HTML
-
Hepatitis B virus (HBV), is a species of the Orthohepadnavirus genus in the Hepadnaviridae family, which replicates via a unique protein-primed reverse transcription mechanism (Nassal, 2008). The pregenome RNA (pgRNA) of HBV is packed into a core capsid and is reverse transcribed into a 3.2 kb relaxed circular DNA (RC-DNA) by P protein, which functions as the viral polymerase and reverse transcriptase (Beck and Nassal, 2007). The mature virion containing RC-DNA can only infect hepatocytes because of its hepatotropism (Baumert et al., 2007). After it is released into the nucleoplasm, RC-DNA is "repaired" and transformed into a plasmid-like covalently closed circular DNA (cccDNA) by cellular factors and enzymes (Urban et al., 2010). Nuclear cccDNA molecules are organized into a chromatin-like structure as viral minichromosomes that synthesizes the subgenomic RNAs (preS/S mRNAs and HBx mRNA) and the greater-than-genome length RNAs (pregenome RNA and precore RNA) via cellular RNA polymerase Ⅱ (Levrero et al., 2009). Because cccDNA cannot be thoroughly eradicated by current therapeutic approaches, chronic HBV-infected hepatocytes persistently secrete Hepatitis B surface antigen (HBsAg), Hepatitis B e antigen (HBeAg), and complete virion particles (Seeger and Mason, 2000). Interferon (IFN) α and nucleos(t)ide analogues have been broadly applied clinically but they also have limitations such as severe side effect and the potential to induce drug resistant strains, which make it impossible to completely eradicate HBV in patients with chronic hepatitis B (CHB) (Yuen and Lai, 2001). Therefore, novel effective therapeutic agents against HBV are urgently required.
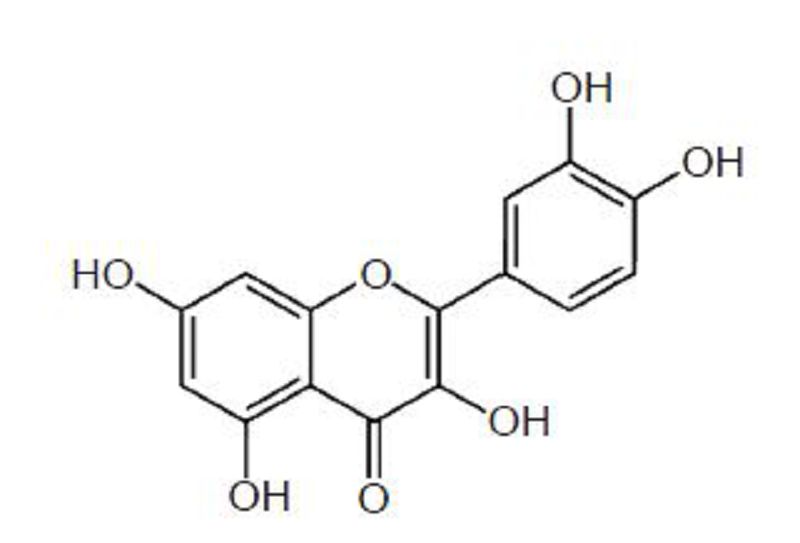
Quercetin (3, 3', 4', 5, 7-pentahydroxyflavanone and its analog 3, 3', 4', 5, 7-pentahydroxy-2-phenylchromen-4-one, Figure 1) is categorized as a flavonol, one of the six subclasses of flavonoid compounds (Kelly, 2011). Quercetin-type flavonols are widely distributed in plants, and the most abundant forms are quercetin glycosides, which consist of quercetin linked with one or two glucose molecules (Hakkinen et al., 1999). Quercetin has shown many potential benefits to human health such as blood pressure control (Larson et al., 2012), antioxidant, anticancer, antidiabetic, anti-infective, and anti-inflammatory (Kelly, 2011). Besides these beneficial effects on human health, quercetin has also exhibited in vitro antiviral activity against some RNA viruses including the human immunodeficiency virus (HIV), poliovirus type 1, parainfluenza virus type 3, respiratory syncytial virus (Kaul et al., 1985; Kelly, 2011), and hepatitis C (Gonzalez et al., 2009). In murine studies, quercetin reportedly protected the lungs from oxidative stress induced by the influenza virus (Kumar et al., 2003), and reduce susceptibility to influenza infection following stressful exercise (Davis et al., 2008).
However, the effect of quercetin on HBV remains unclear. Therefore, in the present study, we evaluated the potential anti-HBV activity of quercetin using two models including the HepG2.2.15 cells, as well as HuH-7 cells transiently transfected with pCH-9/3091 to mimic HBV transcription, protein synthesis, and DNA replication under physiological condition. HepG2.2.15 was derived by stably transfecting HepG2 cells with pDolTHBV-1, a construct carrying four copies of HBV genome (Sells et al., 1987). pCH-9/3091 contains a 1.1× HBV genome driven by the human cytomegalovirus immediate early I protein promoter (Nassal, 1992). The cells were treated with quercetin at varying concentration and time gradients, and then viral antigen secretion as well as viral DNA, transcripts, and heat shock protein levels were analyzed in both models using enzyme-linked immunosorbent assay (ELISA) and quantitative real-time polymerase chain reaction (qPCR). Possible synergistic effects of quercetin and lamivudine (3TC), entecavir (ETV), or adefovir (Ade) against viral replication were also explored.
-
The human hepatoma HepG2.2.15 and HuH-7 cell lines were obtained from the China Center for Type Culture Collection (CCTCC). Cells were cultured in Dulbecco's modified Eagle's Medium (DMEM) supplemented with 10% (v/v) fetal bovine serum (FBS) and antibiotics (100 units/mL penicillin and 100 μg/mL streptomycin, Gibco Life Technologies, USA) at 37 ℃ in a 5% CO2 incubator.
The pCH-9/3091 construct containing 1.1 copies of the HBV genome was kindly provided by Prof. Michael Nassal (University of Freiburg) and transfected into HuH-7 cells. Briefly, HuH-7 cells at a density of 2 × 105 cells/well were seeded in 24-well plates, cultured for 24 h, and then transfected with 0.8 μg of pCH-9/3091 using Lipofectamine 2000 (Invitrogen, USA) according to the manufacturer's instructions.
-
Quercetin dihydrate was purchased from Calbiochem (German). Three clinically used anti-HBV drugs, ETV (entecavir-triphosphate, Moravek Biochemicals, USA), 3TC (lamivudine, Sigma-Aldrich, USA) and Ade (adefovir, Sigma-Aldrich, USA), were used as positive controls. And all compounds were dissolved in dimethyl sulfoxide (DMSO), and then diluted into different concentration. HepG2.2.15 and HuH-7 cells were plated into 24-well plates at a density of 2 × 105 cells/well, cultured or transfected with pCH-9/3091. After incubation for 24 h, the media was gently aspirated and 1 mL of complete media with different concentration of compounds was replaced in each well. The final concentration of DMSO in each well was 0.3%. Treatment with 30 nmol/L ETV and 0.3% DMSO served as positive and negative (non-drug) controls. The treatment day was designated as day 0, and the culture media were replaced every 24 h with fresh medium containing the respective treatments. Supernatants and cells were collected at the indicated time points for further analysis.
-
The toxicity of quercetin against the cell lines was determined using the water-soluble tetrazolium ([WST]-8 [2-(2-methoxy-4-nitrophenyl)-3-(4-nitrophenyl)-5-(2, 4-disulfophenyl)-2H-tetrazolium, monosodium salt]) assay with the cell counting kit-8 (CCK-8, Dojindo Labora-tories, Japan) following the manufacturer's instruction. Briefly, HepG2.2.15 and HuH-7 cells (1 × 104 cells/well) were cultured in 96-well plates, treated with quercetin (1, 10, 25, 50, 100, and 200 μmol/L) for 4 days, and the media were replaced with fresh media containing quercetin daily. On the last day, 10 μL of WST-8 solution was added to each well followed by incubation at 37 ℃ for 2 h, and then the optical density (OD) was measured at 450 nm. Each experiment was performed in triplicate and the cell viability was normalized to the control sample.
-
To quantify the HBV antigen secretion, the supernatants of drug-treated cells were collected at the indicated time points, and the HBsAg and HBeAg levels were determined using a commercial ELISA kit (Kehua Bio-engineering, China). The results were normalized to the control sample.
-
Intracellular nucleocapsid-associated viral DNA was extracted from HepG2.2.15 or transfected HuH-7 cells as described previously (Feng et al., 2013). In brief, 2 × 105 cells were lysed in 200 μL lysis buffer containing 50 mmol/L Tris-hydrochloride (HCl) pH 7.5, 140 mmol/ L sodium chloride (NaCl), and 0.5% NP-40. After centrifugation, the supernatants were incubated with 20 U DNase I (TaKaRa, Japan) and 2 μL RNase A (TaKaRa) at 37 ℃ for 2 h, to completely digest the non-encapsulated DNA and RNA. Then, 200 mmol/L proteinase K (Tiangen, China) was added to degrade the core particles. After extraction and precipitation with phenol-chloroform and ethanol, respectively, the viral DNA was dissolved in Tris-ethylenediaminetetraacetic acid (EDTA, TE) buffer. Extracellular viral DNA was extracted from 200 μL of medium using the TIANamp Virus DNA/RNA kit (Tiangen) according to the manufacturer's instructions.
Total RNA was extracted from HepG2.2.15 and transfected HuH-7 cells using TRIzol reagent (Invitrogen) and reverse transcribed to cDNA using a PrimeScript RT reagent kit (TaKaRa). The q-PCR was performed with the SYBR Premix Ex Taq Ⅱ (TaKaRa) using a LightCycler 96 Real-Time PCR system (Roche, Swizerl and). The cycling program was run at 95 ℃ for 5 min, followed by 45 cycles at 95 ℃ for 10 s, 60 ℃ for 10 s, and 72 ℃ for 10 s. The primers used are listed in Table 1.

Table 1. Primers
-
All results of this study are presented as mean ± standard deviation (SD) of at least three independent experiments. Differences were considered statistically significant when P < 0.05 and P < 0.01 using unpaired twotailed Student's t-tests.
Cells culture and transfection
Compounds and cell treatment
Cytotoxicity assay
ELISA assay of HBV antigen secretion
Quantitative real-time polymerase chain reaction (qPCR)
Statistical analysis
-
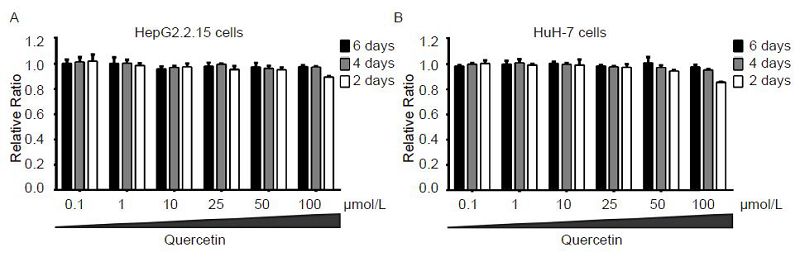
The viabilities of HepG2.2.15 and HuH-7 cells in the presence of varying concentrations (0.1, 1, 10, 25, 50, and 100 μmol/L) of quercetin were examined at different time points (2, 4, and 6 days) using the CCK-8 assay. At these tested concentrations, quercetin showed no cytotoxicity in both cells. Even at the highest concentration, treatment with quercetin for 6 days reduced HepG2.2.15 and HuH-7 cell growth by only 10.4% and 14.3% (Figure 2A, 2B), respectively. Other studies have also reported that quercetin showed a very low toxicity, and its half-maximal (50%) cytostatic concentration (CC50) was more than 100 μmol/L (Romero et al., 2005; Kelly, 2011). Therefore, based on the cytotoxicity assay, 50 μmol/L was chosen as the maximum treatment concentration of quercetin in subsequent experiments.
-
We studied the effect of varying concentrations of quercetin (1, 10, 25, and 50 μmol/L) on HBV antigen secretion by determining the levels of HBsAg and HBeAg in HepG2.2.15 or transfected HuH-7 cells. DMSO and 30 nmol/L ETV were used as controls. Quercetin-treated HepG2.2.15 cells (1 μmol/L) did not show a visible reduction in viral antigens compared to the untreated controls, whereas higher concentrations significantly inhibited both HBsAg and HBeAg secretion dose-dependently (Figure 3A). Notably, quercetin inhibited HBsAg secretion more potently than it did HBeAg secretion (Figure 3A), and 50 μmol/L caused 56.9% and 41.0% reduction of HBsAg and HBeAg, respectively. In HuH-7 cells transfected with pCH-9/3091, 1 μmol/L quercetin also failed to decrease viral antigens. A dose-dependent suppression was observed with quercetin at concentrations of 10 μmol/L and higher, and the decrease in HBsAg was greater than in HBeAg levels (Figure 3B). Compared to the untreated control, the HBsAg and HBeAg levels dropped to 59.6% and 68.5%, respectively (Figure 3B) upon following treatment with 50 μmol/L quercetin. Furthermore, HBV antigen secretion in HuH-7 cells transfected with pCH-9/3091 showed less sensitivity to quercetin treatment.
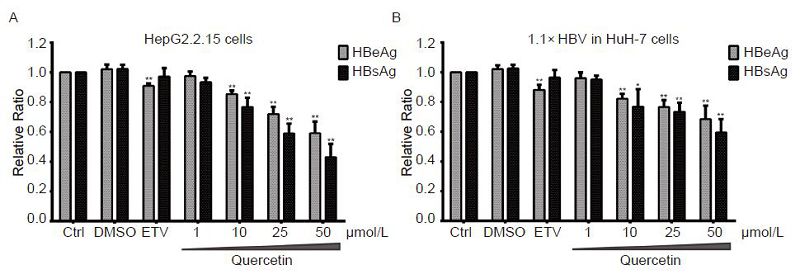
Figure 3. Concentration-dependent suppression of viral antigen secretion by quercetin. HepG2.2.15 and pCH-9/3091-transfected HuH-7 cells were treated with quercetin at indicated concentrations or 30 nmol/L entecavir (ETV). On day 4, culture supernatants were collected and assayed for HBsAg and HBeAg levels by ELISA. All data were normalized to untreated control (Ctrl); *P < 0.05, **P < 0.01.
Next, 50 μmol/L quercetin was used to assess viral antigens secretion over a time course. In both cell models, treatment with quercetin significantly inhibited HBV antigen secretions from day 3, and these effects intensified on day 4 (Figure 4A, 4B). However, no time-dependency inhibition was observed.
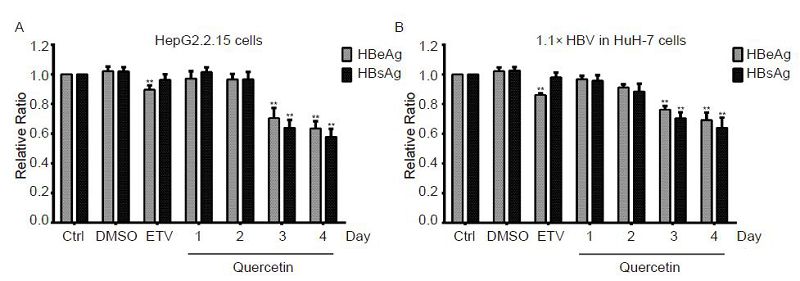
Figure 4. Time-dependent suppression of viral antigen secretion by quercetin. HepG2.2.15 and pCH-9/3091-transfected HuH-7 cells were treated with 50 μmol/L quercetin or 30 nmol/L entecavir (ETV). Culture and ETV-treated supernatants were collected daily and on day 4, respectively. HBsAg and HBeAg concentrations were measured by ELISA. All data were normalized to the untreated control (Ctrl); **P < 0.01.
-
To investigate whether quercetin influenced HBV replication, the intracellular and extracellular viral DNA was extracted from the drug-treated cells and supernatants and then analyzed using qPCR. The quercetin treatments were also performed with concentration (Figure 5) and time (Figure 6) gradients.
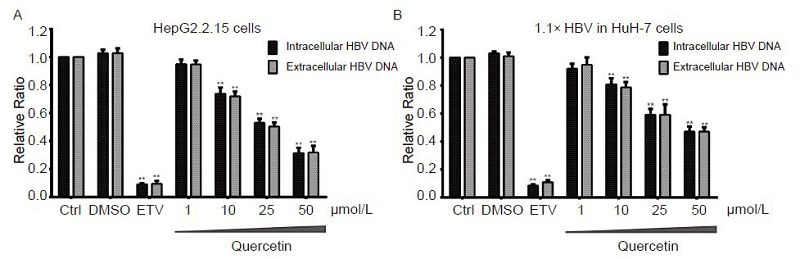
Figure 5. Concentration-dependent suppression of viral DNA level by quercetin. HepG2.2.15 and pCH-9/3091-transfected HuH-7 cells were treated with quercetin at indicated concentrations or 30 nmol/L entecavir (ETV). On day 4, intracellular and extracellular HBV DNA of samples were extracted and analyzed using qPCR. All data were normalized to untreated control (Ctrl); **P < 0.01.
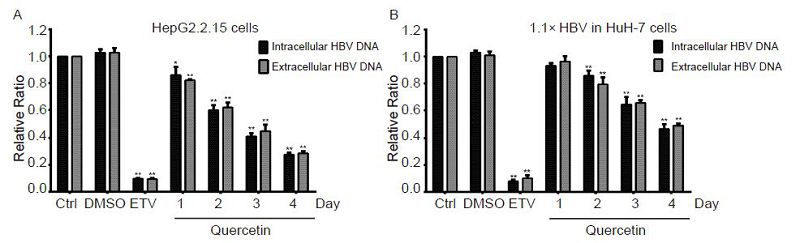
Figure 6. Time-dependent suppression of viral DNA levels by quercetin. HepG2.2.15 and pCH-9/3091-transfected HuH-7 cells were treated with 50 μmol/L quercetin or 30 nmol/L entecavir (ETV). Intracellular and extracellular HBV DNA of samples were extracted daily, and ETV-treated samples were collected on day 4. HBV genomic DNA levels were examined using qPCR. All data were normalized to the untreated control (Ctrl); *P < 0.05, **P < 0.01.
HepG2.2.15 and pCH-9/3091-transfected HuH-7 cells showed a clear dose-dependent suppression of HBV DNA levels following treatment with quercetin (Figure 5). The inhibitory effect of quercetin was observed at concentrations of 10 μmol/L and higher, and the intracellular viral DNA levels of HepG2.2.15 and HuH-7 cells transfected with pCH-9/3091 were significantly decreased by 68.5% and 52.9%, respectively (Figure 5A, 5B) upon at 50 μmol/L. The extracellular viral DNA levels were similarly and significantly inhibited by 68.1% and 52.9%, respectively (Figure 5A, 5B). The time gradient tests revealed a time-dependent decline in the HBV DNA levels following quercetin treatment (Figure 6). HepG2.2.15 and pCH-9/3091-transfected HuH-7 cells treated with 50 μmol/L quercetin showed significant inhibition of viral DNA levels from day 1 and 2, respectively. In both cell models, 30 nmol/L ETV reduced HBV DNA levels by about 90% (Figure 5, 6), which was similar to previously reported studies (Seifer et al., 1998; Bader and Korba, 2010).
To further elucidate its anti-HBV activity, quercetin was compared with clinically used nucleos(t)ide analogs, alone or in combination. The drugs including 3TC, ETV and Ade inhibited specifically the reverse transcription of HBV, resulting in a sharp drop in viral DNA but not transcription and antigen secretion levels (Yuen and Lai, 2001; Trepo et al., 2014). Therefore, intracellular HBV DNA levels were determined and compared in HepG2.2.15 cells following drug treatment. The result showed that 3TC and ETV at 70 and 9 nmol/L, respectively and Ade at 1.2 μmol/L reduced intracellular HBV DNA levels by 52.7%, 80.3%, and 86.7%, respectively (Figure 7). A combination of quercetin (50 μmol/L) and either 3TC, ETV, or Ade enhanced the inhibitory effect on viral replication and intracellular viral DNA levels were decreased further by 77.6%, 89.7%, and 89.3%, respectively (Figure 7).

Figure 7. Combination treatments of quercetin and clinically used anti-HBV drugs. HepG2.2.15 cells were treated with 50 μmol/L quercetin (Q) alone or in combination with lamivudine (3TC, 70 nmol/L), entecavir (ETV, 9 nmol/L) or adefovir (Ade, 1.2 μmol/L) for 4 days. Intracellular HBV DNA of samples were extracted on day 4. HBV genomic DNA levels were examined using qPCR. All data were normalized to untreated control (Ctrl); **P < 0.01.
-
The heat shock proteins (HSP) including HSP40, heat shock cognate protein (HSC) 70, and HSP90-β have been reported to participate in the HBV life cycle as indispensable factors in the initiation of viral reverse transcription and replication (Hu et al., 2004; Nassal, 2008). Therefore, the viral transcription levels of these heat shock proteins were analyzed in the HepG2.2.15 cell model following incubation with 50 μmol/L quercetin for 4 days. The qPCR results showed that 50 μmol/L quercetin significantly reduced the mRNA levels of HSP40, HSC70, and HSP90-β to 25.4%, 54.1%, and 22.5%, respectively (Figure 8). However, 30 nmol/L ETV did not influence the transcription levels.
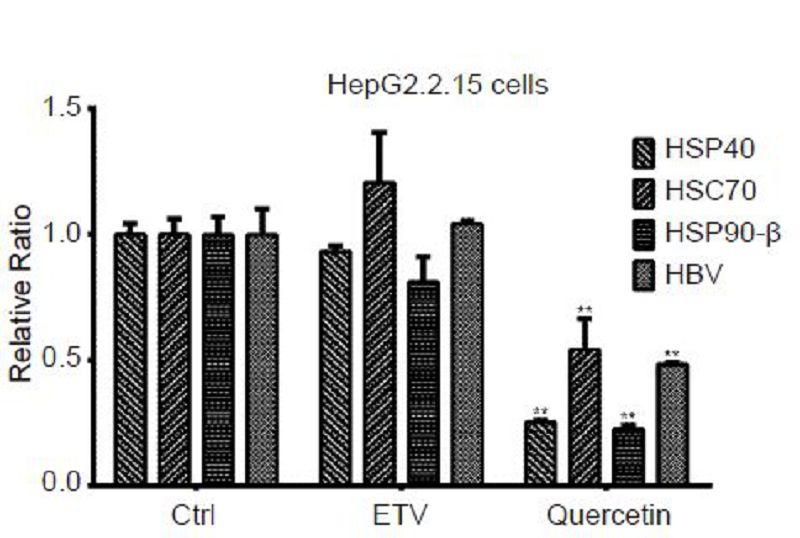
Figure 8. Influence of quercetin on heat shock proteins (HSPs) and hepatitis B virus (HBV) transcripts. HepG2.2.15 cells were treated with 50 μmol/L quercetin or 30 nmol/L entecavir (ETV) for 4 days. Total RNA of samples were extracted, reverse transcribed, and assayed for transcription levels of HSP40, HSC70, HSP90-β, and HBV using qPCR. All data were normalized to untreated control (Ctrl); **P < 0.01.
In addition, the level of HBV transcripts was also analyzed, and 50 μmol/L quercetin significantly inhibited viral transcription by 51.8% (Figure 8) while ETV showed no effect in this test. Since viral antigens are directly involved in HBV transcription, these results may explain why quercetin but not ETV reduced antigen secretion (Figure 3, 4).
Cytotoxicity of quercetin against HepG2.2.15 and HuH-7 cells
Inhibition of HBV antigen secretion by quercetin
Suppression of HBV replication by quercetin
Influence of quercetin on heat shock proteins and HBV transcripts
-
HBV infection is a major viral-induced public health problem worldwide. More than 2 billion people have been affected with HBV, and over 350 million patients have CHB (Liaw and Chu, 2009). People with HBV have an increased risk of developing hepatic decompensation, cirrhosis, and hepatocellular carcinoma (Liaw and Chu, 2009). The estimated global mortality rate is 786, 000 deaths per year (Lozano et al., 2012). The current clinically used therapeutic agents for HBV are mainly IFNα and nucleos(t)ide analogues (Yuen and Lai, 2001; Liaw and Chu, 2009; Trepo et al., 2014); however, both agents have their limitations. IFNα, an immunomodulator, acts the enhance the immune response of the host to HBV antigens expressed on the surface of the hepatocytes, thereby causing or potentiating the lysis of infected hepatocytes (Yuen and Lai, 2001; Yang et al., 2014). However, the clinical application of IFNα is limited by its poor response, high cost, and adverse effects. Presently, several nucleos(t)ide analogues including 3TC, ETV, and Ade have been approved for clinical application (Trepo et al., 2014). These oral agents selectively and potently inhibit HBV polymerase. For example, ETV blocks all three distinct phases of replication mediated by viral polymerase including priming, reverse transcription, and DNA-dependent DNA synthesis (Seifer et al., 1998). Therefore, it reduces the encapsulated HBV DNA level in host cells by 90% at a very low concentration (30 nmol/L, Figure 5, 6). However, ETV failed to eradicate the HBV cccDNA in the nuclei of infected hepatocytes, and did not influence viral transcription in cell models (Figure 8), which resulted in the secretion of high levels of HBV antigens Figure 3, 4). In addition, the viral infection relapse following discontinuation of the medication and occurrence of drug resistance during therapy limit the application of nucleos(t)ide analogs (Yuen and Lai, 2001; Liaw, 2009).
Quercetin is a plant extract derivative and, therefore, has very low toxicity. The result of the cytotoxicity assay showed that the highest dose (100 μmol/L) had no significant toxicity against HepG2.2.15 cells (Figure 2), but the solubility of quercetin in DMSO limited its assay at higher concentrations. In human studies, treatment with quercetin at up to 1000 mg/day for several months showed no adverse effects, and a low dose of 150 mg/ day significantly increased plasma concentrations and demonstrated a biological effect (Kelly, 2011). Hence, quercetin is generally well tolerated in human. Based on the various antiviral effects of quercetin mentioned above, we investigated whether quercetin influenced the replication of HBV. The ELISA (Figure 3, 4) and qPCR (Figure 5, 6) results demonstrate that quercetin indeed inhibited HBV in the both cell models used. Furthermore, quercetin effectively decreased the secretion of HBsAg and HBeAg as well as the viral DNA level, whereas ETV only suppresses the synthesis of viral DNA but not antigen secretion. The decline in HBsAg and HBeAg levels implied that quercetin may influence the viral transcription of subgenomic RNAs, and this hypothesis was supported by the qPCR results in Figure 8. One of the signal transduction pathways influenced by quercetin is the phosphoinositide 3-kinase (PI3K)-Akt (Gulati et al., 2006), which could also regulate the transcription and replication of HBV RNA and DNA, respectively (Guo et al., 2007). Therefore, PI3K might be a potential mediator of the quercetin-induced repression of HBV transcription.
The efficacy of quercetin was compared with those of known anti-HBV drugs, alone or in combination (Figure 7), and it exhibited a similar level of efficacy to 3TC. However, the concentration of quercetin used was higher than that of 3TC, indicating that the inhibition of viral reverse transcription by quercetin was not as potent as that of the known nucleos(t)ide drugs, probably due to its indirect effect on HBV. HSPs including HSP40, HSC70, and HSP90-β convert the inactive P protein of HBV transiently into an active form, which is indispensable for the initiation of viral reverse transcription and replication (Nassal, 2008). The inhibitor of HSP90, geldanamycin (GA) (Hu et al., 2004), and the inhibitor of HSC70, oxymatrine (OMTR) (Wang et al., 2010) repressed the reverse transcription and replication of HBV. In addition, quercetin inhibited the heat shock factor activity, resulting in repression of HSP gene expression (Powers and Workman, 2007) and further attenuation of hepatitis C virus production (Gonzalez et al., 2009). The results in Figure 8 showed that quercetin indeed reduced the transcriptional levels of HSP40, HSC70, and HSP90-β. Hence, the quercetin-induced inhibition of viral DNA is probably mediated by decreasing viral RNAs and host HSP. Because quercetin and the nucleos(t)ide drugs likely affected different factors in the replication cycle of HBV, their combination showed more efficacy than either treatment alone did (Figure 7). Furthermore, these results indicate the potential clinical value of quercetin as an anti-HBV agent.
In conclusion, this study demonstrated that quercetin has promising anti-HBV activity, mediated by the reduction of HBsAg and HBeAg secretion as well as viral DNA level in vitro. Furthermore, treatment with quercetin also decreased HSPs and viral transcription levels. Therefore, these results revealed the potential of quercetin as an effective anti-HBV agent with low toxicity.
-
We appreciate Dr. Michael Nassal for kindly providing the pCH-9/3091 plasmid. This work was supported by grants from the National Major Science and Technology Special Projects for Infectious Diseases of China (2012ZX10004503-008, 2012ZX10001006-002, and 2012ZX10002006-002), and National Natural Science Foundation of China (31300748).
-
The authors declare that they have no conflict of interest. This article does not contain any studies with human or animal subjects performed by any of the authors.
-
ZC, FZ, and KH designed the research. ZC, GS and YH performed the experiments. WG and WS provided experimental support. ZC and FZ drafted the manuscript. All authors read and approved the final manuscript for submission.














 DownLoad:
DownLoad: