-
Influenza A viruses can infect a variety of animals, including poultry, human, pigs, horses, marine mam-mals, and carnivore animals [i.e. dog and cat], and they are classified into different subtypes on the basis of antigenic properties in the two surface glycop-roteins, haemagglutinin (HA) and neuraminidase (NA) (14, 21). Wild aquatic birds are the primary natural reservoir of influenza A viruses, which harbor all currently known 16 HA and 9 NA subtypes (3, 21). Pigs may also play an important role in the evolution and ecology of the influenza A virus (21). The tracheal epithelium of pigs contain both sialic acid linked to galactose by an α-2, 6 linkage (SA α 2, 6 Gal) and sialic acid linked to galactose by an α-2, 3 linkage (SA α 2, 3 Gal) receptors and can be infected with swine, human and avian viruses (9), therefore, pigs have been hypothesized to serve as an intermediate host for the adaptation of avian influenza viruses to humans or as mixing vessels for the generation of genetically reassortant viruses (21).
Currently, three predominant subtypes of influenza virus are prevalent in pig populations worldwide: H1N1, H3N2, and H1N2, and these include classical swine H1N1, avian-like H1N1, human-like H3N2, reassortant H3N2 and various genotype H1N2 viruses (1, 20). Following the 1968 Hong Kong influenza pandemic (H3N2), H3N2 virus was first isolated from swine in 1970 in Asia. Since that time, human-like H3N2 viruses have been detected frequently throug-hout Europe, Asia and North America, and these viruses continue to co-circulate with H1N1 viruses (1, 2, 20). Unlike human viruses, H3N2 swine viruses have different epizootiological patterns in different areas of world (1, 20). Since 1984, reassortant H3N2 viruses have circulated in pig population in Europe, which contain genes of human (HA and NA) and avian (PB2, PB1, PA, NP, M and NS) (1). Before 1998, classical H1N1 viruses were the exclusive cause of swine in-fluenza in North America. However, around 1997-1998, three different genotypes of H3N2 viruses began to emerge in pigs: wholly human-like, double-reassortant viruses containing genes of human and swine viruses and triple-reassortant viruses containing genes of human, swine and avian viruses, and only the triple-reassortant viruses became established in pigs (20, 25).
Reverse genetics, a term used in molecular virology, describes the generation of viruses possessing a genome derived from cloned cDNAs. Reverse genetics for influenza virus has dramatically changed our under-standing of the replication cycles of these viruses (12). In addition, this methodology has allowed genetic manipulation of viral genomes to generate new viruses, which can be used as live, attenuated vaccines or vectors to express heterologous proteins.
In 2005, we isolated one influenza H3N2 virus from pigs with respiratory disease on a farm in eastern China. Genetic analysis showed that the isolate was a triple-reassortant virus among human-like H3N2, human-like H1N1 and classical swine H1N1 viruses. To manipulate the virus genomes on the level of DNA, we established a reverse genetic system for this H3N2 isolate.
HTML
-
Two lung samples were collected from dead pigs with a severe respiratory disease in 2005. Embryonic eggs and (Madin-Darby canine kidney) MDCK cells 1μg/mL TPCK-trypsin were used for the primary isolation of influenza virus. Hemagglutinin inhibition (HI) and neuraminidase inhibition (NI) assays were used for virus typing. Virus-containing allantoic fluid was stored at –80 ℃.
The plasmid pHW2000 containing human pol Ⅰ and pol Ⅱ promoter were obtained from the repository of St. Jude Children's Research Hospital.
-
RT-PCR was performed with segment-specific primers as previously described by Hoffmann et al (7).Viral RNAs were extracted from 200μL infectious allantoic fluid with Trizol LS reagent (Invitrogen) following the instruction of the manufacturer. The RNA was transcribed into cDNA by using AMV reverse transcriptase (Promega, Madison, WI) accor-ding to the protocol with Uni12 primer (Uni12 primer, 5`-AGCAAAAGCAGG-3`) in 40μL. PCR was per-formed with a set of primers specific for influenza A virus genomic fragments. The first cycle of the amplification consisted of a 4-min period at 94 ℃ and was followed by 35 cycles with the following conditions: 94 ℃ for 20 sec, 58 ℃ for 30 sec, and 72 ℃ for 3 min. PCR products were purified with the TaKaRa agarose gel DNA purification kit (TaKaRa). Sequencing was performed by Shanghai Sangon Biological Engineering Technology & Services Co., Ltd (Shanghai, China). Sequences were compiled with the Lasergene sequence analysis software package (DNAStar, Madison, WI, USA).
-
The PCR products from pMD18-T vector con-taining cDNA of the isolate were digested with Bsm B Ⅰ or Bsa Ⅰ, and then cloned into the vector pHW2000. Sequencing was performed to confirm that there were no error mutations.
-
The rescue of isolate from cDNA were generated by DNA transfection as described previously (6). Briefly, 293T human embryonic kidney and MadinDarby canine kidney (MDCK) cells were cultured and used for the transfection experiments. MDCK cells were maintained in MEM containing 10% FBS. 293T human embryonic kidney cells were cultured in Opti-MEM (Gibco) containing 5% FBS. For the trans-fection experiments, 293T cell suspension was seeded into a six-well plate (3 mL each well). The cultured 293T cells (1×106 cells per well each) were used for the transfection experiments. Lipofectamine 2000 (Invitrogen) was used according to the manufacturer's instructions to transfect the cells. Briefly, 293T cells were transfected with a DNA-lipid complex con-taining 1 μg of each plasmid in a final volume of 1 mL. Six hours later, the DNA-transfection mixture was replaced by Opti-MEM. The cells were incubated for an additional 24 h, and then, 200 μL of supernatant was taken and inoculated into 10-day-old embr-yonated chicken eggs or MDCK cells. The allantoic fluid of this first egg passage stock was analyzed by HA assay and RT-PCR. Each viral segment was partially sequenced to confirm the identity of the recombinant virus.
Virus isolation and identification
RT-PCR Amplification and Genomic Sequencing
Construction of expressing plasmids of the cDNAs of the isolate
Cell culture and co-transfection of eight plasmids
-
One virus isolate with hemagglutination activity was isolated. The isolate was determined as H3N2 subtype by HI and NI assays and designated as A/-swine/Shandongi3/2005 (Sw/SD/3/2005). RT-PCR was performed with segment-specific primers to amply 8 gene segments of the isolate (Fig 1).The nucleotide sequences of the isolate have been deposited in the GenBank database under accession numbers EU-116038-EU116046. The BLAST and phylogenetic analysis of Sw/SD/3/2005 genomic sequence demon-strated that the HA, NA, NP, M, PB1 and PA genes shared the highest sequence similarity with those of A/Moscow/10/99 or A/Hong Kong/1144/99 (99%), and the PB2 gene shared the highest homology with that of A/Puert Rico/8/34 (99%), while the NS gene share the highest homology with that of A/Swine/-Shanghai/1/2005 (Table 1). These results showed that A/swine/Shandong/3/2005 was a reassortant virus from human-like H3N2, human-like H1N1 and classical swine H1N1 viruses.
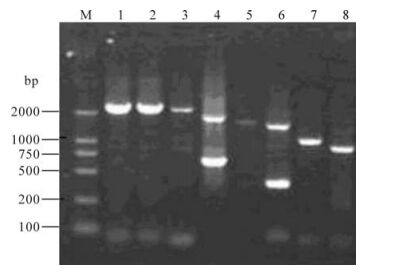
Figure 1. Full length amplification of all eight segments of Sw/SD/3/2005 by RT-PCR. M, DNA ladder; 1, PB2; 2, PB1; 3, PA; 4, HA; 5, NP; 6, NA; 7, M; 8, NS.

Table 1. Genetic similarity between Sw/ZJ/1/2004 and reference strains available in GenBank
HA plays an important role in determining tissue tropism, systemic spread, and pathogenicity of in-fluenza A viruses (14, 21). The full-length sequence of HA gene contain 1 778 nucleotides, and have a coding sequence with 1 707 nt. The deduced amino acid sequences from HA gene have 567 amino acid residues, among which there are a 16 aa signal peptide sequence, a 328 aa HA1 sequence, and a 321 aa HA2 sequence. There is an amino acid motif PEKKQTR↓G at their HA cleavage sites. Glycosylation of viral antigens masks and unmasks antigenic sites and there-fore is an important process in the generation of new viruses (11, 17). Ten potential N-linked glycosylation sites were found in the HA protein, nine of which are located in HA1. The amino acid sequence of Sw/SH/ 3/2005 at residues 226 to 228 was Ile-Ser-Ser, a typical motif of human influenza viruses and a site known to affect receptor-binding properties, which implied that the isolate preferentially binds to SA α 2, 6 Gal of human cell receptors rather than SA α 2, 3 Gal of those of avian (11, 22).
A balance between the activity of HA in virus attachment and NA in virus release is crucial for optimal viral replication (8, 21). The entire sequence of HA gene contain 1 467 nucleotides, and has a coding sequence of 1 410 nt. The deduced amino acid sequences from the HA gene have 469 amino acid residues. There are no amino acid substitutions observed at positions 274 [H→Y) or 294 [N→S) which were reported to confer resistance to oseltamivir in clinical influenza isolates (5) in the NA gene of the Sw/SD/3/ 2005 isolate, suggesting that it is sensitive to NA inhibitors. Neither substitution is observed at position 31 [H→Y) of the M2 protein which were reported to confer resistance to anti-influenza drugs Amantadine and rimantadine in the recent Europe swine viruses, including H1N1, H1N2 and H3N2 viruses (10), in isolate Sw/SD/3/2005, suggesting that it is sensitive to this class of antiviral drugs.
-
The PCR products from the pMD18-T vector con-taining cDNA of Sw/SD/3/2005 were digested with Bsm B Ⅰ or Bsa Ⅰ, and cloned into the vector pHW2000. Eight expression plasmids were constructed containing the eight genomic fragments of Sw/SD/3/2005 (H3N2), designated pHWsd-PB2, pHWsd-PB1, pHWsd-PA, pHWsd-HA, pHWsd-NP, pHWsd-NA, pHWsd-M, and pHWsd-NS, respectively.
-
The rescued viruses were inoculated into 10-day-old embryonated chicken eggs or MDCK cells. The virus-induced cytopathic effect was observed in MDCK cells (Fig. 2). The virus titer was 1:8 by HA assay in the allantoic fluid. The sequences of HA and NA genes of the rescued virus showed the HA and NA genes were identical to those of Sw/SD/3/2005, demonstrating that the rescues by our constructed eight-plasmid transfection system was successful.

Figure 2. virus-induced cytopathic effect by rescued Sw/SD/3/2005 at post-incubation 48h. A: Mock-infected MDCK cell. B: Post-incubation 48h with rescued Sw/SD/3/2005.
The reverse genetics derived R-swsd3/05 was characterized by its growth potential in chicken embryos and in tissue culture (Table 2). The results showed that R-swsd3/05 was stable after passage 3.

Table 2. Comparison of the hemagglutiation unit of the R-swsd3/05 in chicken embryo and MDCK cell
Virus isolate and genetic characterization
Establishment of expressing plasmids
Recovery of Sw/SD/3/2005 by cotransfecting eight plasmids
-
Influenza virus infection in pigs was first described in 1918 in China, and the time coincided with so-called the Spanish pandemic in humans (1). In China, it was documented that four subtypes (H1N1, H1N2, H3N1 and H3N2] of swine influenza viruses were present in pig population (4, 15, 16, 19, 23). Early studies on influenza viruses from 1976 to1982 revealed co-circulation of swine H1N1 and human-like H3N2 viruses in pigs in southern China, and three reassortant H3N2 viruses containing the surface genes HA and NA from human virus origin and the internal genes from swine H1N1virus were detected (19). During 1993-1994, avian-like H1N1 viruses firstly were detected in pigs in Southern China (4), which were co-circulating with classical H1N1 viruses previ-ously detected (18, 19). In 2004, we isolated three reassortant H1N2 viruses that contained the NA gene of human-like H3N2 virus origin and seven other genes from a classical H1N1 virus (16). Recently, inters-pecies transmission of human H1N1, avian H5N1 and H9N2 to pigs have been reported in China (15, 24, 26). Since 2003, high pathogenic avian influenza (HIPV) H5N1 viruses have become endemic in poultry in China, and pose serious threat to poultry, mammalian animals and human health. Since March 2003, we have carrying out influenza A virus (especially H5N1) surveillance in pig populations in China, and our results also showed that avian H5N1 and H9N2 could casually infect pigs resulting in severe respiratory disease (data not shown). In this study, we failed to detect avian H5N1 or H9N2 viruses on the farm where Sw/SD/3/2005 was recovered, although retrospective epidemiology survey showed that chicken death was observed in some neighboring backyard farms at that time.
Genetic reassortment may be a main mechanism to generate new pandemic strains, which is exemplified by the pandemic influenza viruses of 1957 (H2N2) and 1968 (H3N2) derived from reassortments of avian viruses and with the prevailing human viruses (21). Interestingly, the 1957 and 1968 human pandemic viruses also contained genes of avian viruses (23). These data showed that reassortant viruses containing genes of avian origin might have a selective advantage in mammals. Although no reassortant viruses derived from avian, swine, and/or human have yet been found in China, there is evidence suggesting that avian H1, H9, and H5 had been transmitted to pigs in southeast-ern China (15, 26, 28). Because of susceptibility to both avian and human viruses, pigs may serve as ideal intermediate hosts for the generation of potential pandemic influenza viruses by reassortment (1, 9, 21). Southeastern China has been regarded as an epicenter for the emergence of pandemic influenza viruses, where a variety of influenza viruses co-circulate in people, pigs and poultry (14, 18). Introduction of avian viruses (H1N1, H5N1 or H9N2) into pigs and co-infection with human viruses (H3N2 or H1N1) or swine viruses (H1N1 or H1N2) provide a favorable opportunity for the generation of reassortants, which pose a significant pandemic threat. In this study, we detected a triple-reassortant virus which was not found before. This finding should raise concerns about the genetic evolution of influenza virus in pigs and rein-force influenza virus surveillance in pigs in China.
The reverse genetics system for the influenza virus was established in 1999 by two different research groups (6, 13), relying on the cellular enzyme RNA polymerase I for the synthesis of influenza viral RNAs. A RNA polymerase Ⅰ/Ⅱ system with only 8 plasmids was reported by Hoffmann et al (6), and in this system, the synthesis of vRNA and mRNA was achieved from one viral cDNA template, circumventing the need for protein expression constructs. The reverse genetic system of 8 plasmids is a powerful tool to determine the structure-function relationships, pathogenicity, and host cell tropism. Furthermore, the 8 plasmid system also can be applied to the rational development of influenza virus high growth reassortants. In this study, we successfully established the 8 plasmid reverse genetic system of Sw/SD/3/2005, which provides a useful platform to further address the molecular patho-genicity and development of a candidate vaccine.













 DownLoad:
DownLoad: