HTML
-
Laos, officially the Lao People's Democratic Republic, is the only landlocked country in Southeast Asia, which borders Myanmar, Thailand, Cambodia, Vietnam and China (Fig. 1). Laos is a low-income, multi-ethnic country, with a population of estimated 7.23 million in 2018.

Figure 1. The geographic location of Laos in Southeast Asia. The real line with black shadow indicates the border between different countries. The blue shadow indicates the location of the "Golden Triangle". The red shadow indicates the location of the sample site, Vientiane Capital, Laos. The two-letter abbreviations indicate different provinces of Laos. The numbers under country names indicate the HIV prevalence reported by the Joint United Nations Programme on HIV/AIDS, and the numbers under the two-letter abbreviations indicate the prevalence of HIV among patients with different domiciles of origin determined by the present study.
Laos has been regarded as having low human immunodeficiency virus (HIV) prevalence for decades, with the first case identified in 1990 (Toole et al. 2006). According to the report of the Joint United Nations Programme on HIV/AIDS (UNAIDS) in 2017, the HIV prevalence in Laos was 0.3%, the number of people living with HIV was 12, 000, and the AIDS-related deaths were less than 500 (UNAIDS 2017). The specific HIV/AIDS project regarding antiretroviral therapy (ART) in Laos was started in 2001. Up to the end of 2017, the proportions of people living with HIV receiving ART and who have suppressed viral loads were 48% and 45% respectively (UNAIDS 2017). However, its southwest neighbor countries, Thailand, Myanmar and Cambodia, suffered a more serious impact of HIV. The HIV prevalence among these countries were 1.1%, 0.7% and 0.5%, respectively (UNAIDS 2017).
A series of studies regarding HIV had been conducted in Laos. In 1996, a survey performed among 502 pregnant women in Vientiane, the capital of Laos, revealed that the HIV positive rate was 0.4% (Loue et al. 1998). In 2007, the HIV prevalence among 540 men who had sex with men was 5.6% (Sheridan et al. 2009). Besides, most studies regarding HIV in Laos focused on the behavior, vulnerability, and co-infection associated with HIV (Thammavong et al. 2011; Slesak et al. 2012; Hansana et al. 2013; Sychareun et al. 2013; Thanavanh et al. 2013; Paboriboune et al. 2014; Sichanh et al. 2014; Barennes et al. 2015; Vorasane et al. 2017). The HIV epidemic states of Laos in recent years has remained unclear.
With the efforts of local governments and international organizations, the HIV molecular epidemiology of most countries in Southeast Asia has been thoroughly studied (Lu et al. 2008; Liao et al. 2009; Pang et al. 2012; Chen et al. 2014). The predominant HIV strains circulating in the China-Myanmar border region are recombinant forms (RF) of subtypes B, C and CRF01_AE (RF_01BC). In Thailand, these strains are CRF01_AE, B and RF_01B. In Cambodia and Vietnam, the main strains are CRF01_AE. By 27 July 2019, 40, 925 sequences from China had been submitted to the Los Alamos National Laboratory HIV sequence database (HIVDB), and the number was 17, 469 in Thailand, 6898 in Vietnam, 2305 in Cambodia and 1051 in Myanmar (HIVDB 2019). However, this number in Laos was zero, although Laos is surrounded by the countries mentioned above (HIVDB 2019). Therefore, Laos has represented the missing link in studies of HIV molecular epidemiology in Southeast Asia.
To understand the recent molecular epidemiological characteristics of HIV in Laos and to provide some references for the prevention of HIV in Southeast Asia, we performed a retrospective study among 2851 patients visiting a hospital in Vientiane, the capital of Laos, from November 2011 to May 2012. Phylogenetic and phylogeographic analyses based on HIV near full-length genomes were conducted.
-
This study was approved by the Ethics Committee of Kunming Institute of Zoology, Chinese Academy of Sciences (approval number: SWYX-2009021; approval date: January 7, 2009). Written informed consent was obtained from all participants.
-
From November 2011 to May 2012, 2851 patients were recruited from a hospital in Vientiane, the capital of Laos. Basic sociodemographic information was collected, including gender, ethnicity and domiciles of origin. Approximately five milliliters of peripheral blood samples were collected with EDTA2K plus vacuum tubes. The blood samples were mixed with DNA-EZ Reagents G DNA-Be-Locked B for Blood Samples (Sangon Biotech Co., Ltd., Shanghai, B64 2021) and stored in a -80 ℃ freezer until HIV testing and genome amplification experiments were performed.
-
Genomic DNA was extracted from the blood samples using a Blood Genomic DNA Mini Kit (CW Bioteck Co., Ltd., Beijing, CW 2087M). The HIV gag fragment was amplified with nested PCR using the TransTaq DNA Polymerase High Fidelity Kit (Beijing TransGen Biotech Co., Ltd., Beijing, AP131-13). Nested PCR were performed using procedures previously described (Chen et al. 2018). The PCR products were tested with 1% agarose gel, and the possible positive samples were sent for sequencing in Sangon Biotech (Shanghai) Co., Ltd. Amplified gag sequences were blasted in Genbank to ensure whether they were HIV sequences.
-
For the HIV-positive samples, HIV near full-length genome was amplified from the provirus. Three fragments that had overlap were amplified independently with nested PCR. The primers used in the nested PCR are summarized in Supplementary Table S1. The annealing temperature used in the nested PCR was 50 ℃, and the extension time was 4 min. The other conditions and operations were performed in the same manner as with the amplification of the HIV gag fragment.
These three fragments were assembled using seqman in the DNAstar package. The near full-length sequences were hand-curated to ensure the correct open reading frame according to the sequence chromatogram.
-
Primary subtype information of the amplified sequences was acquired using the Recombinant Identification Program that is available in HIVDB. Furthermore, HIV subtype reference sequences relevant to Southeast Asia were downloaded from HIVDB and aligned with the amplified sequences. The maximum likelihood (ML) tree was constructed using MEGA X with 1000 bootstrap replications. For the sequences that did not cluster with subtype reference sequences, bootscan analysis was performed using Simplot 3.5.1 to identify potential HIV recombinants, as previously described (Chen et al. 2017a). For the sequences that shared the same recombinant breakpoints, ML trees were constructed with each subregion using the procedures described above.
-
For the purpose of investigating the dynamics of HIV in Laos, 79 near full-length sequences of HIV CRF01_AE were downloaded from the HIVDB, including two sequences sampled in Central African Republic in 1990, 30 sequences sampled in Thailand from 1990 to 2010, ten sequences sampled in Vietnam from 1997 to 1998, 31 sequences sampled in China from 1997 to 2010, two sequences sampled in Japan in 1993, and four sequences sampled in Myanmar, Indonesia, Philippines and Hongkong from 1993 to 2015, respectively (The GenBank accession number of these sequences are available as required). The downloaded sequences were aligned with the longest common subregion (HXB2: 2790–8995) of the amplified sequences. Bayesian phylogeographic analysis was performed using the BEAST v1.10.4 package as previously described (Chen et al. 2018).
-
Near full-length genomes were submitted to the Stanford University HIV drug resistance database (https://hivdb.stanford.edu/hivdb/by-sequences/). Drug resistance mutations were defined as mutations that reduce viral susceptibility to antiretroviral drugs (low-level resistance, intermediate resistance, high-level resistance) according to the drug resistance mutations list of the Stanford University HIV drug resistance database algorithm version 8.7 (HIVDR 2019).
-
The sequences amplified by this study were submitted to GenBank under accession numbers MG760371 to MG760431.
Ethics Statement
Participants
HIV Testing
HIV Sequence Generation
Phylogenetic Analysis
Phylogeographic Analysis
Drug Resistance Analysis
Sequence Data
-
From November 2011 to May 2012, a total of 2851 patients were recruited from a hospital in Vientiane, the capital of Laos, with a mean age of 32 years old. Most of these patients were Lao Loum ethnicity (85.8%), 58.8% of them were female, and 59.0% of them domiciled in Vientiane (Table 1).
Variables Total patients HIV positive Chi square P value Gender 0.018 0.894 Female 1675 40 (2.4%) Male 1176 29 (2.5%) Ethnicity 4.201 0.122 Lao Loum 2445 65 (2.7%) Lao Soung 264 3 (1.1%) Lao Theung 142 1 (0.7%) Domiciles of origin 14.514 0.024 Vientiane Capital 1682 54 (3.2%) Khammouan 72 2 (2.8%) Bolikamxai 517 10 (1.9%) Xaignabouli 57 1 (1.8%) Xiangkhoang 129 1 (0.8%) Vientiane 209 1 (0.5%) Other provinces* 185 0 *Attapu, Bokèo, Champasak, Houaphan, Louang Namtha, Louangphrabang, Oudômxai, Phoôngsali, Salavan, Savannakhet, Xékong Table 1. The prevalence of HIV among hospital patients in Vientiane Prefecture of Laos.
According to the results of the amplification of the HIV gag fragment, 2.4% of the participants (69/2851) were found to be HIV proviral DNA positive (Table 1). The prevalence of HIV among patients who domiciled from different provinces was statistically distinct (P = 0.024). Patients domiciled from Vientiane had the highest HIV prevalence (3.2%) followed by Khammouan (2.8%) and Bolikamxai (1.9%). The HIV prevalence was not significantly different between female and male, ethnic Lao Loum, Lao Soung and Lao Theung (P > 0.05).
-
From the proviral of the 69 HIV-positive patients, 61 near full-length genomes of HIV were successfully amplified and sequenced. The results of the Recombinant Identification Program that is available in HIVDB showed that 56 sequences were CRF01_AE, and the other five sequences were recombinants form of HIV subtype B and CRF01_AE. Furthermore, the ML tree showed that these 56 sequences clustered with CRF01_AE reference sequences, confirming that they were CRF01_AE (Fig. 2). The remaining five sequences did not cluster with any reference sequences and formed a unique cluster with 100% maximum parsimony bootstrap value, implying that they had a close genetic distance with each other and might be new HIV recombinant forms (Fig. 2).
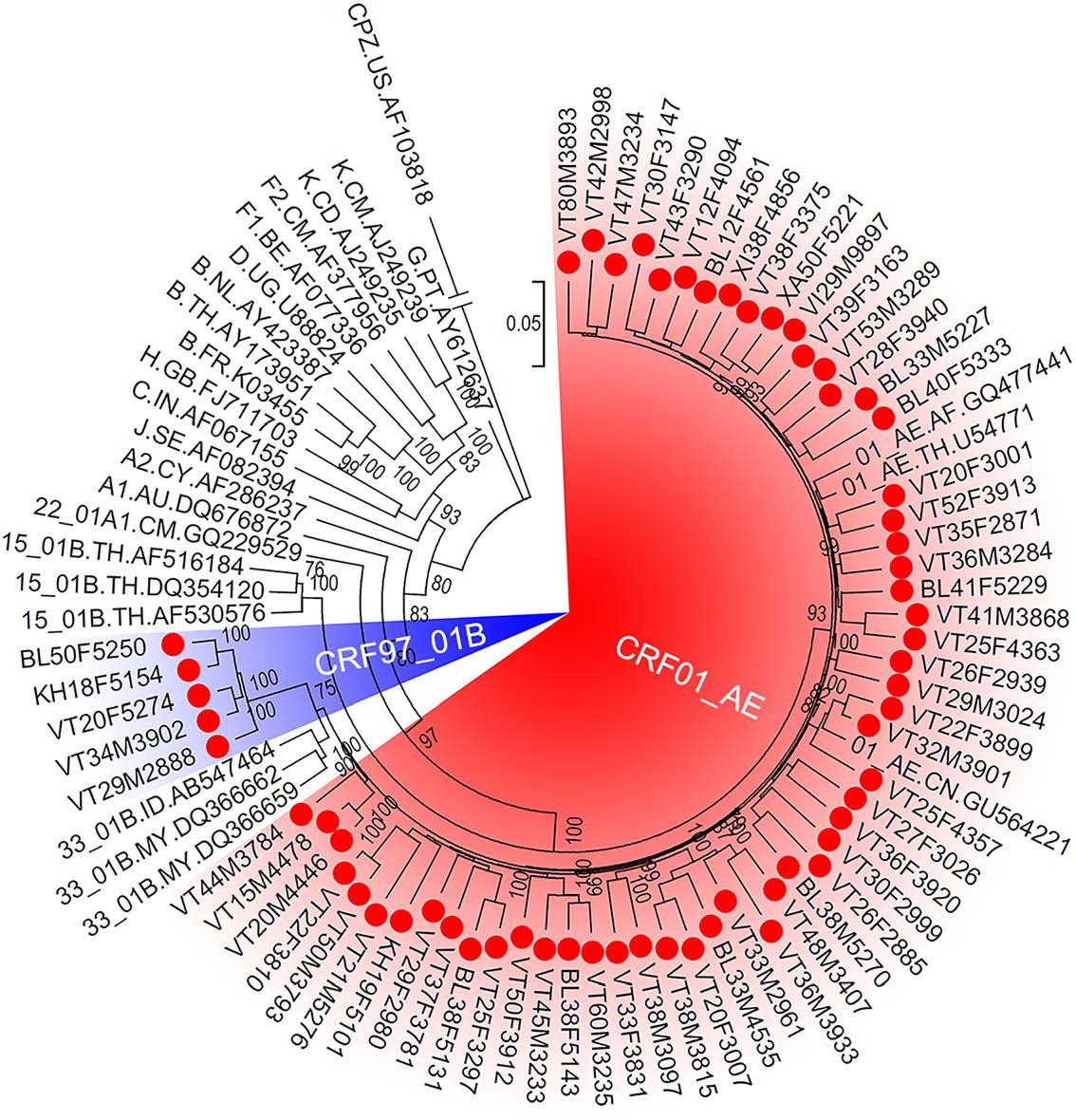
Figure 2. The maximum likelihood tree of HIV near full-length genomes. The red spots indicate the sequences that amplified from patients in Vientiane Capital, Laos. The red and blue shadows indicate HIV sequences of CRF01_AE and the newly identified CRF97_01B, respectively. The support of each branch is shown at each node, and only bootstrap values greater than 75% are shown.
-
To explore whether these five sequences were new recombinant forms and shared the same breakpoints, bootscan analysis was performed using subtypes B, F, G, H, J, K and CRF01_AE as reference subtypes. Bootscanning plots confirmed that they were recombinant forms of two subtype B fragments and two CRF01_AE fragments and shared the same breakpoints (Fig. 3A). The independent ML tree of each segment further confirmed the results of the bootscan analysis (Fig. 3B). According to the HIV nomenclature proposal, these five sequences were designated to be CRF97_01B, being the first HIV circulating recombinant form that identified in Laos. CRF97_01B was constructed of HIV CRF01_AE from 227 to 1170 nt, 2790 to 8995 nt, and subtype B from 1142 to 2794 nt, 8951 to 959 nt (Fig. 3C).
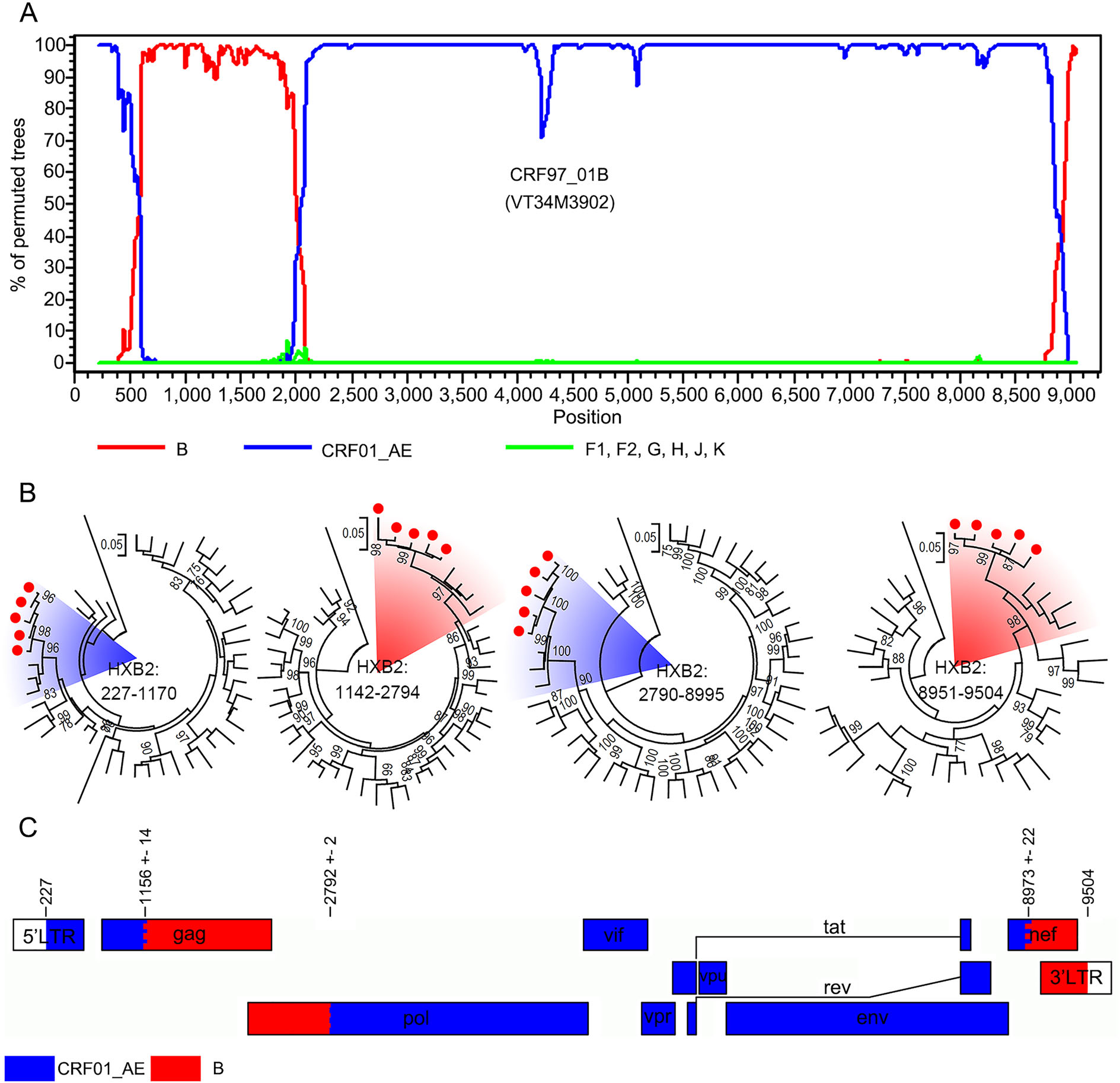
Figure 3. The identification of HIV subtype CRF97_01B. A The bootscan plots of CRF97_01B. The lines with different colors indicate different HIV subtypes. B The maximum likelihood trees based on the sub-regions of CRF97_01B. Subtype references are downloaded from the Los Alamos HIV Sequence Database, including subtype B to K, CRF01_AE and a sequence of chimpanzee. The red circles indicate the sequences that amplified from patients in Vientiane Capital, Laos. The red and blue shadows indicate sequences of CRF01_AE and subtype B. The support of each branch is indicated at each node, and only bootstrap values greater than 75% are shown. C Schematic maps of CRF97_01B. The rectangles with different colors indicate different HIV subtypes.
-
To characterize the dynamics of CRF01_AE in Laos and its bordering countries, Bayesian phylogeographic analysis was performed based on long CRF01_AE fragments. The results of the ML tree showed that CRF01_AE in Southeast Asia first circulated in Thailand (Fig. 4). All sequences formed ten main clusters, and sequences from Laos were distributed in nine of them, implying that HIV in Laos originated from multiple lineages. Sequences of Laos mainly originated from Thailand, and only three sequences originated from China (Cluster 1 in Fig. 4). Additionally, the CRF01_AE segment of CRF97_01B was derived from sequences of Laos in the late-1980s (Cluster 2 Fig. 4).
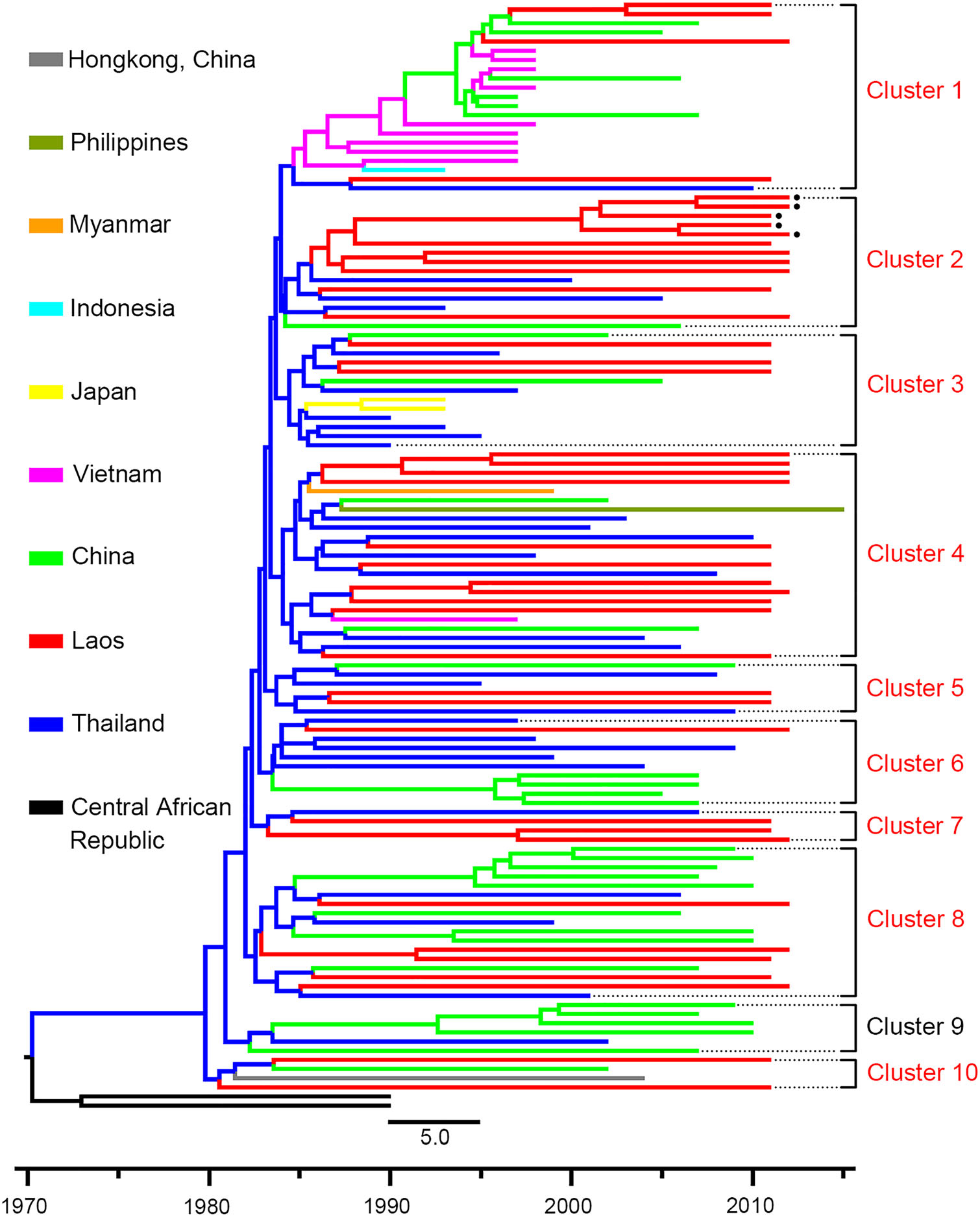
Figure 4. The maximum clade credibility tree based on sequences of HIV CRF01_AE. HIV CRF01_AE sequences (2790–8995 nt according to HXB2) in Southeast Asia are used for Bayesian phylogeographic analysis. Sequences with different origins are shown with different colors. Clusters including sequences from Laos are shown with red typeface. The black spots indicate the sequences of the newly identified CRF97_01B. The black square indicates the node of CRF97_01B, and the number indicates the origin time of CRF97_01B.
-
Among these 61 near full-length genomes, seven contained drug resistance mutations (Table 2). The mutations included A62V, D30N, E138G, E138K, F116Y, G140R, G163R, G190E, G190S, G48R, G73S, M184I, M184V, M230I, M46I, Q151M, T215A, V75I and Y181I. Among these seven sequences, one had high-level resistance to nucleoside reverse transcriptase inhibitors (NRTI), nonnucleoside reverse transcriptase inhibitors (NNRTI) and protease inhibitors (PI); two had high-level resistance to NRTI and NNRTI; one had low-level resistance to NNRTI and middle-level resistance to PI; one had middle-level resistance to NNRTI; and two had low-level resistance to NRTI or integrase inhibitors.
Samples NRTI NNRTI PI IN Mutations Resistant Mutations Resistant Mutations Resistant Mutations Resistant VT36M3933 M184I ABC (L), FTC (H), 3TC (H) E138K, G190E EFV (H), ETR (M), NVP (H), RPV (H) D30N, M46I, G48R, G73S ATV/r (L), FPV/r (L), IDV/r (L), LPV/r (L), NFV (H), SQV/ r (L) VT38M3815 A62V, V75I, F116Y, Q151M, M184V ABC (H), AZT (H), D4T (H), DDI (H), FTC (H), 3TC (H), TDF (M) Y181I EFV (M), ETR (H), NVP (H), RPV (H) VT42M2998 M184I ABC (L), FTC (H), 3TC (H) G190S, M230I EFV (H), ETR (L), NVP (H), RPV (M) VT37F3781 T215A AZT (L), D4T (L) VT27F3026 E138G RPV (L) M46I NFV (M), BL50F5250 E138K RPV (M) VT30F3147 G140R, G163R EVG (L), RAL (L) NRTI Nucleoside reverse transcriptase inhibitors, NNRTI non-nucleoside reverse transcriptase inhibitors, IN integrase inhibitors, PI protease inhibitors, L low level resistance, M intermediate resistance, H high-level resistance, ABC abacavir, FTC emtricitabine, 3TC lamivudine, AZT zidovudine, D4T stavudine, DDI didanosine, TDF tenofovir, EFV efavirenz, ETR etravirine, NVP nevirapine, RPV rilpivirine, ATV/r atazanavir/r, FPV/r fosamprenavir/r, IDV/r indinavir/r, LPV/r lopinavir/r, NFV nelfinavir, SQV/r saquinavir/r, EVG elvitegravir, RAL raltegravir Table 2. The drug resistance mutations of HIV sequences amplified from patients in Vientiane Capital, Laos.
Demographic Characteristics and HIV Prevalence
HIV Subtyping
Identification of New HIV Circulating Recombinant Form
Dynamics of CRF01_AE in Southeast Asia
Drug resistance Characteristics
-
According to the reports of UNAIDS, countries in Southeast Asia exhibit different prevalence of HIV (Fig. 5). The HIV prevalence in Thailand, Cambodia and Vietnam has passed the peaked stage and exhibits a downward tendency, while the HIV prevalence in Myanmar and Laos has reached the peaked stage and exhibits a plateau phase. Previous studies revealed that the HIV prevalence in Laos was 0.4% among pregnant women in 1996 and 5.6% among men who had sex with men in 2007 (Loue et al. 1998; Sheridan et al. 2009). To the best of our knowledge, these studies were the only ones conducted regarding the HIV prevalence of Laos. In the present study, we found that the HIV prevalence among 2851 hospital patients in Vientiane was 2.4%, eight times higher than the report of UNAIDS (Fig. 5). All the results above implied that the HIV prevalence in Laos might not be as low as previously believed. Additionally, it should be noted that the participants in this study were recruited from the hospital, and the HIV prevalence among them was more similar to the general population than to the high-risk groups. Moreover, the HIV prevalence of Laos was determined by amplifying proviral DNA in the present study, and these results might be underestimated compared with the serological test, mainly due to the long period of storage and primer specificity (Assih et al. 2016). All these results indicated that Laos might be suffering a serious impact from HIV.
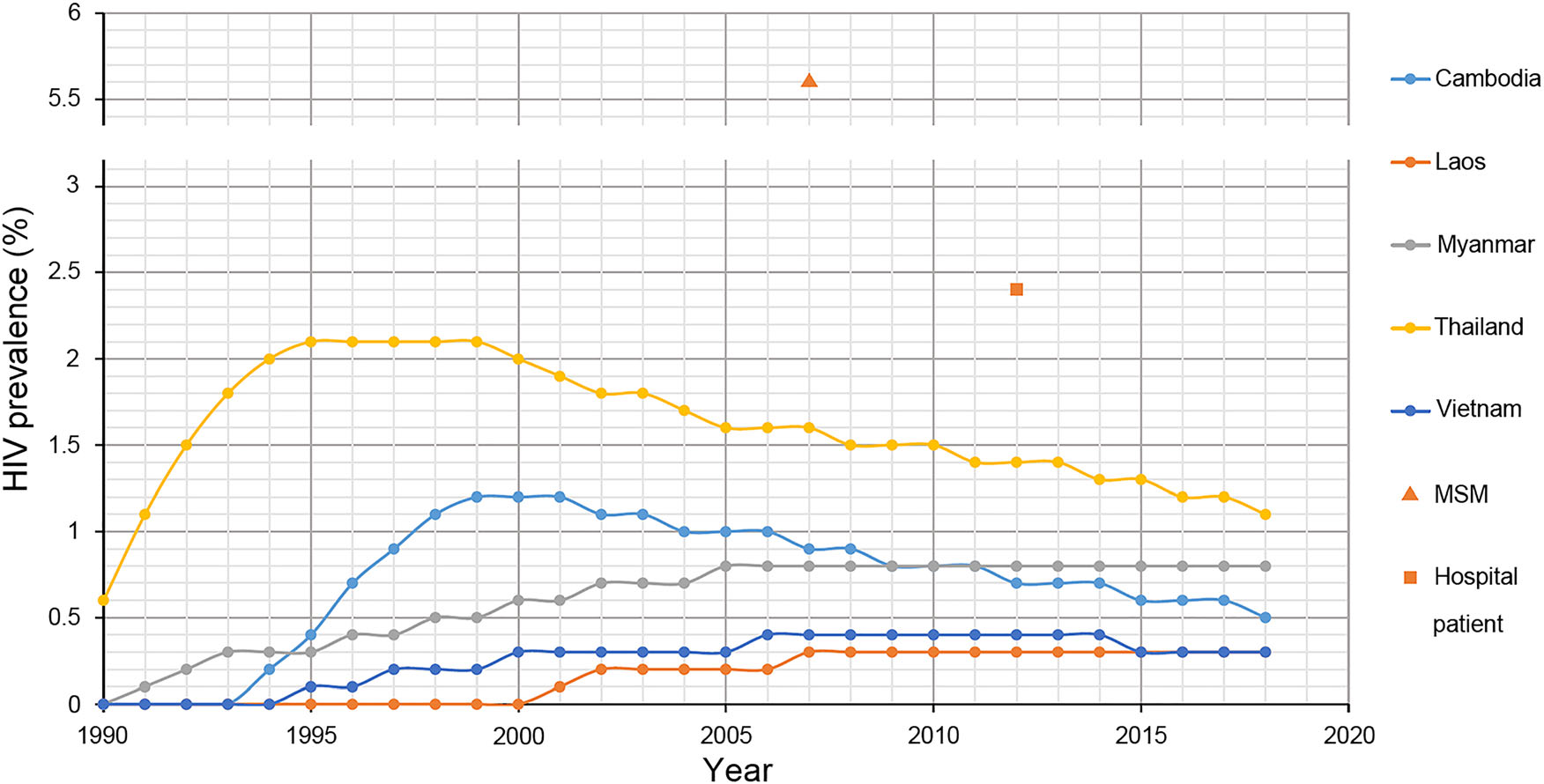
Figure 5. The prevalence of HIV in Southeast Asia. The lines with different colours indicate the HIV prevalence of different countries in Southeast reported by the Joint United Nations Programme on HIV/ AIDS. The line overlapping with the abscissa means that the HIV prevalence was less than 0.1% (UNAIDS 2017). The triangle indicates the HIV prevalence among 540 men who have sex with men in Vientiane Capital, Laos (Sheridan et al. 2009). The square indicates the HIV prevalence among 2851 hospital patients in Vientiane Capital, Laos.
The molecular characteristics of HIV are rapidly evolving in Southeast Asia. HIV subtypes B and CRF01_AE were co-circulating in Thailand in 1980s, and lead the formation of numerous recombinant strains, such as CRF15_01B and CRF34_01B (Liao et al. 2009; Li et al. 2012; Tovanabutra et al. 2003, 2007). Loue et al. (1998) conducted a study among pregnant women in Vientiane and found that the two sequences amplified from HIV-positive samples were identified as CRF01_AE. Unfortunately, these two sequences were not available for analysis. In the present study, we successfully amplified 61 HIV near full-length sequences from patients visiting a hospital in Vientiane and found that CRF01_AE was predominant in this region, which was in accord with the major subtype of the surrounding countries, including Thailand, Cambodia, Vietnam (Phanuphak et al. 2015). Unexpectedly, a new HIV circulating recombinant form that was constructed by subtype B and CRF01_AE was identified in the present study, accounting for 8.2% of the sequences of Laos. Although several subtype B fragments had been amplified and sequenced in the present study, we could not amplify the near full-length sequences successfully before the blood samples were used up, mainly due to primer specificity (Assih et al. 2016). These results revealed that the molecular characteristics of HIV in Laos might have been gaining complexity in recent years.
The prevention of HIV is no longer a unique event in Southeast Asia, and cross-border transmission has been demonstrated to be a crucial factor for the formation of HIV recombinant forms between different subtypes (Zhou et al. 2014). Although there are no available HIV sequences of Laos in HIVDB, the cross-border transmission of HIV subtypes B, C, CRF01_AE, CRF07_BC and CRF08_BC were well studied between other countries in Southeast Asia (Deng et al. 2008; Tee et al. 2008; Liao et al. 2009; Li et al. 2010, 2012; Liu and Zhang 2011; Pang et al. 2012; Chen et al. 2017b). In the present study, we confirmed that CRF01_AE in Laos was introduced from Thailand in the 1980s. Although the CRF01_AE fragments of our newly identified CRF97_01B had a closer relationship with sequences of Laos, we could not draw a conclusion that CRF97_01B were formed by subtype B and CRF01_AE in Laos, as we failed to provide full sequences of subtype B from the present study. To achieve a thorough understanding of dynamics of CRF97_01B and to prevent the cross-border transmission of HIV, more nationwide studies are needed in Laos.
Since 2001, ART for people living with HIV was available in Laos; while, the proportion of more than 95% adherence to ART was 60%, much lower than its neighbor countries, such as China and Vietnam (Tran and Houston 2012; Hansana et al. 2013; Zuo et al. 2019). The present study revealed that the drug resistance prevalence among HIV patients in Laos was 11.5%. As the status of antiviral therapy of the participants was unclear, we could not distinguish the proportion of transmitted and acquired drug resistance. To monitor the drug resistance characteristics of HIV in Laos, more recent and systematic researches are urgently needed.
Two limitations should be considered when interpreting the results of the present study. First, the present study was a retrospective study, and the samples were collected in 2011 and 2012. The results of the present study could only indicate the HIV prevalence situation in 2011 and 2012. Second, although thousands of participants with different domiciles of origin were recruited, the sample site was only in the capital of Laos, and it could not reflect the exact HIV prevalence in various provinces of Laos. Despite these limitations, the present study showed clearly that patients in Vientiane, the capital of Laos, are affected by HIV CRF01_AE and the newly identified CRF97_01B and noted that more nationwide surveys and preventive measures are urgently needed.
In summary, we performed a retrospective study among 2851 hospital patients in Vientiane from 2011 to 2012 and found that the HIV prevalence among this group was 2.4%, eight times more than the report of UNAIDS. Phylogenetic and phylogeographic analysis based on HIV near fulllength genomes revealed that the predominant strain among them was CRF01_AE, which mainly originated from Thailand through multiple introductions. A new HIV circulating recombinant form was also identified, known as CRF97_01B. These results filled an important gap in the molecular epidemiology and evolution of HIV in Southeast Asia and called for immediate measures to prevent Laos from exhibiting the same increase in HIV prevalence as its bordering countries.
-
We thank Professor Qing-Peng Kong (Kunming Institute of Zoology, Chinese Academy of Sciences) for the assistance in collecting samples. We also thank all the participants who participated in this research. This work was supported in part by Grants from the National Natural Science Foundation of China (U1302224, 81601802, 81271892, 81902106). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
-
YTZ and CZ conceived and designed the research. XC, MY and YW performed the HIV provirus amplification experiments. YTZ, CZ and XC analyzed the data and wrote the manuscript. All authors revised and approved the final manuscript.
-
The authors declare that there are no competing interests.
-
Additional informed consent was obtained from all patients for which identifying information is included in this article.
















 DownLoad:
DownLoad: