-
Dear Editor,
Dengue infections are caused by all four serotypes of dengue virus (DENV1–4). In recent years it has been confirmed that DENV-1 and DENV-2 are co-circulated in Guangzhou, Guangdong, China (Lai et al. 2015; Luo et al. 2017). DENV-2 strains were imported from Thailand and Indonesia (Luo et al. 2017; Zhao et al. 2014, 2016), and had become the second local circulated serotype in Guangzhou.
Rapid and accurate diagnosis of DENV is crucial for prompt clinical management during an outbreak. The combined NS1 and IgM/IgG tests using immunochromatographic assays (ICs) are recommended for routine diagnosis of dengue fever. But these assays are inherently inferior to ELISA in sensitivity, and thus their usefulness during acute phase of DENV infection is uncertain.
In August 2018, one female (pt3) who had fever and decreased white blood cell count (1.41 × 109/L) was admitted to our hospital. Skin rash was found on her forearms, but thrombocytopenia was not found (platelet count, 117 × 109/L). We did the NS1 and IgM/IgG rapid testing for DENV (see Supplementary Materials and Methods for details), and found negative for both tests from samples on day 4 and day 5 (Supplementary Table S1). But the DENV RNA was shown to be positive on day 5 and serotype 2 was determined by RT-PCR for the same patient (Supplementary Materials and Methods and Supplementary Table S1). To understand whether the lack of sensitivity of these commercial kits contributed to the false negative results, laboratory data from a total of 15 cases with RT-PCR confirmed DENV2 infection was analyzed (Supplementary Table S1). 10 out of 11 individuals with NS1 rapid testing showed weak or strong positive in consistent with the results of RT-PCR. Most cases (8/10) were negative for IgM/IgG by rapid testing before day 6.
We further analyzed the NS1 and IgM/IgG in sera collected from seven patients using ELISA method, including pt3, on days 5, 6, 32 and 356 (Fig. 1). The false negative of pt3 in NS1 rapid test was associated with a very low NS1 antigenemia (OD = 0.2 or 0.3 on day 5 or day 6) (Fig. 1A). Five patients showed IgM positive during the acute phase (Fig. 1B) but only pt6 was weak positive in rapid test, (Supplementary Table S1) indicating the limitation of IgM rapid test. For IgG testing by ELISA (Fig. 1C), pt2, pt4 and pt5 were positive, however, they were negative by the previous rapid test. Notably, pt3 was still IgM negative on day 6, and only became positive on day 32. Surprisingly, IgG cannot be detected on days 5, 6, 32 and even 356, suggesting a delayed IgM response and a defective IgG response (Fig. 1C). According to the standard diagnostic criterion, the pt3 may be classified as primary infection (Nguyen et al. 2018; Prince et al. 2011; Zhao et al. 2016). But low NS1 antigenemia usually occurs in secondary DENV infection (Duyen et al. 2011), thus this patient is an exception. In another study, no difference was observed in NS1 concentration among the serum specimens from patients with primary and secondary DENV1 infections by NS1 capture ELISA (Sophie et al. 2002). The very low level of NS1 in this patient's serum during the acute phase could explain why the test was negative due to the low sensitivity of rapid NS1 test.
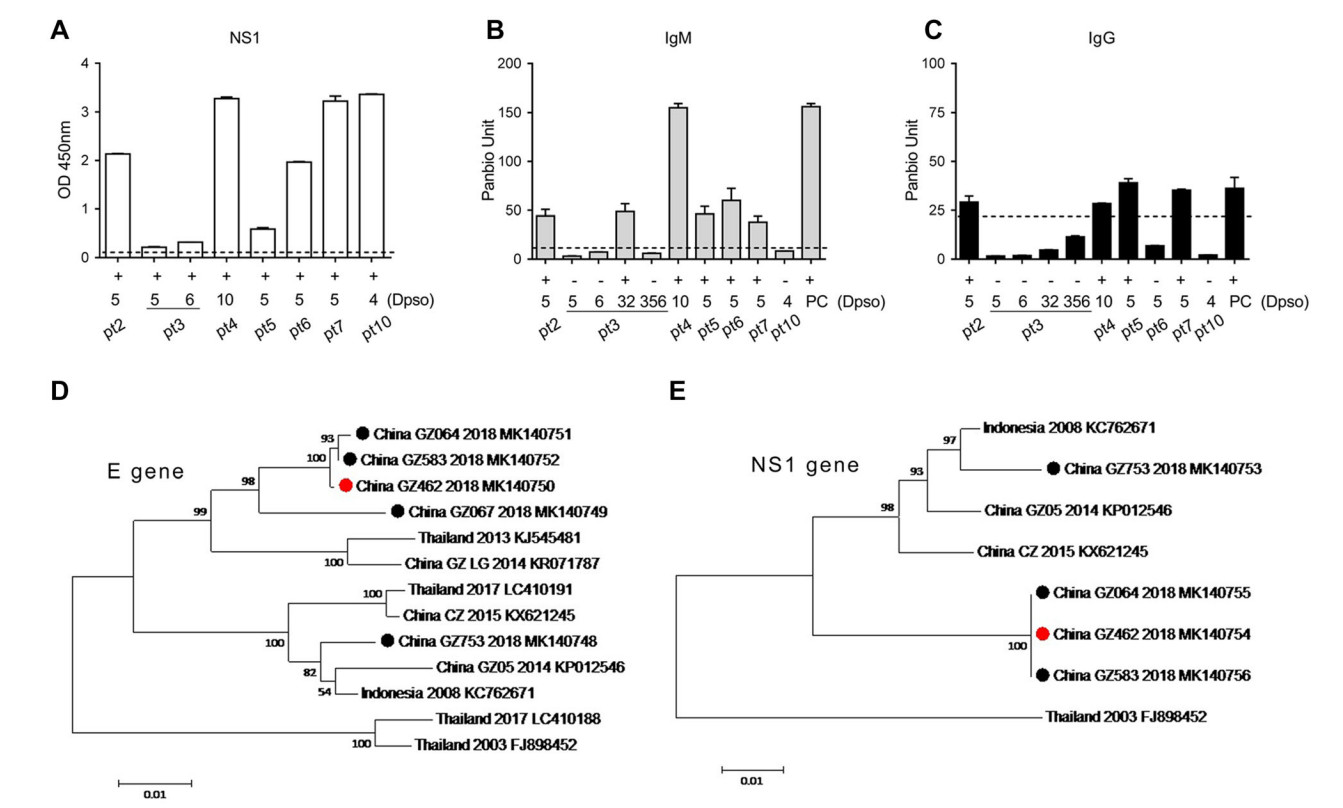
Figure 1. A–C Detections of NS1, IgM and IgG of DENV in seven patients by the commercial ELISA kits. A Serum NS1 from seven DENV2-infected patients using ELISA Kit for DENV NS1 antigen detection (WANTAI, China). B, C IgM or IgG test in serum using PanBio dengue IgM and IgG capture ELISA. Each sample was tested in duplicates. The dotted lines indicate the cutoff value for OD or Panbio unit. PC, positive control. Dpso, Days post the symptom onset.
To find out if the variation of virus strain resulted in the patient's deficient antibody response or low NS1 antigenemia, we sequenced the coding gene of the envelope (E) protein (see Supplementary Materials and Methods for details). Among the seven collected serum samples, five fragments of E glycoprotein were successfully amplified by RT-PCR from pt2, pt3, pt6, pt7 and pt10, and sequenced. The phylogenetic analysis revealed that there were two clusters of DENV2 strains co-circulating in which one (GZ064, GZ583, GZ462 and GZ067 isolated from pt6, pt2, pt3 and pt7, respectively) was closely related to the Thailand strain (KJ545481, 2013), and the other (GZ 753 from pt10) was closely related to the Indonesia strain (KC762671, 2008, Fig. 1D). By alignment of E sequence of four isolates in cluster one with the Thailand strain (KJ545481, 2013), two or three mutations were identified in 2018 isolates (Table 1). For example, the China_GZ064 strain had two substitutions: C121Y and V358A. It was noted that the sequences of GZ462 from pt3 and GZ583 from pt2 were completely identical, despite pt3 had lower NS1 OD value in serum compared to that of pt2 (OD: 0.2 versus 2.0, Supplementary Table S1 and Fig. 1A).
Isolates/patients Country Year Accession No. Amino acid position 45 121 141 162 170 182 322 358 Thailand Thailand 2013 KJ545481 L C V I I T V V GZ-LG China 2014 KR071787 - - - - - A - - GZ064/pt6 China 2018 MK140751 - Y I V - - I A GZ067/pt7 China 2018 MK140749 P - I - V - I - GZ462/pt3 China 2018 MK140750 - - I V - - I - GZ583/pt2 China 2018 MK140752 - - I V - - I - Table 1. Amino acid in E proteins of DENV2 isolates in cluster one circulating in Guangzhou, 2018.
To study whether the NS1 gene mutation might have affected the secretion of NS1, we also did sequence analysis of full-length of NS1 (Fig. 1E and Supplementary Materials and Methods). Three sequences obtained from one DENV2 cluster (pt6, pt2 and pt3) were identical and pt3 showed no mutation in NS1 gene. A previous study showed that the mutation E350D in Chikungunya virus 6KE1 protein reduced the recognition by the mAb and affected the performance of a rapid E1-antigen test (Tuekprakhon et al. 2018). It was also reported that the serum NS1 level varied between infections by different DENV2 strains in a mouse model (Watanabe et al. 2012). In our study we did not find any mutation in either E or NS1 genes, and thus diminished NS1 secretion was a result of viral mutation. Our study also showed that at least two DENV2 strains co-circulated in the summer in Guangzhou, Guangdong Province of China in 2018. The sequences of NS1 gene were conserved among the clinical isolates in the same cluster, whereas E gene had some mutations.
As for the first line of screening test for DENV infection, the sensitivity of rapid NS1 test could vary according to the fluctuation of NS1 antigenemia levels (Bosch et al. 2017; Pang et al. 2017; Shan et al. 2015). Our study displayed that very low level of NS1 secretion in serum from DENV-infected patient with special immune status resulted the false negatives in two successive rapid tests. These combined the negative results of antibodies detection obstruct the diagnosis of DENV infection for doctors in clinic. In summary, we observed the increased number of DENV2 infection in clinic and found one case with unexpected delayed IgM response and defective IgG response, accompanied with low level of NS1 protein. This unusual antibody response to DENV2 and NS1 antigenemia limited the sensitivity of NS1 and IgM/IgG rapid tests for DENV diagnosis. For the clinically suspected cases, it would be a good practice to perform multiple diagnostic tests to avoid misdiagnosis.
-
We thank Dr. Yongjun Guan for the consulting of the capture ELISA and the isolation of antibody. This Project was supported by funds from 2017 Guangzhou major projects of collaborative innovation in health care (201704020229), the State Key Laboratory of Pathogen and Biosecurity (SKLPBS 1839).
HTML
Acknowledgements
-
The authors declare that they have no competing interests.
-
The study was approved by the Ethical Committee of the Guangzhou Eighth People's Hospital (20131224). The samples were collected and used in strict accordance with relevant guidelines and regulations.
















 DownLoad:
DownLoad: