2006; Bruijning-Verhagen et al. 2012; Kambhampati et al. 2015). The high numbers of children admitted to hospital with rotavirus and norovirus infections determined the crucial role of these viruses in the etiology of nosocomial infections. Indeed, there was a strong relationship between the parameters of AGE etiology and those of nosocomial infection, with a linear correlation coefficient of 0.94.
An unusually high ratio of RVA infection in the total viral population detected among children with AGE (35.7%) was observed in children hospitalized during the summer of 2010, which was not typical for the summer months in areas with a temperate continental climate. This is likely attributable to the abnormally high temperatures (35 ℃–40 ℃) recorded in the European part of Russia during July and August, 2010, which was accompanied by air pollution (Revich et al. 2015) and people spending more time near open reservoirs and in public bathing places, and using ice from the freezer more frequently, thereby promoting the spread of rotaviruses. Notably, in some countries of Central and Eastern Africa, characterized by warm climates, the increase in the incidence of rotaviral enteritis in children occurs during dry (Mayindou et al. 2016) or dry and warm seasons (Omore et al. 2016).
Currently, Russia is on the verge of introducing universal RVA vaccination. From this perspective, the experience of other countries should be taken into account, where, after the introduction of vaccination and a reduction in morbidity, a significant increase in the incidence rate occurred again, accompanied by changes in infection severity in other age groups, likely due to variations in circulating genotypes (Leshem et al. 2014; Abdel-Haq et al. 2016; Shim et al. 2016). Changes in circulating RVA genotypes against the background of universal RVA vaccination have been documented in both developing and developed countries (Gurgel et al. 2009; Afrad et al. 2013b; Pitzer et al. 2015); however, there is no convincing evidence that these changes depend on RVA vaccination or the composition of the vaccines used.
Discussion and comparison of the distribution of RVA genotypes is relevant in the context of the composition of existing RVA vaccines. Rotarix (GlaxoSmithKline) and RotaTeq (Merck) vaccines are used worldwide. The Rotarix vaccine consists of an attenuated G1P[8] human RVA strain, whereas RotaTeq contains 5 bovine-human mono-reassortant RVA strains, comprising either human RVA VP7 (G1, G2, G3 and G4) or VP4 (P[8]) gene segments, introduced into a bovine RVA strain G6P[5] to provide the genetic background. Currently, RVA strains that are homotypic, or partially heterotypic, in relation to vaccines, including G1P[8], G2P[4], G3P[8], G4P[8], and G9P[8] (Patton 2012), prevail among strains circulating worldwide; Moscow is no exception, as demonstrated in this study. The Moscow urban area is a place of intersection, where migration, business, and tourist flows occur, and this would be expected to manifest in significant similarity in RVA genotype distribution relative to that in other regions of Europe. Indeed, among the viral genotype structures present in the cities of the both European and Asian parts of Russia (Moscow, Nizhny Novgorod, St. Petersburg, Chelyabinsk, Tyumen, Makhachkala, Khabarovsk, Novosibirsk, Omsk) and in a number of bordering countries (Ukraine, Moldova), the genotypes G4P[8] and G1P[8] were dominant during 2005–2014 (Podkolzin et al. 2009; Zhirakovskaia et al. 2012; Lobzin et al. 2017; Sashina et al. 2017). In addition, the most noticeable change in the RVA genotype structure noted in the period 2012–2016 in Nizhny Novgorod (the increase in the proportion of the G9P[8] genotype up to 25.9%–45.9%), was also reflected in our results. Hence, the 2012–2013 period in Moscow also was characterized by a high proportion of RVA strains with the G9P[8] genotype (30%), which are closely related to Turkish and Nizhny Novgorod strains (GenBank accession numbers MF494853 and KC677698, respectively). However, in a number of world regions, some strains (e.g., G12P[6], G9P[4], G8P[6], G9P[6] etc.) that are fully or partially heterotypic with respect to vaccines are medically important (Afrad et al. 2013b; Doro et al. 2014). Despite the available data on the cross-protective activity of rotavirus vaccines, including those against fully heterotypic strains (Leshem et al. 2014; Velasquez et al. 2014), it is critical to study the antigenic variety among circulating RVA strains. It cannot be completely ruled out that vaccination can selectively influence viral populations, leading to fluctuations in strains distribution and their immune escape. In this context, the circulation of strains Moscow-38/2013/G6 and Moscow-804/2014/P[9] in the Moscow region is particularly interesting. The closest nucleotide sequences to these strains belong to strains with the established G6P[9] genotype isolated in the following countries: USA (1998, GenBank accession number, EF672609), Hungary (1998, AJ488134), Hungary (1997, AJ488140), Hungary (1998, AJ488141), Tunisia (2008, JX271004); and G3P[9] isolated from Russia in 2010 (JQ289055) and 2011 (JX027618). These strains are partially or completely heterotypic with respect to vaccines (Fig. 4).
Thus, the main cause of hospitalization with AGE and nosocomial gastrointestinal infections of children aged less than 5 years in the two Moscow hospitals was RVA followed by noroviruses. The predominant RVA genotype between 2009 and 2014 in Moscow was G4P[8], which was detected in 38.7% cases; genotypes G1P[8], G9P[8], G3P[8], G2P[4] were also detected. Simultaneously, the proportion of untypable rotavirus strains was high (20%) which determines the need to increase the sensitivity of rotaviruses genotyping methods. The presence of circulating RVA strains with rare genotypes, which are partially or completely heterotypic with respect to rotavirus vaccines, indicates the need for continuous surveillance and monitoring of circulating RVA genotypes.
-
The study of the distribution of RVA G/[P]-genotypes was at the expense of the Russian Science Foundation (Grant No. 16-15-10332).
-
EF and EM designed the experiments. VK, AM, AO, TS, GB, NB and AL carried out the experiments. EF and EM analyzed the data. EF, EM, VK and YA wrote the paper. NF and VZ checked and finalized the manuscript. All authors read and approved the final manuscript.
-
The authors declared that they have no conflict of interests.
-
The study protocol was approved by the Medical Ethics Review Committees of the hospitals in Russia. Written consent was provided by the parents.
Conflict of interest
Animal and Human Rights Statement
-

Table Supplementary Table S1. Primers and probes for multiplex real-time RT-PCR-detection of AdV, EnV, RVA, NoV, AstV, SaV, ReV, RVC, genotyping of rotaviruses and sequencing of VP7 and VP4 genes fragments.
References
Bakhtoiarov GN, Kiselev IS, Zverev VV, Faizuloev EB. Evaluation of real-time multiplex PCR effectiveness for group A rotavirus genotyping. Zh Mikrobiol Epidemiol Immunobiol. 2014:43-49. (In Russian)
Iturriza-Gomara M, Cubitt D, Desselberger U, Gray J. Amino acid substitution within the VP7 protein of G2 rotavirus strains associated with failure to serotype. Journal of Clinical Microbiology. 2001;39:3796-3798.
World Health Organization (WHO). Manual of rotavirus detection and characterization methods, October 2009.
Marova AA, Oksanich AS, Kaira AN, et al. Experience of application of multiplex qPCR for differential diagnostics of intestinal viral infections. Zh Mikrobiol Epidemiol Immunobiol. 2012:39-45. (In Russian)
Simmonds MK, Armah G, Asmah R, et al. New oligonucleotide primers for P-typing of rotavirus strains: Strategies for typing previously untypeable strains. J Clin Virol. 2008;42:368-373.















 DownLoad:
DownLoad: