-
Porcine reproductive and respiratory syndrome virus (PRRSV) is a member of the family Arteriviridae (1) and is characterized by respiratory disease in young pigs and severe reproductive failure in sows, including abortion, stillbirths and weak piglets. PRRS has caused immense economic losses in the pig industry and is considered to be one of the most important infectious diseases of pigs in the world (2). PRRSV has been recognized as one of the most important pathogens of pigs throughout the world. The virus was first confirmed in China in 1996 and since then the virus has become widespread all over the country (3). The PRRSV genome is a singlestranded, nonsegmented, positive-sense RNA, about 15.5 kb in length. The viral genomic RNA is polycistronic, comprising nine open-reading frames (ORFs) (4). Two large ORFs (ORF1a/ORF1b), occupying the 5`-terminal three-fourths of the genome, encode the viral replicase polyprotein, which is predicted to be autoproteolytically cleaved at 12 sites, ultimately producing 13 non-structural proteins (5), while the seven ORFs located at the 3`-terminus of the genome are postulated to encode viral structural proteins.
Among the nonstructural proteins, the multidomain protein nsp2 is the largest PRRSV replicative protein. Genetic analysis of both Coronaviridae and Arteriviridae (1) originally identified the protein to be spanning amino acids (aa) 384 to 1 363 (nsp2 aa 1 to 981); this was subsequently projected to comprise aa 384 to 1 578 (nsp2 aa 1 to 1 196) of strain VR-2332 ORF1a. Three major domains could be determined through the alignment of arterivirus nsp2 proteins: an N-terminal cysteine proteinase domain (PL2), a functionally unspecified middle region, and a hydrophobic transmembrane (TM) region near the C terminus (8, 9). However, the exact C-terminal cleavage sites have yet to be determined. The function of PRRSV nsp2 in the virus life cycle is currently not known, while Equine arteritis virus (EAV) nsp2 has been shown to be involved in the generation of double-membrane vesicles together with nsp3 and to serve as a cofactor for the nsp4 protein (10). Studies have identified the presence of a 51-nt deletion in the nsp2 gene of NA Type 1 viruses, which is located between nt 2 199 and 2 250 in ORF1a of the Lelystad virus genome (6, 7). Eighteen of the 20 NA-type 1 viruses possess the same 51-nt deletion (6).
Nucleotide and putative amino acid sequence analysis have revealed that Nsp2 is the most variable region of PRRSV (11, 12). Genetic variability, including point mutations and the insertion or deletion of nucleotides, has been demonstrated in the Nsp2 gene, not only between North American and European genotypes, but also among viruses within the same genotype (6). However, the biological and clinical significance of Nsp2 diversity remains to be elucidated. The aims of this study were: (ⅰ) Establish a genetic database for the nsp2 gene of PRRSV isolated from Ningxia Hui Nationality Autonomous Region (Ningxia) of China, (ⅱ) Identify relationships between sample date and the evolution of PRRSV in China, (ⅲ) Determine the phylogenetic relationship between the Ningxia strains and those from other regions of China.
HTML
-
The seventeen field clinical samples from pigs with reproductive failure (comprising three pigs from Zhongwei (ZHW), Heying (HY), Shizuishan (SZSH), Wuzhong (WZH) and Yingchuan (YCH) counties respectively, and two pigs from Dabao Country (DB)) in the Ningxia region of China in 2007, and reference strain of PRRSV CH-1a (GenBank access number: AY032626) used in this study were obtained from virology department of Lanzhou Veterinary Research Institute, Gansu of P.R. China. The sense primer (S1) was 5'-TGGGCGGTTGAGCAGGTC-3' and the reverse primer (R1) was 5'-GCAAGGCCCGCCAGAGTCG-3'. This generated the complete region of nsp2 gene. Each sequence of the PCR products was determined by both enzyme analysis and sequencing.
-
The viral RNA of all clinical samples were extracted using a QIAamp Viral RNA Mini Kit (Qiagen, Germany), following the manufacturer's instructions. In brief, after lysis of the specimens, the mixture was applied to a spin column as described by the manufacturer's protocol. Reverse transcription was performed at 42℃ for 1.5 h with 5μL total RNA, and then PCR was carried out in a reaction volume of 50 μL containing 1μL of each primer (S1 and R1, 50 pmol/μL), 5 μL of 10×PCR Buffer, 4 μL 2.5 mmol/L dNTPs, 3 μL of 25 mmol/L MgCl2, 1μL of RNase Inhibitor (40U/μL), 0.5 μL of Taq (5U/μL), 10 μL of reverse transcription production and 24.5 μL ddH2O. Amplification conditions were 94℃ for 4 min, then 30 cycles that each consisted of denaturing at 94℃ 1min, annealing at 57℃ 40 s, and extension at 72℃ for 1 min, the sample was then heated at 72℃ for 10 min, and a final stage of 4℃ for 5 min. The amplified PCR products were analyzed by agarose gel electrophoresis and purified using the QIAquick gel extraction kit (Qiagen) per the manufacturer's protocol. All nucleotide sequences were obtained from clinical sample RNA by direct sequencing of PCR products.
-
Sequence data were analyzed using the method described by Chen et al (13). Briefly, the amplified sequences of the nsp2 gene were translated and compared with ch-1a, SD2006 and SD (3) sequences from mainland China available in GenBank. Multiple sequence alignment was carried out using the sequence analysis software Lasergene1 (DNASTAR Inc., Madison, WI).
Virus strain and the primers
RNA extracted and RT-PCR
Sequence analysis
-
Partial Ningxia PRRSV field isolates were used for RT-PCR analysis, and the various patterns obtained are shown in Fig. 1. The isolated PRRSV nsp2 product length wasabout 630bp, consistent with the expected length. The length of the Classical strain (ch-1a) PCR products was about 710bp. This result indicated that compared to ch-1a strain, all of the isolates possessed the same deletion in the nsp2 gene.
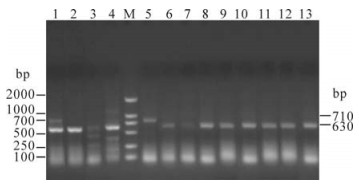
Figure 1. RT-PCR results for partial PRRSV isolates of the nsp2 gene. 1-4, The products of nsp2 gene for ZHW, ZHW1, SHZSH and HY respectively. M: molecular size marker (DL2000); 5, products of nsp2 gene for ch-1a. 6-13, The products of nsp2 gene for HY1, YCH, YCH1, WZH, WZH1, DB1, DB2 and HY1, respectively. All the products were electrophoresed on a 1.2% agarose gels.
-
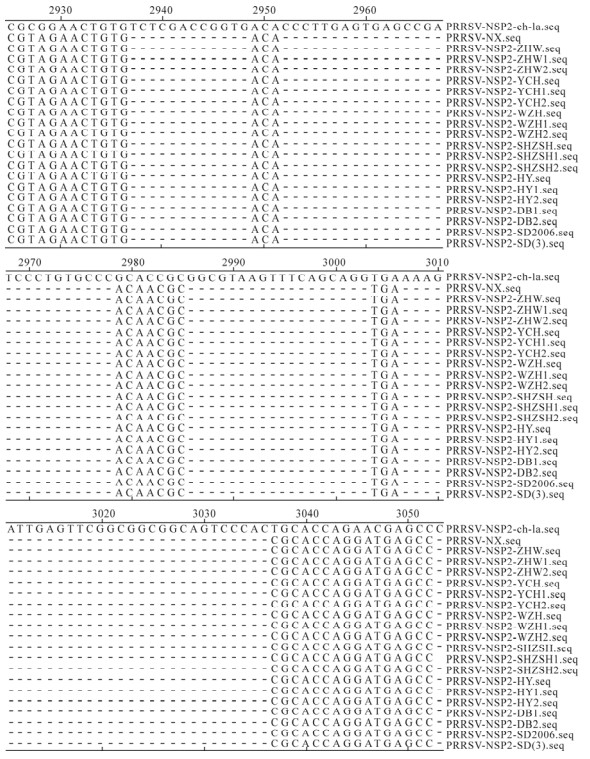
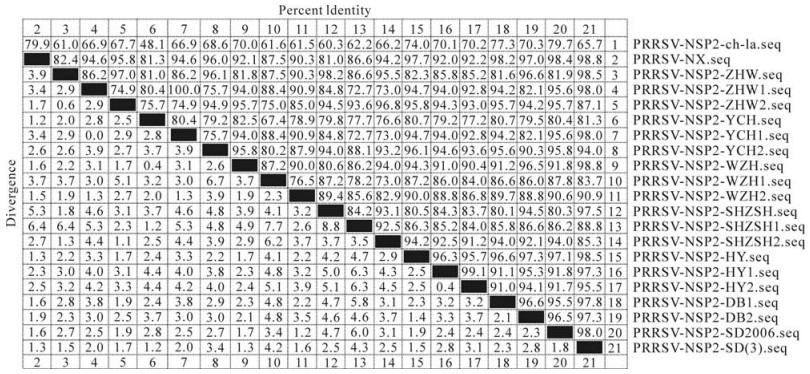
Relative to strain ch-1a, the nsp2 genes of seventeen isolates have 87 nucleotides deleted at positions 2 937 to 2 948, 2 952 to 2 978, 2 986 to 3 003 and 3 007 to 3 036, respectively; these deletions are the same as those for the SD (3) and SD2006 strains (Fig. 2). Sequence analysis indicated that homology between Ningxia and ch-1a was 60.3%-79.9% in nucleotide sequence, and homology between Ningxia and the common SD strain was 80.3%-98.8% in nucleotide sequence. The nsp2 genes of seventeen isolates had 74.9%-100% nucleotide sequence identity with each other (Fig. 3).
PCR product analyses
Sequencing and phylogenetic analysis of Nsp2 genes
-
PRRSV has been recognized as one of the most important pathogens of pigs throughout the world and the virus has become widespread in China since it was first confirmed in 1996. Even though several PRRSV vaccines are available, control of the disease remain difficult (14). In this study, 8 pig farms in the Ningxia were diagnosed with PRRSV infection on the basis of clinical signs and RT-PCR. Seventeen field isolates of PRRSV obtained from 8 farms with severe disease were identified as North American type PRRSV on the basis of sequence analysis (in press). To our knowledge, this is the first report of the emergence of North American type PRRSV strain in herds of pigs in the Ningxia Hui National Autonomous Region of China.
The ORF1a gene of PRRSV encodes a single poly-protein, predicted to be cleaved at eight sites to form nine Nsps (15). Nsp2 contains conserved catalytic cysteine and histidine residues at the amino terminus and is predicted to encode a cysteine protease (16). Amino acid deletions in Nsp2 have been identified in several PRRSV isolates (3). The areas in Nsp2 with deletions and amino acid hypervariability are predicted to be immunologically important (1); Nsp2 might function as an immunodominant protein under selective pressure by the host immune system (17, 18). In our study, we find that the nsp2 genes of seventeen isolates have an 87 nucleotides deletion at positions 2 937 to 2 948, 2 952 to 2 978, 2 986 to 3 003 and 3 007 to 3036, respectively, relative to strain ch-1a. This deletion site is same as the SD (3) and SD2006 strains. Sequence analysis indicates that homology between the Ningxia and ch-1a strain was 60.3%-79.9% in nucleotide sequence, and homology between the Ningxia and SD strains was 80.3%-98.8% in nucleotide sequence. The nsp2 genes of seventeen isolates had 74.9%-100% nucleotide sequence identities with each other. It was also indicated that Ningxia PRRSV may originated from a popular strain in 2007, but the association between pathogenicity and variation of Nsp2 in Ningxia and the other popular strain should be studied further.
All of those results demonstrated that PRRSV variation strain exists in Ningxia. Detailed analysis of these virus strains can provide data for understanding the epidemiology of PRRSV, and at the same time provide information for preservation and manipulation of the disease. Finally identification of the characteristics of the Ningxia Hui Nsp2 sequences may offer important insight into the properties of PRRSV in this region.













 DownLoad:
DownLoad: