-
By Oct. 29, 2020, in a matter of 10 months since the beginning of the COVID-19 outbreak, more than 43, 000, 000 confirmed cases of COVID-19 had been reported, with more than 1, 100, 000 deaths (WHO 2020). The origin and intermediate host of SARS-CoV-2, the virus responsible for COVID-19, have not been identified yet. However, it has been confirmed that this virus is mainly transmitted by respiratory droplets and direct contact (Liu Y et al. 2020). Moreover, SARS-CoV-2 viral nucleic acid was detected in the feces of patients (Wu Y et al. 2020), which has been observed in other coronavirus infection cases (Guan et al. 2015; Mackay and Arden 2015), and a rise in immunoglobulin M (IgM) antibody levels was confirmed in newborns, suggesting that SARS-CoV-2 can also be transmitted through the fecal-mouth route and vertically from mother to child (Dong et al. 2020). Humans of all ages are susceptible to SARS-CoV-2 infection, which is spreading more efficiently than influenza. Therefore, it is crucial and pressing to search for effective treatment strategies to cope with such an unprecedented health emergency.
There are currently no approved drugs or vaccines for COVID-19, highlighting the huge unmet medical need to develop treatments. Convalescent plasma therapy has been proven successful in conferring protection against SARS-CoV, MERS-CoV, avian influenza, and Ebola virus infections, considerably alleviating clinical symptoms and reducing mortality (Bloch et al. 2020). The available data also support the feasibility of this treatment for SARS-CoV-2 (Devasenapathy et al. 2020). However, a large-scale convalescent plasma transfusion program is conditioned by the limited availability of convalescent plasma and the lack of appropriate risk assessment (Shi et al. 2020). On the other hand, neutralizing monoclonal antibodies (mAbs) have attracted extensive interest as promising candidates to fight against emerging viruses. The administration of mAbs for passive immunization can provide immediate and specific protection and complement the development of preventive vaccines, thus potentially having a significant impact on the control of the COVID-19 pandemic. The successful use of mAb therapy (mAb114) during the Ebola virus outbreak highlighted the effectiveness and safety of mAbs (Corti et al. 2016). More importantly, the therapeutic potential of antibodies targeting coronaviruses was well recognized during the SARS outbreak (Wang C et al. 2020).
HTML
-
The Coronaviridae family is subdivided into four genera: α, β, γ, and δ (Fehr and Perlman 2015). Among the seven coronaviruses that have been reported to infect humans, HCoV-229E and HCoV-NL63 belong to genus α; the remaining five viruses are all classified into genus β, including HCoV-OC43, HCoV-HKU1, SARS-CoV, MERS-CoV, and SARS-CoV-2. Specifically, infections of HCoV-229E, HCoV-NL63, HCoV-OC43, and HCoV-HKU1 mainly cause mild respiratory diseases, while infection by SARS-CoV-2 may eventually lead to severe pneumonia and death, similarly to infection by SARS-CoV (~ 10% mortality rate) and MERS-CoV (~ 36% mortality rate), both of which have induced worldwide outbreaks in the past two decades (Zaki et al. 2012; Ying et al. 2015).
Previous studies revealed that the SARS-CoV-2 genome has 29, 891 nucleotides and encodes a single open reading frame (ORF) flanked by two untranslated regions (UTRs). Two-thirds of the genome are proteolytically cleaved into 16 putative nonstructural proteins (nsps) and four main structural proteins, which are the spike protein (S), membrane protein (M), envelope protein (E), and nucleocapsid (N) and are encoded at the 3′ end of the viral genome (Chan et al. 2020).
Host cell entry is mediated by the S protein by binding of its S1 subunit to angiotensin-converting enzyme 2 (ACE2) receptors on the cell surface, which subsequently triggers conformational changes on the S2 subunit, thus allowing the virus to enter the host cells via endocytosis. S1 is composed of the N-terminal domain (NTD) and receptor-binding domain (RBD), which is crucial to determine tissue tropism and host ranges (Walls et al. 2020). SARS-CoV-2 uses ACE2 for entry and the serine protease TMPRSS2 for S protein priming, which facilitates SARS-CoV-2 entry into host cells (Markus et al. 2020; Dong et al. 2020). After entry, SARS-CoV-2 is uncoated and the single positive (+) strand RNA is released into the cytoplasm and subsequently translated to produce polypeptides, which are cleavaged into nsps by papain-like protease and Mpro or 3CLpro. These nsps include helicase and RNA-dependent RNA polymerase (RdRp), which will participate in the process of RNA synthesis. As well as nsps, structural and other proteins are encoded by viral genome after RNA synthesis. Eventually, the structural protein and genome are assembled into a new virion and released by exocytosis (Sanders et al. 2020).
-
Currently, the serological characteristics of COVID-19 patients are partially documented. Xu et al. examined the IgM and IgG levels in 17, 368 individuals, including patients, related hospital workers, and residents in an involved community, to assess the cumulative prevalence of SARS-CoV-2 infection in China (Xu et al. 2020). They reported that the seropositivity in Wuhan varied between 3.2% and 3.8% and in a subcohort of 242 patients with infection confirmed by a viral nucleic acid RT-PCR test, the cumulative seroprevalence rates of IgM and IgG antibodies against the recombinant antigens containing the nucleoprotein and a peptide from the S protein of SARS-CoV-2 on the 7th day after the onset of symptoms were 44% and 56%, respectively, and both reached over 95% by day 20 and day 16, respectively (Xu et al. 2020). Moreover, Zhao et al. showed that among 173 patients the seroconversion rate for total antibodies (Ab), IgM, and IgG within one week since the onset was 93.1%, 82.7%, and 64.7%, respectively, and after two weeks the proportion greatly increased to 100.0% (Ab), 94.3% (IgM), and 79.8% (IgG) (Zhao et al. 2020).
Wang et al. further reported that mean antibody neutralization titers against SARS-CoV-2 live virus were ~ sevenfold higher in the sicker patients and that patients with severe COVID-19 had more robust binding antibodies to both the N protein and S trimer (Wang P et al. 2020). Even though antibodies against the SARS-CoV-2 N protein do not provide protective immunity against infection (Gralinski and Menachery 2020), specific antibodies against the S protein do (Gao et al. 2020). Additionally, another study demonstrated that RBD-specific IgG accounts for half of the S protein-antibody response in COVID-19 patients during recovery, indicating that RBD is the primary immunogen (Liu L et al. 2020).
Nevertheless, sera collected from SARS and COVID-19 patients exhibited limited cross-neutralization, as demonstrated by Hou et al. using luciferase reporter viruses. The study showed that convalescent COVID-19 sera failed to cross-neutralize SARS-CoV in vitro and rare cross-neutralizing antibodies were generated after SARS-CoV-2 infection (Hou et al. 2020), which was also reported in other studies (Ou et al. 2020; Anderson et al. 2020). Thus, recovery from SARS-CoV-2 infection may not confer resistance to SARS-CoV and vice versa.
Additionally, Okba et al. detected MERS-CoV-S cross-reactive antibodies in the serum of a COVID-19 patient, but not MERS-CoV-S1 cross-reactive antibodies, which is consistent with their earlier findings for MERS-CoV serology (Okba et al. 2020). This finding revealed a higher specificity of S1 for SARS-CoV-2 antibodies, which could be due to the high degree of conservation in the CoV S2 subunit compared to S1. Thus, S1 is more specific than S as an antigen for SARS-CoV-2 serological diagnosis.
Clinical application of convalescent plasma in COVID-19 patients demonstrated that neutralizing antibodies provide important specific immune defense against viral infections. It has been reported neutralizing antibodies in asymptomatic patients decay faster than symptomatic individuals (Long et al. 2020; Ibarrondo et al. 2020). In a large cohort investigation, Stefansson et al. tested 2102 samples collected from 1237 persons in Iceland, and observed antibodies persisted for as long as 4 months after diagnosis (Baum et al. 2020). Moreover, 14 serum specimens from 17 COVID-19 patients at 6–7 months post-infection showed significant neutralizing activities in Tan's research (Tan et al. 2020). It is necessary to conduct further studies about the dynamics of neutralizing antibodies and the duration of humoral immunity. The differences among these researches might be resulted from many factors, such as enrolled cases of different ethnic groups and geographical regions, varied sampling and testing methods and so on.
Collectively, these findings provide strong empirical support for the serological diagnosis and management of COVID-19 patients, as well as antigen selection for candidate vaccines and targeted antibody development.
-
Therapeutic neutralizing antibodies need to go through a long process from isolation to clinical application. In vitro assays of neutralizing activity, in vivo evaluation of protective effect, and safety and clinical trials are necessary steps for antibody development. Researchers are endeavoring to develop such mAbs or their functional fragments as prophylactic or therapeutic agents to prevent or treat SARS-CoV-2 infection. Therefore, compared to the usual timelines, the development of mAbs in this pandemic setting could be reduced in 5–6 months. Moreover, the previous researches on SARS-CoV and MERS-CoV antibodies provide a basis for these studies (Wang HY et al. 2020).
47D11 was the first SARS-CoV-2-neutralizing human monoclonal antibody to be reported, which was screened from readily available hybridoma libraries of genetically modified mice that had been immunized with SARS-CoV and MERS-CoV S proteins (Wang C et al. 2020). In vitro experiments showed 47D11 could neutralize authentic SARS-CoV and SARS-CoV-2 with IC50 values of 0.19 μg/mL and 0.57 μg/mL, respectively, indicating that antibodies isolated from SARS survivors may play a huge role in our fight against SARS-CoV-2 (Table 1).
Source Antibodies Cross reactivity Blocking receptor binding Target Complex crystal (PDB ID) Binding affinity (Kd nmol/L) Neutralizing effect (IC50 μg/mL or nmol/L) Current status Refs. Convalescent SARS patients 47D11 Yes No RBD NA 9.6b 0.57† μg/mL Under research Wang C et al. (2020) S309 Yes No RBD (Site B) 6WPS 0.001b 0.079† μg/mL Under research Pinto et al. (2020) CR3022 Yes No RBD (Site C) 6W41 < 0.1b No Under research Yuan et al. (2020b) Immunized llama VHH-72-Fc Yes Yes RBD NA 38.6a 0.2* μg/mL Under research Wrapp et al. (2020a) Convalescent COVID-19 patients P2C-1F11 No Yes RBD (Site A) NA 2.12a 0.03† μg/mL Under research Ju et al. (2020) P2B-2F6 No Yes RBD (Site A) 7BWJ 5.14a 0.41† μg/mL Under research CA1 No Yes RBD (Site A) NA 4.68 ± 1.64a 4.981† μg/mL Under research Shi et al. (2020) CB6 (JS016) No Yes RBD (Site A) 7C01 2.49 ± 1.65a 0.835† μg/mL Phase-I B38 No Yes RBD (Site A) 7BZ5 70.1a 0.177† μg/mL Under research Wu Y et al. (2020) H4 No Yes RBD (Site A) NA 4.48a 0.896† μg/mL Under research 414-1 No Yes RBD (Site A) NA 0.31c 1.75† nmol/L Under research Wan et al. (2020) 505-3 No Yes RBD (Site A) NA 0.08c 3† nmol/L Under research BD-23 NA Yes RBD (Site A) 7BYR 4.3a 8.5† μg/mL Under research Cao et al. (2020) BD-368-2 NA Yes RBD (Site A) NA 0.82a 0.015† μg/mL Under research CC6.29 No Yes RBD (Site A) NA 1.20a 0.002* μg/mL Under research Rogers et al. (2020) CC6.30 No Yes RBD (Site A) NA 1.71a 0.001* μg/mL Under research CC12.1 No Yes RBD (Site A) 6XC2 5.92a 0.022* μg/mL Under research Yuan et al. (2020a) CC12.3 No Yes RBD (Site A) 6XC4 54.3a 0.026* μg/mL Under research 4A8 NA No NTD (Site D) 7C2L 1.00b 0.39† μg/mL Under research Chi et al. (2020) 2–4 NA Yes RBD (Site A) 6XEY NA 0.057† μg/mL Under research Liu LH et al. (2020) 2–15 NA Yes RBD NA NA 0.0007† μg/mL Under research 4–8 NA No NTD NA NA 0.009† μg/mL Under research 2–43 NA Yes RBD NA NA 0.003† μg/mL Under research COV2-2196 No Yes RBD NA NA 0.015† μg/mL Under research Zost et al. (2020) COV2-2130 No Yes RBD NA NA 0.107† μg/mL Under research S2M11 No Yes RBD (Site A) 7K43 66a 1.66† μg/mL Under research Poh et al. (2020) S2E12 No Yes RBD (Site A) 7K4N 1.6a 5.29† μg/mL Under research LY-CoV555 No Yes RBD NA NA NA Phase III (pause) Jones et al. (2020) REGN10933 No Yes RBD 6XDG 0.0140a 0.043† nmol/L Phase III Baum et al. (2020) VelocImmune mice REGN10987 No Yes RBD 6XDG 0.0298a 0.041† nmol/L Phase III Phage library n3088 NA No RBD (Site C) NA 12.6b 3.3* μg/mL Under research Wu YL et al. (2020) n3130 NA No RBD (Site C) NA 32.5b 3.7* μg/mL Under research n3086 NA No RBD NA 88.97b 26.6* μg/mL Under research n3113 NA No RBD NA 57.01b 18.9* μg/mL Under research H014 Yes Yes RBD (Site C) NA 0.33b 1* nmol/L Under research Lv et al. (2020) †The data were derived from authentic virus neutralization experiments
* The data were derived from the pseudovirus neutralization experiments
aThe data obtained using surface plasmon resonance (SPR)
bThe data obtained using biolayer interferometry (BLI)
cThe data obtained using enzyme-linked immunosorbent assay (ELISA)
*The data were derived from the pseudovirus neutralization experiment (other data derived from authentic virus neutralization experiments)Table 1. Characteristics of representative antibodies to SARS-CoV-2.
Many neutralizing antibodies derived from B cells of patients infected with SARS-CoV have been reported until now, such as S309 (Pinto et al. 2020), CR3022, CR3014 (Yuan et al. 2020b), 80R (Hwang et al. 2006), and m396 (Prabakaran et al. 2006), some of which showed potent cross-reactivity to SARS-CoV-2. S309 can neutralize both SARS-CoV and SARS-CoV-2, while CR3022, CR3014, 80R, and m396 can only neutralize SARS-CoV. IC50 value of S309 for SARS-CoV-2 neutralization was 0.079 μg/mL. Interestingly, CR3022 also showed cross-reactivity with SARS-CoV-2 RBD (Table 1).
In addition, Wrapp et al. obtained a single-domain antibody (VHH), VHH-72, derived from a llama immunized with SARS-CoV and MERS-CoV S proteins, that can bind tightly to the RBD region of SARS-CoV S protein and neutralize SARS pseudoviruses. By further engineering this VHH into a bivalent Fc-fusion protein, they showed that this novel bivalent molecule could also neutralize SARS-CoV-2 pseudoviruses with an IC50 of approximately 0.2 μg/mL (Wrapp et al. 2020a) (Table 1).
As with other human pathogens, mAb can be easily isolated from the B cells of patients infected with SARS-CoV-2 by the antigenic protein bait. Thus, the protein bait needs to be functionally relevant and pure enough to isolate antibodies that are relevant to the epitopes on the native spike. The RBD domain and S1 subunit of SARS-CoV-2 are often selected as protein bait for fishing out the antibodies. Severe cases seem to have a much higher neutralizing antibody response and hence more likelihood of obtaining potent neutralizing antibodies (Zhao et al. 2020). However, it is unclear about the relationship between the severity of diseases and antibody response. It will take a while to figure out the cause and consequence of the two.
Ju et al. first analyzed the immune responses of eight SARS-CoV-2-infected patients after exposure to virus stimulation, including the level of viral-specific antibodies in plasma and the proportion of specific B cells in memory cells, finding that the antibody response levels among these infected patients varied greatly (Ju et al. 2020). Further, this group isolated 206 monoclonal antibodies capable of recognizing SARS-CoV-2 RBD and obtained their paired heavy- and light-chain gene sequences. Using a pseudovirus evaluation system, the antibody inhibition rate of 12 strains exceeded 50% and that of the other seven strains exceeded 80%. Among them, P2C-1F11 and P2B-2F6 were elite antibodies with IC50 of 0.03 μg/mL and 0.05 μg/mL, respectively (Table 1).
Moreover, Shi et al. used SARS-CoV-2 S protein recombinant RBD as bait to screen specific memory B cells in peripheral blood mononuclear cells (PBMCs) from peripheral blood of convalescent patients with COVID-19 (Shi et al. 2020). The variable region sequences of these B cell receptors were cloned into a vector containing a constant region to produce a series of IgG antibodies. Among them, CB6 was selected for testing in animal models and the results showed that this antibody not only reduced the virus titer but also inhibited the pathological lung damage in the monkeys challenged with SARS-CoV-2.
Meanwhile, a team led by George Fu Gao isolated four human mAbs (B38, H4, B5 and H2) from a rehabilitated patient, all of which have shown in vitro neutralization of SARS-CoV-2 and two of which (B38 and H4) block the binding of RBD to the cellular receptor ACE2 (Wu et al. 2020b) (Table 1).
On the other hand, Wan et al. enriched more than 1000 antigen-specific B cells in plasma from 11 COVID-19 rehabilitation patients by using the recombinant S1 or RBD antigen of SARS-CoV-2 and cloned 729 pairs of naturally paired antibody genes of single B cells (Wan et al. 2020). The team mainly reported 11 different neutralizing antibody sequences, eight of which showed an IC50 within 10 nmol/L and the best one, 414-1, with an IC50 of 1.75 nmol/L. Besides, 553-15 could be combined with other neutralizing antibodies and increase their neutralizing ability (Table 1).
Meanwhile, Cao et al. used high-throughput cell sequencing to identify 14 highly active neutralizing antibodies from a screening of 8558 viral protein-binding antibody sequences, the most effective of which was a neutralizing antibody called BD-368-2 (Cao et al. 2020). Unlike traditional methods, large-scale data obtained from high-throughput single cell can help researchers observe the clonal enrichment of B cells before antibody expression in vitro and calculate the degree of enrichment based on the number of cloned cells observed. The researchers suggest that enriched B cell clones are more likely to produce novel binding SARS-CoV-2 neutralizing antibodies with high affinity. In addition, by using bioinformatics methods to predict the structure of CDR3H, researchers found mAbs with highly similar CDR3H structures to the SARS-CoV neutralizing antibody m396 showed a high neutralization potency for SARS-CoV-2. The largely conserved combination of the VDJ gene segment and the likely overlapping epitopes of those mAbs suggest the existence of stereotyped B cell receptors against SARS-CoV-2 (Cao et al. 2020). A new study supports this idea: Yuan et al. compiled a list of 294 SARS-CoV-2 RBD-targeting antibodies where information on IGHV gene usage is available and found that IGHV3-53 is the most frequently used IGHV gene among these antibodies (Yuan et al. 2020a) (Table 1).
Furthermore, Chi et al. isolated and identified mAbs from 10 recovering COVID-19 patients. Three of these mAbs showed neutralizing activity against SARS-CoV-2. 4A8 showed high potency against both SARS-CoV-2 and its pseudovirus and it works by binding to a unique region in the NTD rather than RBD of the S protein (Chi et al. 2020) (Table 1).
To study the antibody response against SARS-CoV-2 and discover mAbs, the Scripps research team adapted their pipeline to rapidly isolate and characterize mAbs from convalescent donors (Rogers et al. 2020). A group of SARS-CoV-2 donors who had previously tested positive on the swab test was recruited for PBMC and plasma collection. Assessed by the neutralizing activity of convalescent serum against SARS-CoV and SARS-CoV-2 and selected eight donors for mAb discovery. They classified single antigen-specific memory B cells and recovered and cloned the corresponding variable genes using a high-throughput production system that could achieve antibody expression and identification within 2 weeks. More than 1000 mAbs were isolated from three convalescent donors by memory B cell selection using SARS-CoV-2 S or RBD recombinant proteins. Notably, the most potent neutralizing antibodies were those directed to the core region of RBD, known as the receptor-binding motif (RBM), including two antibodies, CC6.29 and CC6.30, that neutralize SARS-CoV-2 pseudovirus with an IC50 of 2 ng/mL and 1 ng/mL, respectively (Table 1).
Liu LH et al. (2020) isolated 61 SARS-CoV-2 neutralizing antibodies from 5 severe COVID-19 patients. Among them, nine antibodies showed very high neutralization potency, with IC50 values ranging from 0.7 to 9 ng/mL. The nine antibodies are diverse and can be divided into three categories, four against RBD (2–15, 2–7, 1–57, and 1–20), three against NTD (2–17, 5–24, and 4–8), and two against the undetermined region of S trimer (2–43 and 2–51). Notably, 2–15, binding to the top region of RBD, exhibits the most competitive against ACE2 and the most neutralizing activity to SARS-CoV-2 (Table 1).
In addition, Wu YL et al. (2020) developed a new technique to screen and reconstruct the full human heavy-chain variable region antibody skeleton. Using this technique, a whole human nanometer antibody library based on natural germline genes was designed. The RBD domain and S1 protein subunit of SARS-CoV-2 were selected as antigens, and nano-antibodies targeting multiple epitopes were selected from the antibody library. Among them, antibodies n3130, n3088, n3086, and n3113 showed moderate neutralization activities, inhibiting SARS-CoV-2 pseudovirus infection in a dose-dependent manner with IC50 values of 26.6, 18.9, 3.3, and 3.7 μg/mL, respectively. The combination of n3088 or n3130 with n3113 showed synergistic neutralization of SARS-CoV-2, with IC50 values of 0.51 and 0.70 μg/mL, respectively. Similarly, Lv et al. recently established a coronavirus antibody library by using phage display technology and identified an antibody, H014, with broad-spectrum neutralization of the coronavirus family by high-throughput screening (Lv et al. 2020). Pseudovirus neutralization assays revealed that H014 has a potent neutralizing activity, with an IC50 of 3 nmol/L and 1 nmol/L against SARS-CoV-2 and SARS-CoV pseudoviruses, respectively (Table 1).
With the advancement of research, many antibody drugs have entered clinical trials. It is worth mentioning that on June 5, 2020, the China Food and Drug Administration officially approved the first clinical trial of a novel therapeutic antibody, mAb CB6 derived from a convalescent patient, which is now at the edge of phase I. Additionally, on June 11, 2020, REGN announced the clinical trial of its antibody cocktail therapy for the treatment and prevention of COVID-19. It contains two antibodies, REGN10933 and REGN10987, which were obtained by regeneration from the VelocImmune transgenic mouse platform and COVID-19 rehabilitation patient blood based on the screening of the efficacy and binding ability of the S protein (Baum et al. 2020). Moreover, more than 10 antibody drugs such as LY-CoV555 (phase III), AZD7442 (phase I), CT-P59 (phase I) and TY027 (phase I) are already in clinical trials (https://clinicaltrials.gov/), giving us the hope of dealing with the COVID-19.
-
The spike (S) glycoprotein of SARS-CoV-2, which forms homotrimers protruding from the viral surface, can mediate virus entry into host cells (Wrapp et al. 2020b). The S protein is a class I fusion protein that consists of two subunits, S1 and S2. The S1 subunit is responsible for binding to host cell receptors and the S2 subunit mediates the fusion of viral and cellular membranes (Wrapp et al. 2020b).
Besides its pivotal role for virus entry, S protein is also the main target of neutralizing antibodies and the focus of therapeutic and vaccine design efforts. The S protein amino acid sequence homology between SARS-CoV-2 and SARS-CoV is 80% and between SARS-CoV-2 and the bat SARS-related CoV (SARSr-CoV) RaTG13, 97.2% (Zhou et al. 2020). Previous studies revealed that SARS-CoV-2 has a higher affinity for hACE2 than SARS-CoV. Compared with the RBD of SARS-CoV, the binding conformation of ACE with SARS-CoV-2 RBD is more compact. In addition, the substitution of several residues in SARS-CoV-2 RBD can make two virus-binding hotspots at the RBD-ACE2 interface more stable. Therefore, SARS-CoV-2 RBD has a higher binding affinity for ACE2 (Shang et al. 2020). As RBD-specific antibodies have a brilliant neutralizing ability for different virus strains, SARS-CoV-2 RBD can provide a basis for the study of antibody targets for highly specific and high-affinity antibodies (Wrapp et al. 2020b).
S glycoprotein is in a pre-fusion conformation before contacting the cell receptor and can promote virus fusion with the host cell membrane through several structural rearrangements (Wrapp et al. 2020b). The binding of S protein to the host receptor destroys the stability of the pre-fusion trimer, resulting in the shedding of the S1 subunit and the transformation of the S2 subunit into a highly stable post-fusion conformation. It should be noted that in order to attach the host receptor, the RBDs in S1 subunits require hinge-like conformational movements, with one RBD fully exposing the determinants of receptor binding, while the other two are hidden. Thus, the predominant state of the trimer exists as one "up" (receptor-accessible) and two "down" (receptor-inaccessible) conformations (Fig. 1).
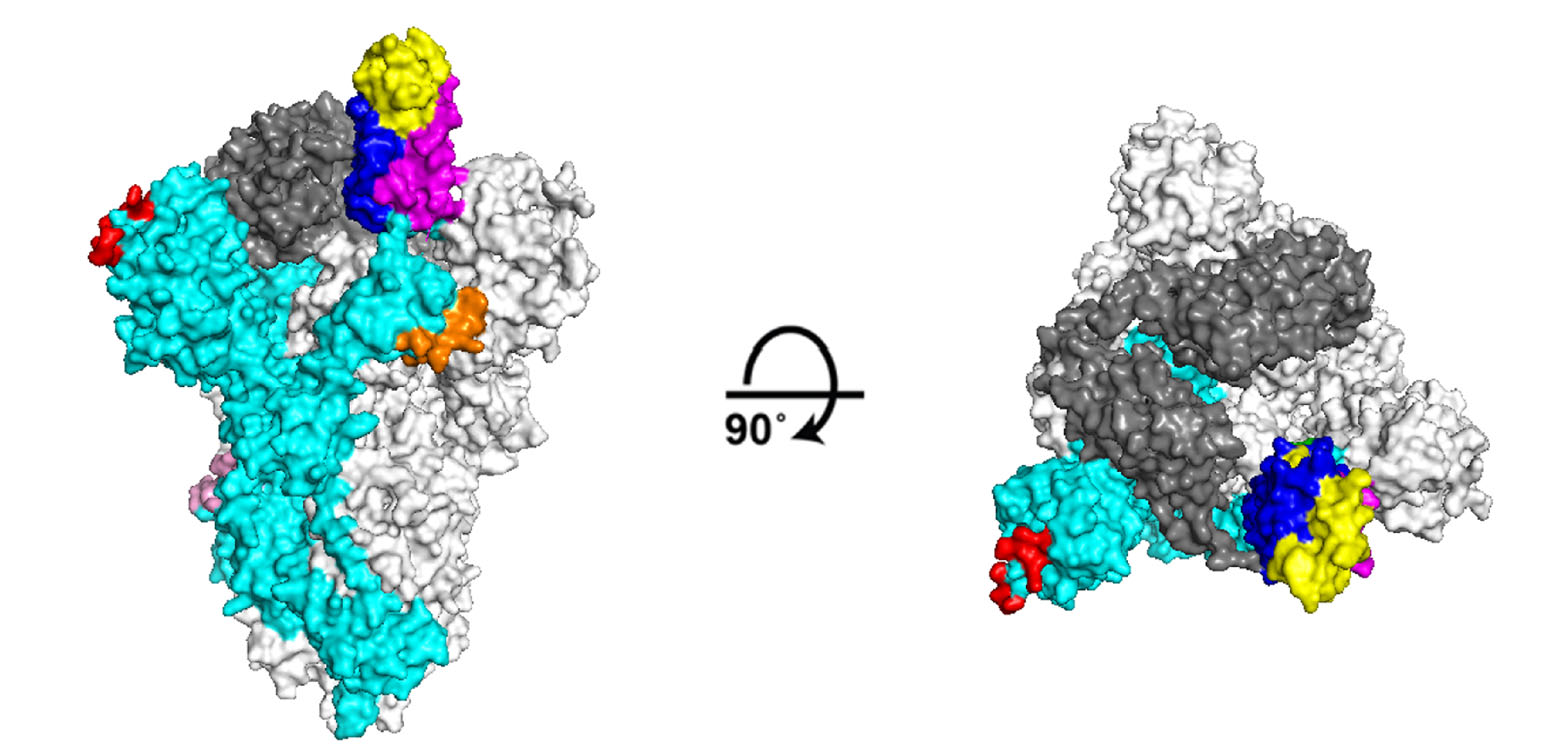
Figure 1. Structure of SARS-CoV-2 S trimer in the prefusion conformation displayed as molecular surfaces. Side (left) and top (right) views of the S trimer with a single RBD in the up" conformation is shown (PDB ID 6VYB). The monomer corresponding to the "up" conformation is marked as cyan, and the monomers corresponding to the "down" state are marked as gray (RBD are marked as dark gray). Site A is highlighted in yellow, site B is highlighted in magenta, site C is highlighted in blue, site D is highlighted in red, site E is highlighted in orange, site F is highlighted in pink.
-
RBM, the core region of the RBD, is the binding site of hACE2 (Shang et al. 2020). The X-ray crystal structure and Cryo-EM structure reported by recent studies have shown that antibodies CB6 (Shi et al. 2020), P2B-2F6 (Ju et al. 2020), BD23 (Cao et al. 2020), B38 (Wu et al. 2020b), 2–4 (Liu LH et al. 2020a) and S2E12 (Poh et al. 2020) are all targeted to this region. These epitopes are clustered at the top of the S protein. When the S protein is in the "down" configuration, these epitopes are partially covered by the adjacent RBD. When the S protein is in the "up" configuration, they are completely liberated. This explains why hACE2 and some antibodies can only bind to the "up" conformation. To better describe these binding epitopes, we denominate this region site A (residues 420–426 and 431–502) (Fig. 1). The buried RBD surface area of CB6 is 1088 Å and both the VH and VL of this antibody are heavily involved in antigen binding, explaining its superior affinity. P2B-2F6 binds to the RBD with a buried surface of 626 Å, involving 14 heavy- and three light-chain residues. The epitope residues include K444, G446, G447, N448, Y449, N450, L452, V483, E484, G485, F490, and S494. However, the only overlapping residues recognized by P2B-2F6 and hACE2 are G446 and Y449, largely due to angle differences as they approach the RBD. This also explains why P2B-2F6 can bind to both "up" and "down" conformations without clashing with the other S protomer. According to the results of 3D reconstruction by Cryo-EM, a single BD-23Fab is observed per S trimer with an "up" conformation, and only the VH of BD-23 is involved in binding to the RBD. Particularly, the binding is also facilitated by an N-linked glycan on N165 of adjacent protomer NTD. Just like CB6, both the VH and VL of B38 participate in antigen combination and their buried surface area is 713.9 Å and 497.7 Å, respectively. This wide range of binding patterns predicts their stronger binding and neutralizing activity. Although the RBD-binding epitopes of these antibodies are slightly different, they can all block binding to ACE2 by occupying the RBM, even if at different degrees. However, because of the variability of this region between SARS-CoV-2 and other coronaviruses, antibodies that bind to site A are generally not cross-active. Undeniably, the main problem for these antibodies is how to avoid virus escape.
Another major antigenic site is shared by both the "up" and "down" conformations, which we named site B (residues 326–366) (Fig. 1); it is the target of antibody S309 (Pinto et al. 2020). Noticeably, S309 binds better to the "down" conformation than to the "up" conformation, possibly because the flexibility of the "up" conformation of the S trimer affects its binding. Site B harbors a conservative region in the RBD (residues 333–362) as well as a small overlap of RBM regions (residues 440–444) between SARS-CoV and SARS-CoV-2, which may account for the similar affinity of S309 for these two viruses.
Unlike S309, the epitope of CR3022, a neutralizing antibody obtained from a SARS convalescent patient and cross-reacting with SARS-CoV-2 S protein, is only accessible when at least two of the RBDs in the S trimer are in the "up" conformation (Yuan et al. 2020b). This specific epitope is located on the S trimeric interface, consisting of three segments in the primary sequence (residues 369–392, 427–430, and 516–520). H014 also only binds to the RBD in the "up" conformation. The epitope residues include S383, T387, D405, V407, A411, P412, V503, and Y508 (Lv et al. 2020). These two epitopes are partially overlapped and both are located on the inner region of RBD. This inner region is not exposed in the "down" conformation. In this case, we call this region site C (residues 367–419, 427–430, and 503–520) (Fig. 1). It is interesting to note that, even when one RBD in the SARS-CoV-2 S trimer is in the "up" conformation, the binding of CR3022 to the RBD can still be sterically hindered. According to the structural model analysis, steric hindrance is resolved if the targeted RBD is slightly rotated in the "double-up" conformation. Though site C is a cross-reactive site, CR3022 likely binds more tightly to SARS-CoV because its epitope harbors a glycan absent in SARS-CoV-2. Compared with CR3022, the binding conditions of H014 are relatively loose: H014 can be combined with the S trimer in the "one-up", "double-up" or even "three-up" conformations. Moreover, other antibodies such as n3088 and n3130 were also found to neutralize SARS-CoV-2 by targeting site C (Wu et al. 2020a, b ). These small-sized single-domain antibodies fit in the lateral space between two adjacent RBD protomers and this property renders them a stronger neutralization ability than CR3022.
Some antibodies target quaternary epitopes on the top of the S protein that included elements of the RBDs from two adjacent S1 protomers, such as S2M11 (Poh et al. 2020) and 2–43 (Liu LH et al. 2020). Specifically, CDRH1, CDRH2 and the HCFR3 of S2M11 are docked into the RBM crevice whereas CDRH3 spans the interface between the RBM and helices (residues 339–343, 367–374) as well as residue 436 of an adjacent RBD. Cryo-EM reconstruction of the 2–43 Fab in complex with the S trimer at 5.8 Å resolution reveals a quaternary epitope involving RBD from one subunit and another RBD from the next. This particular combination also results in 2–43 being unable to bind to a single RBD.
Some late studies have shown that binding epitopes of neutralizing antibodies also exist in other regions besides the RBD. Chi et al. reported the 4A8 neutralizing antibody binds to the NTD region of the S protein (Chi et al. 2020). Structural analysis shows that the variable region of 4A8 interacts with its two flexible fragments (residues G140–R156 and R246–W258), thus stabilizing the structure. The interaction interface, which we named site D, is composed of a network of extensive hydrophilic interactions (Fig. 1). In another study, Liu LH et al. (2020) reported three other NTD binding antibodies, 2–17, 5–24 and 4–8. However, the molecular motion prevented visualization of the interaction at high resolution.
Poh et al. identified two immunodominant linear B cell epitopes, S14P5 and S21P2, on the SARS-CoV-2 S glycoprotein. Peptide S14P5 is located close to the RBD (residues T562-A579) and peptide S21P2 partially overlaps with an epitope identified in a recent publication (Poh et al. 2020) and contains a part of the fusion peptide sequence (residues P818–V835). Sites containing these newly discovered epitopes are denominated site E and site F, respectively (Fig. 1).
So far, most effective neutralizing antibodies are binding to RBD, especially site A. According to the existing research results, antibodies targeting site A apparently have the best antiviral activity. These antibodies not only have a stronger affinity, they are also the key to blocking receptor binding. Therefore, we believe that the vaccine design should pay more attention to the effectiveness of RBD based constructions.
We have summarized the recently discovered antigen-binding epitopes on S protein and proposed 6 sites (A–F), which can help us understand the diversity of antibody recognition. However, due to the conformational change of the S protein, particularly the "up" or "down" states of RBD, the way antibodies recognize the S protein is more complicated than we now realize. Their recognition is dynamic, requiring both spatial and temporal moment to exert their binding and neutralizing activities. We are still in the early phase of understanding the antibody response to SARS-CoV-2. There is still much work to be done, especially the potential for antibodies from the first wave to protect against the next.
-
Numbers of clinical trials for COVID-19 are underway. Including drugs exploration, such as chloroquine, hydroxychloroquine, ribavirin, remdesivir, etc (Sanders et al. 2020). Meanwhile, research teams from companies and universities around the world are developing more than 110 vaccines against SARS-CoV-2 based on different technologies (Thanh Le et al. 2020). The development of vaccines and drugs must have an in-depth understanding of the key SARS-CoV-2 proteins and their interactions with host factors.
Compared to drugs and vaccines, antibodies are a more direct and effective therapy during the outbreak and can be applied prophylactically. Recently, the development of antibodies for COVID-19 has entered a phase of rapid development and, hopefully, it will be possible to apply them in clinical practice soon.
At the same time, we should pay more attention to the occurrence of "antibody-dependent enhancement" (ADE) (Bloch et al. 2020). ADE is a well-recognized phenomenon whereby viral entry into cells is enhanced by virus/antibody complex binding to membrane Fc receptors or complement receptors, resulting in a worsening of the disease (Yager 2020). As several COVID-19 vaccine candidates enter human trials and the therapeutic use of convalescent plasma is explored, the debate continues amongst clinicians over whether ADE of SARS-CoV-2 infection represents a relevant concern. Antibodies can engage the host immune system through their constant domains that bind to Fc gamma receptors on host immune cells. These interactions can enhance immunity and help clear the pathogen or infected cells, but they can also lead to disease exacerbation in coronavirus infections. This problem would impede vaccine development but would not interfere with the clinical use of potent neutralizing antibodies that can be modified to prevent Fc gamma receptor interactions and thus preserve their protection against viral pathogens (Robbiani et al. 2020).
On the other hand, current genomic survey data indicate that single-nucleotide variations (SNVs) of SARS-CoV-2 are abundant. In a recent study, a team from Zhejiang University reported the functional characteristics of 11 viral isolates from COVID-19 patients, all with at least one mutation (Yao et al. 2020). This study demonstrated the functional potential of mutations currently occurring in the SARS-CoV-2 genome for affecting viral pathogenicity. Indeed, several articles reported that the mutation of aspartic acid to glycine at site 614 (D614G) of the S protein resulted in increased infectivity of SARS-CoV-2 (Daniloski et al. 2020; Korber et al. 2020; Zhang et al. 2020) and increased viral load in patients and also suggested that the prevalence of the G614 virus is related to increased mortality (Becerra-Flores and Cardozo 2020). Furthermore, in vitro studies showed that the infectivity of D614G-mutant pseudovirus on hACE2-293T and human lung epithelial cells is ~ nine- and ~ eight fold higher than that of wild-type SARS-CoV-2, respectively. In addition, Daniloski et al. also found that the G614 pseudovirus has an increased ability to infect Huh7.5-ACE2, A549-ACE2, and Caco-2 cells (2.4–7.7-fold higher) (Daniloski et al. 2020). Zhang et al. further elucidated that this phenomenon is not caused by an enhanced affinity of ACE2 for the S protein but by increased stability between the S1 and S2 subunits, which elevates the number of functional S proteins on the surface of the virus, thus increasing the chances of virus-cell binding. Nonetheless, they observed that the G614 and D614 pseudoviruses are similarly susceptible to neutralizing antisera, indicating that antibody-mediated control of G614 viruses is still feasible (Zhang et al. 2020).
In conclusion, mAbs targeting SARS-CoV-2 S protein with effective neutralizing activity are the focus of therapeutic intervention for COVID-19 development. Many studies have reported the function and structure of SARS-CoV-2 neutralizing antibodies targeting the RBM region and inhibiting the association between the S protein and ACE2. However, the application of antibodies targeting the RBM region alone may induce drug-resistant mutations in the virus. Baum and colleagues demonstrated that under the therapeutic pressure of a single antibody, the virus produces multiple mutations that evade the blocking action of that antibody which even binds to the highly conserved regions (Baum et al. 2020). They also used the VSV-SARS-CoV-2 system to screen for cocktail antibodies that could effectively prevent escaped mutants of the SARS-CoV-2 because both antibodies can simultaneously non-competitively bind to different regions of RBD. Indeed, with several antibodies that bind to different epitopes working together, virus escape would be greatly reduced and the neutralization activity would be synergistically increased.
Due to the increasing severity of the pandemic worldwide, a growing number of teams are involved in the study of SARS-CoV-2 neutralizing antibodies. However, there are noticeable variations among different assays used by different laboratories. It would be hard to directly compare the potency of antibodies from different pieces of literature. This review, which summarizes typical antibodies since the outbreak, provides researchers with an overview of the field.
-
This work was supported by Grants from the National Science and Technology Major Project (Grant No. 2018ZX10733403).
-
The authors declare that they have no conflict of interest.
-
This article does not contain any studies with human or animal subjects performed by any of the authors.














 DownLoad:
DownLoad: