HTML
-
Human herpesviruses (HHVs) are common enveloped double-stranded DNA viruses, belonging to Herpesviridae family (McGeoch 1989). There are nine herpesviruses that can infect humans, including herpes simplex virus 1 and 2 (HSV-1 and HSV-2, also known as HHV-1 and HHV-2), varicella-zoster virus (VZV or HHV-3), Epstein-Barr virus (EBV or HHV-4), human cytomegalovirus (HCMV or HHV-5), human herpesvirus 6A and 6B (HHV-6A and - 6B), human Herpesvirus 7 (HHV-7) and Kaposi's sarcomaassociated herpesvirus (KSHV, also known as HHV-8) (Lan and Luo 2017). According to genetic, biological and pathogenesis characteristics, nine HHVs are grouped into three sub-families, alpha- (HSV-1, HSV-2 and VZV), beta- (CMV, HHV-6A/B and HHV-7), and gamma-families (EBV and KSHV) (Norberg 2010; Lan and Luo 2017).
HHVs are globally distributed, and more than 90% of adults are ever infected by one or multiple HHVs (Lan and Luo 2017). HHVs typically establish latent infection in host, and undergo lytic reactivation in certain pathophysiological conditions (Schleiss 2009). Primary infection or reactivation of latent infections of HHVs can cause various clinical complications from mild or asymptomatic infections to deadly diseases such as aggressive lymphomas and sarcomas. Infection with different HHVs often lead to various diseases, such as herpes labialis, genital herpes (HSV-1, HSV-2) (Anderson et al. 2014), chicken pox (VZV) (Gershon et al. 2015), infectious mononucleosis (EBV and CMV) (Sarid and Gao 2011), burkitt lymphoma and nasopharyngeal carcinoma (EBV), and exanthema subitum or roseola (HHV-6A/B, and HHV-7) (Santos 2016). HHV infection often causes severe complications in immune-compromised patients, such as encephalitis (HSV-1 and HSV-2), pneumonia (CMV, HHV-A/B) and Kaposi's sarcoma (KSHV). Furthermore, CMV and/or EBV infection and co-infection with HHV-6 and/or HHV-7 are often associated with graft rejection (Sanchez-Ponce et al. 2018). Given effective antiviral therapy regimes are clinically available for various HHV infections, and the clinical manifestations caused by some HHVs are similar and non-specific, prompt and accurate diagnosis of HHV infection is of great significance, in particular for the organtransplant recipients and patients with severe neurological symptoms.
The conventional HHVs typing methods contain nucleic acid detection and non-nucleic acid detection. Common non-nucleic acid detections (mainly enzyme-linked immunosorbent assay) have good sensitivity, but are often prone to false positive results (LeGoff et al. 2014). The nucleic acid detection is often based on nucleic acid amplification with single-plex or multiplex PCR. It needs nine independent reactions to detect and distinguish all HHVs using single-plex PCR methods. Multiplex PCR methods often use fluorophores, or fragment sizes or melting curves of the products to distinguish different types of targets. Because of the limit in fluorescence channels and distinguishability of melting curves, at least two independent reactions are required to distinguish six-nine HHVs using multiplex PCR methods. Therefore, current nucleic acid detection of all HHVs is relatively expensive, time-consuming and laborious. Here, we developed a novel single-tube multiplex qPCR assay for detection of all HHVs. The new assay is sensitive, specific, and suitable for the clinical diagnosis of various HHVs.
-
To determine the sensitivity of the 2-D multiplex qPCR, 4 plasmids containing specific target fragments of HSV-2, EBV, HHV-7, and KSHV were synthesized by BioSune Biotechnology Co., Ltd. (Shanghai, China). Other plasmids containing genomic segments of HSV-1, VZV, HCMV, HHV-6A/B and GAPDH were obtained by PCR amplification, and TA cloning using pMD18-T plasmid vector (Dalian, Takara). The concentration of each plasmid was quantified by the Nanodrop 2000 ultra-micro spectrophotometer (Thermo, USA).
The specificity of the multiplex PCR was assessed using 14 common human viruses, including adenovirus (VR- 930), enterovirus (VR-1076), influenza A and B viruses (VR-333 and VR-789), parainfluenza viruses 3 (VR-93), HCoV-229E (VR-740), HCoV-OC43 (VR-1558), RSV-A (VR-1540) and RSV-B (VR-1400), human rhinovirus (VR- 489), DENV-1 (16007), DENV-2 (16681), DENV-3 (16562) and DENV-4 (1036). In addition to DENV-1 (16007), DENV-2 (16681), DENV-3 (16562) and DENV-4 (1036), all the above strains were purchased from the American Type Culture Collection (ATCC). Some clinical samples confirmed positive for other DNA viruses, including HPV52, HPV58, HPV66, HPV81 and HBV were also used to further evaluate the specificity of the multiplex PCR.
To evaluate the performance of the multiplex qPCR assay, a total of 149 whole-blood specimens and 21 vesicular fluid samples that were previously collected from suspected people with herpesvirus infection by First Affiliated Hospital of Kunming Medical University and Taizhou Fourth People's Hospital were used. The study was approved by Ethics Committee of Yunnan Provincial Biomedical (2020-01-LY-006), and the use of previously stored clinical samples was exempt from informed consent by the review board.
DNA was extracted from 200 μL whole blood or body fluids and eluted in 100 μL Nuclease-free H2O by QIAamp Blood DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer's instructions.
-
Primers and probes specific for the GAPDH were designed using the online tool PrimerQuest of IDT (https://sg.idtdna.com/Primerquest/Home/Index). The primer and probe sequences of HSV-1, HSV-2, VZV, EBV, HCMV, HHV- 6A/B, HHV-7 and KSHV were retrieved or modified from previous researches (Stamey et al. 2001; Sugita et al. 2008; Weidmann et al. 2008; Slavov et al. 2016). The melting temperature (Tm) of each amplicon was predicted using the online tool Oligo Calc: Oligonucleotide Properties Calculator (Northwestern University, Chicago, IL, USA), (http://biotools.nubic.northwestern.edu/OligoCalc.html) based on the amplicon sequence (Kibbe 2007). Actual Tm values of amplicons were measured by the melting curve analysis with Light Cycler 96 Real-Time PCR System (Roche Diagnostics, Germany). The detailed information of the primers and probes are shown in Supplementary Table S1.
-
The 2-D multiplex qPCR assay was performed using SuperReal PreMix (Probe) (Tiangen, Beijing, China). The reaction mixture contained 2 × SuperReal PreMix buffer (including chemically modified HotStar Taq DNA polymerase, antibody-blocked Taq DNA polymerase, dNTPs, MgCl2, etc.), 0.4 μmol/L fluorescent dye SYTO 9 (Life technologies, Carlsbad, CA, USA), 9 pairs of specific primers and 9 corresponding probes labeled by HEX, Texas red and CY5. The concentrations of primers and probes were shown in Supplementary Table S1. Three μL of DNA templates was included in each 25 μL reaction mixture. The reaction program was initiated with enzyme activation and denaturation at 95 ℃ for 15 min, followed by 40 cycles of denaturation at 95 ℃ for 3 s, and annealing and extension at 60 ℃ for 20 s, followed by a melting curve analysis from 65 to 95 ℃ with a speed of 0.007 ℃ per second. During PCR amplification, fluorescence signal was collected from each fluorescence channel at each annealing and extension step. In melting curve analysis, fluorescence signal was measured in FAM channel.
Specimens and Nucleic Acid Extraction
Primers and Probes Design
The 2-D Multiplex qPCR Assay
-
To develop the 2-D multiplex qPCR assay, specific primers and probes for nine HHVs, as well as one human housekeeping gene GAPDH were used. The probes for HSV-2, HHV-7 and GAPDH were labeled with Texas red, for EBV, HHV-6 (covering HHV-6A and -6B) and KSHV with CY5 and for HSV-1, VZV and HCMV with HEX (Supplementary Table S1). To distinguish the targets detected by probes with same fluorescence dye, the specific products were designed to have different Tm values with a Tm interval of at least 1 ℃. To assess whether the presence of herpesvirus variants occurring in the amplicon area of different herpesvirus strains alters the expected Tm, we downloaded all available genome sequences of nine herpesviruses from GenBank (Oct 26, 2020), and predicted the Tm values of all unique sequences in amplicon area. Nine herpesviruses appear to be very conserved with few variants at least in the region of amplicon (data not shown), and predicted Tm value has very little variability (Supplementary Table S1). To obtain the actual Tm values of each herpesvirus, we performed amplification and melting curve analyses using each herpesvirus plasmid with ten replicates. The actual Tm values were determined as 87.9 ℃, 79.7 ℃, 82.0 ℃ for HSV-2, HHV-7, GAPDH; and 82.8 ℃, 79.0 ℃, 84.8 ℃ for EBV, HHV-6 (covering HHV-6A and -6B), KSHV; 83.1 ℃, 86.3 ℃ and 84.0 ℃ for HSV-1, VZV and HCMV, respectively (Fig. 1 and Supplementary Table S2). The actual Tm values of each amplicon were very stable with a very small variation (Supplementary Table S1), and well matched the predicted Tm values for most herpesviruses except HSV-1 and HHV-7 that had a slight difference in Tm values between the predicted and actual values (Supplementary Tables S2). Importantly, the herpesviruses detected by the same fluorescent channel can be well distinguished from each other by their corresponding Tm values.
-
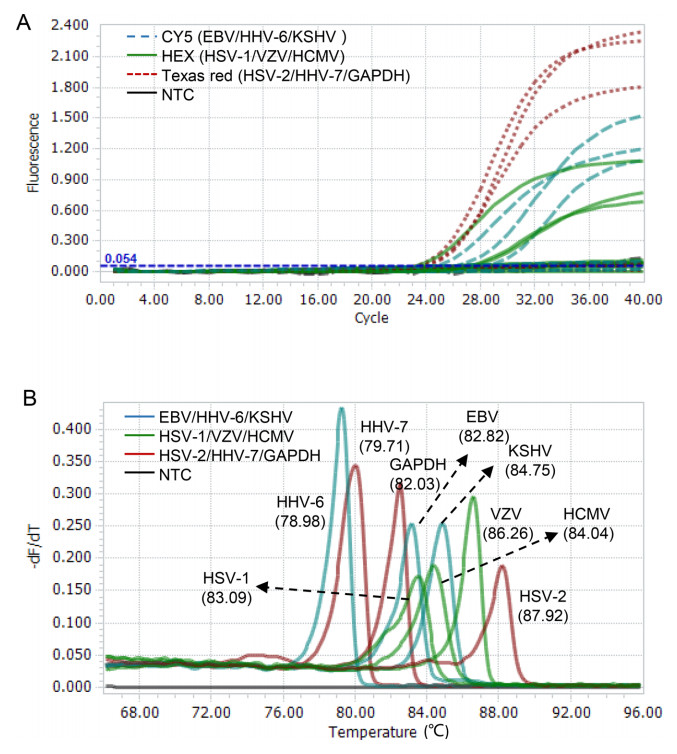
We assessed the performance of the 2-D multiplex qPCR assay using nine plasmids containing specific genomic segments of eight HHVs (HHV-6A and -6B sharing the same sequence) and a human gene GAPDH. The targets bound by the probes with same fluorescence dye (measured by the same fluorescence channel) were well distinguished by their corresponding Tm values when pooled templates were used (Fig. 2A). On the contrary, three amplicons with similar Tm values were easily distinguished by different fluorescent colors of probes (Fig. 2B). When combining the fluorescent color of probe and Tm value of amplicon, each gene was clearly distinguished in this multiplex qPCR assay (Fig. 2).
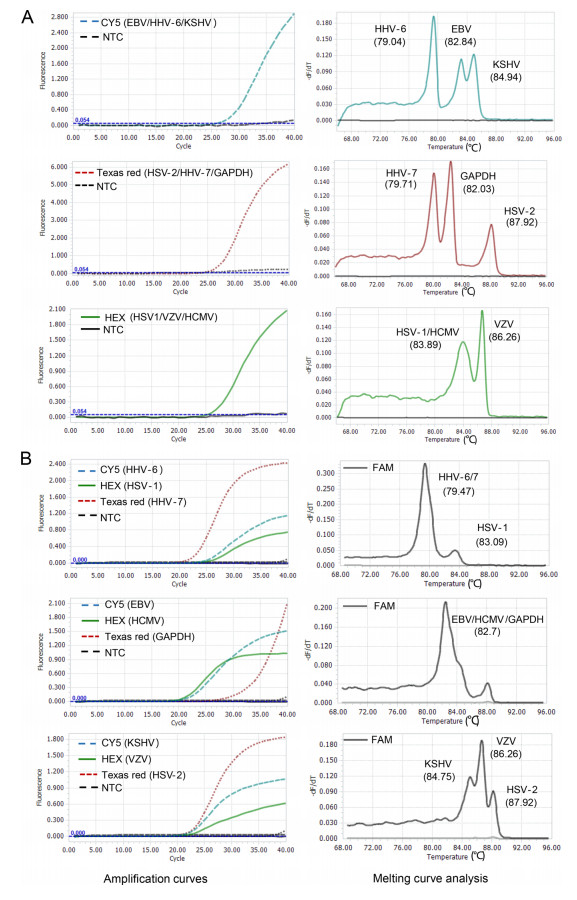
Figure 2. Proof-of-concept validation of the 2D-multiplex qPCR assay. A The co-existing targets detected by the same color probe are distinguished by different Tms. B Amplicons with similar Tms are distinguished by different color probes. The fluorescence channels used are shown in each plot. NTC non-template control.
To evaluate the capacity of the single-tube multiple qPCR assay in detection of co-existing targets, we tested all possible combinations (total: 36) of any two of the 9 plasmids. The co-existing targets sharing the same (fluorescent color) amplification curve but having two different Tm peaks, or having two amplification curves with different fluorescent colors but sharing similar Tm values, was very easily to be distinguished (Supplementary Fig. S1A and 1B). For the co-existing targets having two different amplification curves and two different Tm peaks (Supplementary Fig. S1C), they can be preferentially determined by their Tm values. To facilitate the judgment of Tm values of specific targets, a pooled template containing HSV-2, HHV-7 and GAPDH, which have Tms of 87.9 ℃, 79.7 ℃, and 82.0 ℃, respectively, was used as control for Tm references. By comparison with the reference Tm values, all co-existing targets having different color amplification curves and different Tm peaks were able to be correctly determined (Supplementary Fig. S1C).
-
To evaluate the sensitivity of the multiplex qPCR assay, tenfold serial dilutions of each plasmid from 3 × 107–3 × 101 copies/μL were tested (Supplementary Fig. S2). The results showed that the sensitivity was 300 copies per 25 μL reaction for HSV-1, HSV-2, VZV, EBV, HCMV, HHV-6A/B and GAPDH, 30 copies per 25 μL reaction for HHV-7 and KSHV. Specificity test showed that except GAGDH, there was no amplification for other fourteen common human viruses including seven DNA viruses by the multiplex qPCR assay, indicating that the new assay is specific to HHVs (Supplementary Fig. S3). We further tested whether human genomic DNA (gDNA) affects the specificity and amplification of the assay. The results showed that human gDNA does not inhibit the amplification of the multiplex qPCR assay (Supplementary Fig. S4).
-
To evaluate the accuracy of the single-tube multiplex qPCR assay for the detection of nine HHVs, 170 clinical samples collected from First Affiliated Hospital of Kunming Medical University and Taizhou Fourth People's Hospital were tested. From 149 whole-blood samples, the multiplex qPCR assay detected 30 positive samples for only one of nine HHVs, including 19 EBV, 6 HCMV, 1 HHV-7, and 4 KSHV (Table 1). In addition, 14 samples were detected as coinfection by two HHVs, including 5 by HSV-2 and HCMV, 8 by EBV and HCMV, and 1 by HHV-1 and KSHV (Table 1). From 21 vesicular fluid samples, the multiplex qPCR assay detected 14 positive samples for only one of nine HHVs, including 4 HSV-1, 3 HSV-2 and 7 VZV (Table 2). In addition, 5 samples were detected as coinfection by two HHVs, including 1 by HSV-2 and EBV, 1 by HSV-2 and HCMV, 1 by HSV-2 and HHV-7, 1 by VZV and EBV, and 1 by VZV and KSHV (Table 2). In particular, we detected one sample to be infected by three HHVs (HSV-1, HSV-2 and HCMV) and one sample with four HHVs (HSV-2, VZV, HHV-7 and KSHV) (Table 2). To evaluate the performance of the single-tube multiplex qPCR assay, all these samples were also tested by the single qPCR assay. The results of the single-tube multiplex qPCR assay were completely consistent with those of the single qPCR assay, indicting a 100% consistence rate with the latter (Tables 1 and 2). Among these samples, 73 (42.9%) samples were positive for at least one of all nine HHVs, and the EBV was the most commonly detected HHVs (Table 3).
Herpesvirus 2-D Multiplex qPCR assay (%) Single qPCR assay (%) Concordance rate (%) Positive EBV 19/149 (12.8) 19/149 (12.8) 100 HCMV 6/149 (4.0) 6/149 (4.0) 100 HHV-6A/B 0/149(0) 0/149 (0) 100 HHV-7 1/149 (0.7) 1/149 (0.7) 100 KSHV 4/149 (2.7) 4/149 (2.7) 100 HSV-2 & CMV 5/149 (3.4) 5/149 (3.4) 100 HSV-2 & EBV 8/149 (5.4) 8/149 (5.4) 100 EBV & HCMV 8/149 (5.4) 8/149 (5.4) 100 HHV-7 & KSHV 1/149 (0.7) 1/149 (0.7) 100 Negative 97/149 (65.1) 97/149 (65.1) 100 Total 149 /149 (100) 149 /149 (100) 100 Table 1. Evaluation of the single-tube nonuple qPCR assay for nine human herpesviruses and the reference gene GAPDH using 149 whole-blood samples.
Herpes virus 2-D Multiplex qPCR assay (%) Single qPCR assay (%) Concordance rate (%) Positive HSV-1 4/21 (19.0) 4/21 (19.0) 100 HSV-2 3/21 (14.3) 3/21 (14.3) 100 VZV 7/21 (33.3) 7/21 (33.3) 100 HSV-2 & EBV 1/21 (4.8) 1/21 (4.8) 100 HSV-2 & HCMV 1/21 (4.8) 1/21 (4.8) 100 HSV-2 & HHV-7 1/21(4.8) 1/21(4.8) 100 VZV & EBV 1/21 (4.8) 1/21 (4.8) 100 VZV & KSHV 1/21 (4.8) 1/21 (4.8) 100 HSV-1 & HSV-2 & HCMV 1/21 (4.8) 1/21 (4.8) 100 HSV-2 & VZV & HHV-7 & KSHV 1/21 (4.8) 1/21 (4.8) 100 Negative 0/21(0) 0/21 (0) 100 Total 21/21 (100) 21/21 (100) 100 Table 2. Evaluation of the single-tube nonuple qPCR assay for nine human herpesviruses and the reference gene GAPDH using 21 vesicular fluid samples.
Proportion in all samples (%) Proportion of positive samples (%) HHV single infection rate 25.9 60.3 HHVs double infection rate 15.9 37 HHV multiple infections rate 1.2 2.7 Common HHV single infection rate EBV 11.2 26 VZV 4.1 9.6 CMV 3.5 8.2 KSHV 2.3 5.5 HSV-1 2.3 5.5 HSV-2 1.7 4.1 High proportion of HHV among multiple infections HSV-2 10.6 24.7 EBV 10.6 24.7 CMV 8.8 20.5 VZV 1.8 4.1 HHV-7 1.8 4.1 KSHV 1.8 4.1 HSV-1 0.6 1.4 A total of 170 samples were tested Table 3. Detection rates of various HHVs.
On the other hand, in order to determine a suitable cutoff of the reference gene for the assay, we analyzed the Ct values of GAGDH in clinical samples. Because HSV-2 and HHV-7 share same fluorescent channel with GAGDH, 146 tested samples (including 97 negative samples but excluding 24 samples positive for HSV-2 and HHV-7) were used. The mean Ct value of GAGDH gene was 26.06 ± 3.66 with a scope of 19.06–35.13. For caution, a Ct value of 36 is recommended as the cutoff of GAGDH gene for the assay.
-
We further analyzed the Tm values of amplicons of clinical samples. For each herpesvirus, the Tm values of clinical samples had small variation, and were very similar to those of plasmid standard (Supplementary Table S2). These indicate that the multiplex qPCR assay has enough discrimination for the herpesviruses detected by the same fluorescent channel.
Design of the Novel Single-Tube 2-D Multiplex qPCR Assay
Proof-of-Concept Validation of the Novel SingleTube 2-D Multiplex qPCR Assay
Sensitivity and Specificity of the Novel SingleTube Multiplex qPCR Assay
Evaluation of the Novel Single-Tube Multiplex qPCR with Clinical Samples
Tm Value Variability of Amplicons from Clinical Samples
-
Herpesviruses are double-stranded DNA viruses that are distributed throughout the world and more than 90% of adults are ever infected with one or more HHVs. Infection with some HHVs can cause serious clinical symptoms. EBV and KSHV have been identified as carcinogenic by the International Agency for Research on Cancer (Sarid and Gao 2011). Previous studies showed that adults with normal immune function can also experience symptoms of mononucleosis when infected with EBV and HCMV (Sanchez-Ponce et al. 2018). Organ transplantation or immunodeficiency patients are more likely to be infected with multiple types of HHVs. Therefore, establishing a rapid, accurate and sensitive HHVs typing test is essential to diagnosis of HHV infection and clinical treatment (Floss and Dolff 2019).
In recent years, considerable progress has been made in the field of virus diagnosis by introducing molecular diagnostic methods such as PCR, LAMP and RPA (Kuhn and Frank-Kamenetskii 2008; Yoshikawa et al. 2014; Krumbholz et al. 2019; Zhou et al. 2019; Lu et al. 2020). The LAMP- and RPA-based assays are largely limited in multiplex detection. Currently, many PCR-based methods used for diagnosis of infectious diseases were designed to detect single pathogen or less than six pathogens in a single tube. Infections with some viruses are usually associated with similar clinical symptoms especially in the early stages of infection, but often cause different clinical outcomes. Therefore, increasing the targets in the single-tube multiplex PCR assay will facilitate clinical diagnosis of these viruses, and enable precise treatment.
Nowadays, qPCR is the commonly used multiplex nucleic acid detection methods. The strategy to reach the goal of multiplex detection is mainly involved in specific target discrimination by fluorescence colors or melting curve-based methods (Wan et al. 2016; He et al. 2020). Due to the limitation of the number of fluorescence channels of the instrument, the fluorescence color-based assays are usually less than quadruple in a single tube reaction (Sanchez-Ponce et al. 2018; Pyoria et al. 2020). To overcome the limitation of the instrument, a multi-color combination probe coding method was previously developed (Huang et al. 2011). However, this assay is difficult to distinguish co-existing target genes that generate similar amplification curves by different single-color probes, or are detected by a single-color probe and some double-color or triplex-color probers. On the other hand, the optimized amplicon of qPCR assay is 80–300 base pairs with the Tms of 80 ℃ –90 ℃ (Li et al. 2017). The narrow range of Tm limits the flux of melting curve-based methods. In order to obtain a higher throughput in a single-tube qPCR reaction, the idea of combining fluorescent colors with the probe Tm was first proposed in 2001 (Wittwer et al. 2001), and later verified by some other studies (Elenitoba-Johnson 2001; Liao et al. 2013). However, this method is difficult in probe design and has relatively high cost of probe synthesis.
In this study, we used the previously published 2-D multiplex qPCR method to successfully detect 9 types of herpesviruses, which can provide more options for the early diagnosis of HHVs (Li et al. 2020). Compared with the conventional multiplex qPCR methods, the novel method is simpler, less time-consuming, and labor-saving. This 2-D multiplex qPCR method has high sensitivity with 30–300 copies of various HHVs in 25 μL single-tube reaction, and does not cross-react with 20 other human viruses, including DNA and RNA viruses. The robustness of the novel assay was evaluated using 170 clinical samples. The novel assay showed a high consistency (100%) with the single qPCR assay for HHVs detection. Using the 2-D multiplex qPCR assay, we identified nine clinical samples co-infected with HSV-2 and EBV, six samples with HSV-2 and HCMV, eight with EBV and HCMV, one with HSV-2 and HHV-7, one with VZV and EBV, and one with VZV and KSHV (Table 1). In particular, we also identified a sample co-infected with three HHVs including HSV-1, HSV-2 and HCMV, and another sample with four HHVs including HSV-2, VZV, HHV-7 and KSHV. Although whether the two unique cases with three and four HHVs co-infection had more severe clinical outcomes was not recorded, previous studies well demonstrated that coinfection with multiple HHVs could cause more severe clinical outcomes than single HHV infection (Handous et al. 2020) and often occurred in immunosuppressed or immunocompromised individuals (Garib et al. 2013; Chen et al. 2016). For example, co-infection with HSV and HHV-6 increased the mortality of patients with encephalitis (Tang et al. 1997), and co-infection with HCMV and HHV-6 caused serious clinical outcomes with profound lymphopenia and pneumonia rash and increased the risk of bacterial and fungal co-infections (Handous et al. 2020). Therefore, it is important to develop a multiplex qPCR assay for the detection of HHVs co-infection. The novel 2-dimensional multiplex HHV qPCR assay provides a useful tool for rapid diagnosis of HHV infection.
In summary, we used the 2-D multiplex qPCR method to establish a nonuple qPCR assay for herpesvirus detection. The assay can detect the infection and co-infection of herpesviruses. This rapid, accurate, and cost-efficient multiplex HHV qPCR assay will facilitate the diagnosis of herpesvirus infection and guide timely clinical treatment.
-
This work was supported by the grants from the National Science and Technology Major Project of China (2019YFC1200603, and 2017ZX10103009-002).
-
CZ and Y-XL conceived and designed the study. CZ and Y-YL supervised the project. Y-YL, ZW, and SL collected and screened the samples. Y-XL, L-LZ, YM and HL performed the experiments. Y-XL and CZ interpreted the results. Y-XL and CZ wrote the manuscript. XJ and L-QZ provided critical suggestions on the results and contributed to revision of the manuscript. All authors read the manuscript and approved the submitted version.
-
These authors declare that they have no conflict of interest.
-
The use of patient samples in this study has been approved by the Yunnan Provincial Biomedical Ethics Review Committee.
















 DownLoad:
DownLoad: