-
Ebola virus is an enveloped, nonsegmented, negative-stranded RNA virus. Together with Marburg virus, they are classified in the order of Mononegavirales and the family of Filoviridae (29). Ebola virus is further grouped into four species: Zaire, Sudan, Ivory Coast and Reston. Infection by Ebola viruses can cause severe hemorrhagic fever characterized by widespread tissue infection and damage in multiple organ and tissue in human and other primates, and result in high mortality, up to 90% by Zaire species (29). Currently there is no effective vaccine or treatment. against Ebola infection in human.
The viral genome of Ebola is approximately 19Kb in length, and encodes seven genes: NP, VP35, VP40, GP, VP30, VP24, and L (30). The GP gene encodes two products through transcriptional editing (31, 36). The non-edited ORF encodes a smaller, secreted glycoprotein (sGP) which can be detected in large amount in the serum of the early phase of infected patients (31, 36). However the role of sGP in viral replication and pathogenesis is still not clear. The -1 ORF of the GP gene encodes a membrane-anchored glycoprotein (GP) which is the sole structural protein that forms the spikes on the virion surface (32).
GP is synthesized as the pre-GP in the ER, fully glycosylated in Golgi (GP0), and transported to the plasma membrane where virus budding occurs (13, 25, 32). GP0 is cleaved in the Golgi apparatus by furin-like proteases to generate two subunits, GP1 and GP2, which are linked by a disulfide bond (17, 24, 25). The native form of GP is a homotrimer of GP1/GP2 heterodimer on virions (32). GP1 is highly glycosylated containing both N-and O-linked carbohydrates, especially in the mucin-like region (13). GP1 is believed to be responsible for receptor binding, while GP2 for mediating membrane fusion and viral entry during Ebola infection. An X-ray structure of GP1/GP2 complex, believed to be at the native conformation, was recently reported. Together with the biochemical and extensive functional analysis of GP1, the receptorbinding domain (RBD) and some critical residues in RBD have been elucidated (4, 15, 16, 20, 21, 23).
In contrast, characterization of the Ebola GP2 subunit has been limited in scope. Nevertheless, the available information reveals that the Ebola GP2 subunit shares several characteristic features with other viruses. It was proposed that GP2 is structurally similar to the transmembrane subunits (TM) of retroviruses, particularly that of Rous sarcoma viruses (11, 37). The putative fusion peptide is located near the N-terminus. Following the putative fusion peptide is a region of heptad repeats (N-helical region) which are implicated in formation of coiled-coil structures. Another predicted amphiphathic helical region (C-helical region) is located to the C-terminal end of the GP2 ectodomain. The X-ray structures of the core GP2 ectodomain show that the core GP2 ectodomain forms a trimer in which a long central three-stranded coiled-coil is surrounded by shorter C-terminal helices that are packed in an antiparallel orientation into hydrophobic grooves formed by the inner coiled-coil (19, 39). This structural feature is shared by several viral fusion proteins such as HA2 of influenza virus, gp41 of human immunodeficiency virus, suggesting a common membrane fusion mechanism mediated by different viral glycoproteins. These proteins have been classified as the class Ⅰ fusion proteins (8). Mutational analysis of the hydrophobic residues in the N-and C-helices of GP2 indicated that some of these residues are indeed important in mediating Ebola entry (34), consistent with the structural features of the core GP2 ectodomain.
In this study, we examined the role of the charged residues in the GP2 helical regions in Ebola entry using a HIV pseudotyping system as a surrogate functional assay. We found that substitution of many charged residues in these regions greatly impaired the ability of Ebola GP to mediate efficient viral entry, indicating that these charged residues play an important role in viral entry. Elucidating the role(s) of these residues in protein folding and function of GP may provide insightful information on the molecular mechanism of Ebola entry and on the development of potential entry inhibitors against Ebola infection.
HTML
-
COS-7 cells and HeLa cells were maintained in Dulbecco's modified Eagles Medium supplemented with 10% fatal calf serum with penicillin and streptomycin. Human embryonic kidney 293T cells were maintained in the same medium plus 300 μg/mL Geneticin. A mouse monoclonal antibody, 12B5-1-1 (12), which recognizes the GP1 of Ebola Zaire GP (EBOZ) was obtained from Dr. Mary Kay Hart. The HIV anti-p24 monoclonal antibody was obtained from NIH AIDS Research and Reference Reagent Program. An anti-myc 9E10 monoclonal antibody was purchased from BAbCO.
-
Ebola Zaire GP (EBOZ-GP) gene was synthesized by multiple rounds of overlapping-PCR according to the EBOZ genome sequence submitted to Genebank by Sanchez A (Genebank accession number L11365). Totally 38 oligonucleotides, each about 75 nucleotides in length, were designed and used in the multiple round of PCR. In addition, the c-myc epitope coding region was appended to the carboxyl-terminus of GP to facilitate protein detection. Two silent mutations, both A to T, at the positions 738 and 1437 of EBOZ-GP gene, respectively, were introduced by the oligonucleotides to create an Xba Ⅰ and Hind Ⅲ sites in the GP gene for the convenience of cloning.
-
The synthesized EBOZ-GP gene was cloned into pcDNA3.1(+), a mammalian expression vector, after KpnI and BamH Ⅰ digestion, and named pEBOZ-GP. To examine GP expression, 10 μg of the pEBOZ-GP DNA was transfected into 293T cells in 100-mm plates by a modified calcium phosphate protocol as previously described (26). Forty-eight hours post transfection, the cells were lysed in Triton X-100 lysis buffer (50mmol/L Tris-HCl pH7.5, 150mmol/L NaCl, 5 mmol/L EDTA, 1% Triton X-100 plus protease inhibitors cocktail: leupeptin 10 μg/mL, aprotinin 5 μg/mL and PMSF 2 mmol/L). The clarified cell lysates were subjected to SDS-PAGE and Western blotting. The PVDF membrane was probed with EBOZ GP1 monoclonal antibody-12B5-1-1(1:5 000 dilution, final concentration 0.2 μg/mL) in 1%BSA-0.1%Tween-20-Tris-buffered saline (pH7.5) for one hour, washed three times by 0.1% Tween-20-TBS(pH7.5), then incubated with peroxidase-conjugated goat antimouse antiserum (Pierece) as the secondary antibody, and the proteins were detected by chemiluminescence method according to manufacturer's instructions (Pierce).
-
All GP2 mutants used in this study were generated by site-directed mutagenesis with pEBOZ-GP as the template, using Stratagene's Quick-Change Mutagenesis kit. All of the mutations were confirmed by DNA sequencing (University of Chicago CRC DNA Sequencing Facility).
-
Pseudotyped virion stocks were generated by co-tansfecting DNAs of pEBOZ-GP or GP2 mutant with the HIV vector (pNL4-3-Luc-R--E-), into 293T cells (1, 9). Briefly, 2 μg of DNAs of pEBOZ-GP or GP2 mutants and 2 μg DNA of HIV vector (pNL4-3-Luc-R--E-) was co-tranfected into 1×106 293T cells in six-well plates using Lipofectamine 2000 (Invitrogen). The media containing the pseudotyped virions were collected at 24 h and 48 h post-transfection, and pooled, clarified from floating cells and cells debris by low-speed centrifuge and filtered through 0.45 μm filter. The stocks were kept in -80℃ until use.
To examine the incorporation of the EBOZ GP or its mutants into the pseudotyped HIV virions, 2mL of viral stocks were layered onto a 3-mL cushion of 20% (w/v) sucrose in PBS and centrifuged at 55 000 r/min for 25 min in a SW55 rotor. The pelleted virions were lysed with Triton X-100 lysis buffer, and the preparations were subjected to SDS-PAGE and Western blotting to detect the EBOZ GP or mutant proteins. The HIV p24 in the virions was detected using an anti-p24 monoclonal antibody.
-
The cells of 293T, HeLa, or COS-7 cells were seeded in six-well plates 24 h prior to infection, and 1 mL of the HIV pseudotyped viral stocks was incubated with the target cells. After 5 h of incubation, the viral stocks were removed, and 2 mL of fresh medium was added to each well. Forty-eight hours post-infection, the cells were lysed in 200 μL cell culture lysis reagent (Promega), and luciferase activity of each lysate was measured with the Luciferase Assay Kit (Promega) using a FB12 luminomiter (Berthold Detection System), as previously reported (3).
-
To detect surface expression, 293T cells were transfected with DNAs of EBOZ-GP or its mutants by lipofectamine 2000. Forty-eight hours post-transfection, the cells were dissociated and suspended in FACS buffer (1%BSA-0.1% Sodium Azide in PBS). Approximately 1×106 293T cells were incubated with 0.5 μg of EBOZ-GP monoclonal antibody 12BF5-1-1 on ice for 30 min, washed with FACS buffer and stained with 0.5 μg FITC-conjugated goat anti-mouse IgG (Pierece). The stained cells were analyzed on a FACScan machine (Becton Dikinson). Dead cells that positively stained with propidium iodide were excluded from the analysis.
Cell lines and antibodies
Synthesis of EBOZ GP Gene
EBOZ-GP expression
Mutagenesis of EBOZ GP2
Generation of pseudotyped virion
Pseudotyped HIV infection assay measured by luciferase activity
FACS analysis of cell surface expression of EBOZ-GP and mutants
-
Thirty-eight overlapping oligonucleotides, each about 75 bases in length, were designed and used to synthesize the full length of the glycoprotein gene of Ebola Zaire species by multiple rounds of PCR (Fig 1). The final product is 2 067 bp in length including a myc-tag coding region at the carboxylterminus. Sequencing analysis indicated that there were 14 nucleotide substitutions, introduced either by nucleotide primers or PCR. Among them 10 substitutions are silent mutations including two designed ones that create two restriction endonuclease enzymes sites, Xba Ⅰ and Hind Ⅲ. The other four substitutions, C1093G, A1142G, A1231G, and C1421T, lead to amino acid substitutions of L365V, K381R, T411A, and A474V, respectively. All four residue substitutions are in the mucin-like region of GP1, and three of them are conserved substitutions. Deletion of this region by others (13, 14, 22, 33) and us (unpublished data) indicated that the mucin-like region of GP1 can be deleted without affecting viral entry. Therefore, these substitutions are not corrected.
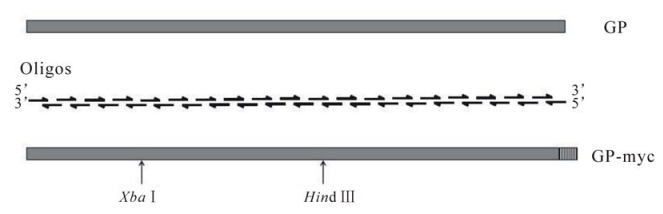
Figure 1. A schematic depiction of the strategy used to synthesize the EBOZ GP gene. The GP gene of Ebola Zaire species (EBOZ)-GP) was synthesized using 38 overlapping oligonucleotides and multiple-round PCR. The coding region of c-myc epitope (GSEQKLISEEDLG) was appended to the carboxyl-terminus of EBOZ GP to facilitate GP protein detection. The Xba Ⅰ and Hind Ⅲ sites as indicated in the figure were engineered by introducing silent mutations at positions of 738, 1437, respectively, to facilitate cloning of the synthesized EBOZ GP gene.
To examine expression of the synthetic gene, the synthesized EBOZ-GP gene was cloned into a mammalian expression vector, pcDNA3.1(+), and the construct was transfected into human embryonic kidney 293T cells by a standard calcium phosphate precipitation method. Cell rounding, detachment, and floating were observed approximately 20 h post-transfection, a well documented EBOZ-GP induced phenomenon by others{Chan, 2000 #387; Simmons, 2002 #349; Takada, 2000 #373; Yang, 2000 #389}. Following SDS-PAGE of the cell lysates and Western blotting, using a monoclonal anti-GP1 antibody 12B5-1-1 as the primary antibody, a band with an apparent molecular weight of 130 kDa, which corresponds to the size reported by others (32), was easily detected from the cells transfected by EBOZ GP construct (Fig. 2A, lane 2), while the mock transfected cells did not have the band (lane 1), indicating that EBOZ GP protein is well expressed in 293T cells. Also, we were able to detect GP2 from the cells transfected by the EBOZ GP construct, using an anti-myc MAb 9E10 (data not shown).
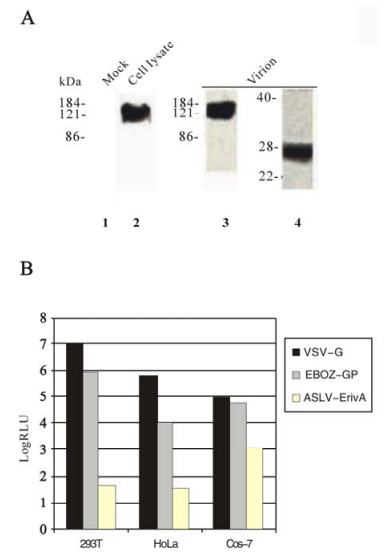
Figure 2. Analysis of expression and viral incorporation of GP, and GP-mediated infection in target cells. A: Expression and viral incorporation of GP protein. Expression: Transient expression of the synthetic GP gene was examined by Western blotting following SDS-PAGE. At 48 h posttransfection, 293T cells were lysed with Triton lysis buffer and aliquots of samples were subjected to SDS-APGE and Western blotting using the anti-GP MAb as the primary antibody, lane 1, mock transfected, lane 2, GP-transfected. Viral incorporation: The synthetic GP gene was co-transfected with the HIV vector into 293T cells, and the media were collected and centrifuged using a sucrose cushion. The pelleted virions were lysed and subjected to SDS-PAGE and Western blotting. Lane 3, Western blotting using the anti-GP MAb antibody as the primary, lane 4, Western blotting using the anti-myc MAb as the primary. B: GP-mediated HIV infection measured by luciferase activity. The HIV virions pseudotyped with Ebola GP, VSV-G (positive control), or EnvA (negative control) were incubated with three different cell lines (293T, HeLa, and Cos-7), the luciferase activities of the challenged cells were measured 48h post-infection, following the protocol as described in the Materials and Methods.
-
To set up an efficient Ebola pseudotyping system to study entry mechanism of Ebola viruses, we tested two retroviral-based pseudotyping systems: Murine leukemia virus (MLV) and human immunodeficiency virus (HIV), which have been reported by others as successful surrogate systems to study Ebola entry {Chan, 2000 #391; Jeffers, 2002 #314; Wool-Lewis, 1998 #348}. We found that the HIV-based system is more efficient than the MLV-based system for Ebola pseudotyping, since the MLV-based system with EBOZ GP gave a much lower signal over the background (data not shown) than the HIV-based system in our hands. Thus, we routinely use HIV-based pseudotyping system for Ebola pseudotyping.
To produce HIV/GP pseudotyped virions, 293T cells were co-transfected with two plasmids, one of them contains the HIV genome (env minus) with a luciferase reporter gene (pNL4-3-Luc-R--E-), and the other contains the EBOZ GP gene. The HIV vector co-transfected with the vesicular stomatitis virus glycoprotein (VSV-G) gene or subgroup A avian sarcoma and leucosis virus (ASLV-A) glycoprotein (EnvA) were used as the positive and negative controls, respectively. To examine incorporation of Ebola GP to the HIV virions, the supernatant from 293T cells cotransfected with the HIV vector and the GP gene was filtered, pelleted, and subjected to SDS-PAGE and Western blotting. A band with an apparent molecular weight of about 130 kDa which corresponds to GP1 was detected, using the anti-GP1 MAb 12B5-1-1 as the primary antibody (Fig. 2A, lane 3). In addition, the myc-tagged GP2 subunit with a molecular weight of approximately 26 kDa was detected in the virion pellet, using anti-myc MAb 9E10 (Fig. 2A, lane 4). The HIV capsid protein p24 was detected by Western blotting using an anti-p24 MAb (data not shown). Therefore, we demonstrated that GP can be efficiently incorporated to HIV particles.
To test the infectivity of the Ebola GP pseudotyped virions, three different cell lines, 293T, Hela, and COS-7, were challenged with GP, VSV-G, or EnvA pseudotyped HIV virions, following a protocol described in the Materials and Methods. Forty-eight hours post incubation, the luciferase activity of these cells was determined as a measure of viral infection. As expected, the EnvA pseudotyped HIV did not infect any of these cells since they do not express the receptor for ASLV-A, Tva{Bates, 1993 #426}. Thus the level of luciferase activity was used as the background in these experiments. It appears that COS-7 cells gave a higher background of luciferase activity than 293T or Hela cells (Fig. 2B). The GP pseudotyped HIV virions gave approximately 10 000 fold in 293T cells, 500 fold in Hela cells, and less than 100 fold in Cos-7 cells, respectively, of luciferase activity over the background (Fig. 2B). However, this cell-dependent discrepancy in viral infection is not a unique property of the GP-mediated viral infection since the VSV-G pseudotyped HIV virions displayed the same trend (Fig. 2B). Nevertheless, our results showed that the synthetic EBOZ GP gene is fully functional in mediating viral infection, as expected. Because 293T and Hela cells gives much higher signal to background ratio, these cell lines were used in the infection experiments in this study.
-
The core structure of the EBOZ GP2 is a long three-stranded coiled-coil surrounded by an extended linker and a shorter C-terminal helices that pack in an antiparallel orientation into the hydrophobic grooves formed by the central coiled-coil (19, 39). The role of the hydrophobic residues in the helical regions of Ebola GP2 has been examined previously by others, and it was found that some substitutions of these residues disrupt the stability of the coiled-coil, thus dramatically affect the membrane fusion function of this protein (34). In contrast, the role of the charged residues within the helical regions of Ebola GP2 in viral entry has not been reported. Sequence alignment of the GP2 proteins among different Ebola species indicates that almost all of the charged residues in the helical regions are conserved. For example, all of the charged residues within the N-helix are conserved, and all except two in the C-helix are also conserved (Fig. 3A).
Figure 3. The charged residues of the helical regions in GP2 targeted for mutational analysis. A: Sequence alignment of GP2 regions (residues 559 to 623) in different Ebola species. The charge residues of GP2 targeted for mutational analysis are highlighted. EBOZ5: Zaire 95, Genebank accession number P87666; EBOG4: Gabon 94, Genebank accession number O11457; EBOZM: Mayinga, Genebank accession number Q05320; EBOEC: Eckron 76, Genebank accession number P87671; EBOIC: Ivory Coast 94, Genebank accession number Q66810; EBORS: Reston Siena/Philippine-92, Genebank accession number Q89853; EBORE: Reston, Genebank accession number Q66799; EBOSB: Sudan Boniface, Genebank accession number Q66814; EBOSM: Sudan Maleo-79, Genebank accession number Q66798. B: Alanine-scanning mutagenesis of the charged residues in the coiled-coil and linker region of EBOZ GP2. The EBOZ GP2 residues 555-632 are shown. The charged residues substituted by alanines are indicated below the targeted residues. Sequence divergence among the charged residues of the different Ebola species are shown on the top of the sequence.
To examine the roles of the charged residues in the Ebola GP2 helical regions, we performed an alanine-scanning analysis of the charged residues within the N-helix, the linker, and C-helix of EBOZ GP2. All together, 21 substitution mutants were generated, including 8 in the N-helix, 6 in the linker, and 7 in the C-helix (Fig. 3B). Each of the substitution mutants was examined for protein expression in 293T cells, incorporation into HIV viral particles, and the ability to mediate viral entry by luciferase assay.
It was found that all 21 mutants were expressed in a level comparable to that of wild type EBOZ-GP in 293T cells (Fig. 4A, top panel). Furthermore, all of the mutant GP proteins except D614A (lane 16 in Fig. 4A, middle panel) were able to be efficiently incorporated into HIV particles (Fig. 4A, middle panel). These results suggest that substitutions of these charged residues do not have a detectably adverse effect on GP protein expression, processing, and protein folding. In these experiments, the HIV p24 levels were used as an estimate for quantifying HIV virions in the supernatants. As shown in Figure 4A (bottom panel), comparable amounts of p24 were detected in all lanes, suggesting that similar numbers of viral particles were produced in these experiments.
Figure 4. Analysis of expression, viral incorporation, and infectivity of the GP2 mutants. A: Expression and viral incorporation of GP mutant proteins. Total protein expression of GP2 mutants in 293T cells (cell lysate), and viral incorporation into HIV virions (virions) were examined as described in Fig. 2A. HIV viral p24 was used as a measure of relative HIV particles in the media. B: and C:. Relative levels of viral infectivity mediated by the GP2 mutants (the synthetic wt GP as 100%) using luciferase activities as the reporter, following the protocol described in the Materials and Methods. Bars, standard deviations. 293T cells were used as target cells in B. HeLa cells were used as target cells in C.
To examine the effect of these mutations on GP function, the pseudotyped HIV virions bearing the mutant GP proteins were incubated with 293T and Hela cells, respectively, and the luciferase activities of the infected cells were determined. All of the eight substitution mutants in the N-terminal helix displayed defective phenotypes in viral entry, giving less than 20% of the wt GP activity in 293T cells (Fig. 4B). Among them three mutants (R559A, R580A, and R587A) had the most adverse effect (less than 5% of wt GP). In contrast, mutations in the linker or the C-terminal helical regions exerted various degrees of effect. In the linker region, three substitution mutations (D607A, E611A, and H613A) had less effect (more than 20% of wt GP) than the other three mutants (R596A, H602A, and D614A). It is likely that the impaired incorporation of D614A (see Figure 4A) is responsible for its inability to mediate viral entry. Similarly, four substitution mutants in the C-terminal helix (K617A, D621A, K622A, and H638A) had less effect than the remaining three mutants (D624A, D629A, and D632A). The infectivity data of these mutants in Hela cells (Fig. 4C) are totally consistent with that in 293T cells. Together these results clearly demonstrated that the charged residues in the GP2 helical regions play an important role in mediating efficient Ebola entry.
-
Twenty substitution mutants of the charged residues, as demonstrated in Figure 4A, did not adversely affect GP expression and its incorporation into HIV virions, or MLV virions (data not shown), suggesting that these mutations of GP did not greatly alter the overall conformation on viral particles. These results suggest that the charged residues in these regions do not play a critical role in maintaining the native structure of GP, consistent with the notion that in the native form, these regions may not adapt well-defined structures on viral or cell surface. In contrast, one mutant, D614A, had a slight defect in incorporation into the HIV particles, suggesting that D614 plays a role in maintaining the native structure of GP. To further investigate the role of this residue in GP folding and function, two additional substitution mutants, D614E, and D614R, were generated, and their effect on GP expression, viral incorporation, and viral infection was determined. Substitution of D614 by another acidic residue, glutamate, appeared to have some adverse effect on viral incorporation (Fig. 5A, viral lysate, lane 3), but greatly impaired the ability of GP to mediate viral infection (less than 15% of wt GP, Fig. 5B). Substitution of D614 by arginine, a basic residue, reduced the efficiency of viral incorporation (Fig. 5A, viral lysate, lane 4), and abolished the ability of GP to mediate viral entry (Fig. 5B). These results suggest that both the negative charge and the size of D614 are important for proper protein folding and thus maintaining the optimal function of GP.
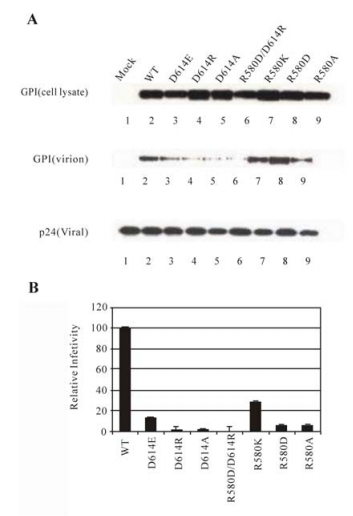
Figure 5. Analysis of D614 mutants in protein expression, viral incorporation, and infectivity. A: Expression and viral incorporation of D614 mutant proteins. B: Relative levels of viral infectivity mediated by the D614 mutants. The figure is similarly labeled like Figure 4.
The X-ray structure of Ebola GP2 indicates that the β-COOH group of D614 is proximal to the guanidino group of R580 (4.14Å), suggesting that these residues may interact via an intra-molecular salt bridge (19, 39). In an attempt to examine this possibility, two point mutants of R580 (R580K, and R580D), and a double mutant in which D614 is substituted by the positively charged arginine, and R580 is substituted by the negatively charged aspartate, respectively, (D614R/R580D), were generated. The effect of these substitutions on GP expression, viral incorporation, and viral infection was determined. Substitutions of R580 either by a negatively charged residue (R580D) or a positively charged residue (R580K) did not adversely affect GP expression or incorporation into virions (Fig. 5A, cell lysate and viral lysate, lanes 7 and 8). As expected, replacing R580 with the negatively charged aspartate (R580D) abrogated the ability of GP in mediating efficient viral entry, while replacing it with the positively charged lysine (R580K) maintained at least 25% of wt GP level (Fig. 5B), again suggesting that the positive charge and the size of R580 are important for maintaining the optimal function of GP. Therefore, it is not surprising that the double mutant (D614R/R580D) was defective in viral incorporation (Fig. 5A, viral lysate, lane 6) and in mediating viral infection (Fig. 5B), although it appeared to be expressed normally (Fig. 5A, cell lysate, lane 6). In addition, surface expression of this and other D614 mutant proteins was examined by FACS analysis, and it was found that all of them were comparable with the wt GP (data not shown). These results demonstrated that D614R can not be compensated by a second site substitution (R580D).
-
Among the aforementioned 21 charged residues in the GP2 helical regions, 11 are positively charged (R, K, and H), and 10 are negatively charged (D and E). Although the X-ray structure of the GP2 core does not reveal direct ionic interactions between them, an intriguing possibility is that some of these residues are involved in forming ionic bonds with the residues of the opposite charges of GP2 in the intermediate conformations of GP protein in Ebola entry. This hypothesis was tested using nine GP double mutants all of which contain substitution of R587 by an aspartate, and each also contains a "compensatory" second site substitution by an arginine of the negatively charged residues within the GP2 helical region, respectively. As shown in Figure 6A, none of these double mutants was defective in GP expression, nor in viral incorporation, suggesting that these substitutions did not greatly disrupt the native conformation of GP. Nevertheless, none of these mutants was able to confer a detectable level of the GP-mediated viral entry (Fig. 6B), indicating that R587D could not be "compensated" by any second site substitutions of the negatively charged residues in the helical regions.
Figure 6. Analysis of R587 mutants in protein expression, viral incorporation, and infectivity. A: Expression and viral incorporation of R587 mutant proteins. B: Relative levels of viral infectivity mediated by the R587 mutants. The figure is similarly labeled like Figure 4.
Synthesis and expression of the EBOZ GP gene
Establishment of an efficient Ebola pseudotyping system
Substitutions of the charged residues in the GP2 helical regions abrogated viral entry
The role of D614 in maintaining the native conformation of GP protein
The defect of mutant R587D could not be corrected by "compensatory" second site substitution of the acidic residues within the GP2 helical regions
-
One important finding of the current study is that many charged residues in the GP2 helical regions are critical in mediating efficient Ebola entry. We have shown that substitution of these charged residues by alanine scanning did not appear to adversely affect GP expression, processing, or viral incorporation, suggesting that these residues are unlikely essential in maintaining the native conformation of GP (the fusion-inactive state). Further, the available structure of the core GP2 ectodomain, which is believed to represent the fusion-active conformation of GP, does not give clue in revealing the role of these residues in membrane fusion or GP structure in this state. Thus this finding raises an intriguing possibility that these charged residues may play an important role in forming the intermediate conformation(s) in membrane fusion and Ebola entry.
Structural and functional analysis has revealed the common features of Ebola GP with other class Ⅰ fusion proteins including the HA of influenza virus, the HIV gp120/gp41 and that of other retroviruses, and the F proteins of paramyxoviruses (2, 5-7, 10, 18, 38, 40). One of the central features of these proteins in membrane fusion and viral entry has been illustrated by the major structural rearrangements and conformational changes of HA from the native, metastable fusion-inactive state to the stable, fusion-active state upon low pH treatment (5, 7, 40). Although the native structure of the GP2 has not been reported yet, the reported GP2 core structure is shown to be a trimeric coiled-coil formed by the N-terminal helices surrounded by a shorter C-terminal helices in an antiparallel orientation (19, 39), which is highly similar to that of the influenza virus HA2 and the HIV gp41. Since it appears that the Ebola entry, like that of influenza virus, is low pH-dependent (27, 35), a low pH environment in endosomes following viral internalization may induce a series of conformational changes on GP2, from the native conformation (fusion-inactive state) to the fusion-active state. Indeed, some substitutions of the hydrophobic residues in the GP2 helical regions impaired the ability of GP to mediate efficient viral infection (34), consistent with their role in forming the fusion-active conformation of GP2. The structural similarity of the core GP2 trimers to other class Ⅰ fusion proteins suggest a common fusion mechanism shared by Ebola and other viruses (8).
However, different class Ⅰ fusion proteins display a great diversity in primary sequences in the helical regions which are involved in formation of the six helical bundles. This sequence divergence may suggest that different fusion proteins use different strategies in activating fusion proteins and in forming the intermediate conformations in membrane fusion and viral entry. Indeed, it has been suggested that influenza virus HA employs a 'leash in the grove', rather than a helix-bundle, mechanism of membrane fusion (8). Curiously, functional analysis of the charged residues in the helical regions in membrane fusion and viral entry, to our knowledge, has not been systemically carried out in any class Ⅰ fusion protein. One possible reason for the lack of effort or interest is that the six helical bundle structures, for example, the core GP2 structure of Ebola, do not give clues regarding the role(s) of the most charged residues. We reasoned that these residues of Ebola GP2 might play important roles in protein folding and function in membrane fusion and viral entry. Indeed, most of the substitution mutants of the charged residues in the regions greatly impaired the ability of GP to mediate Ebola entry, even though these mutations did not appear to grossly affect expression, processing, and incorporation into viral particles (Fig. 4). Therefore, the current study revealed a critical role(s) of these charged residues in Ebola entry. It is interesting to point out that although there are many conserved charged residues in the helical regions in gp41 proteins of different HIV and SIV strains, similar analysis of the charged residues in the HIV gp41 helical regions indicated that substitution mutations at the C-helical region, in stark contrast to that in Ebola GP2 as reported in this study, did not greatly impair the ability of HIV glycoprotein in mediating viral entry (Jiang and Rong, unpublished data). These results suggest a possible mechanistic difference in Ebola GP-mediated membrane fusion from that of HIV gp41 in viral entry. Consistent with this hypothesis, others have noted that mutations of the N-helical region in GP2 exerted different effect on fusion activity compared to that in the HIV-1 gp41, and that a C-terminal peptide of GP2 was not as effective as that of gp41 in inhibiting membrane fusion and viral entry (34). In addition, the C-helical region of GP2 is much smaller than that of HIV gp41. Sequence alignment of the N-1 and C-1 helical regions did not indicate many conserved charged residues among the F proteins from paramyxoviruses (28). Thus, further investigation for diversity of fusion mechanisms utilized by various class Ⅰ fusion proteins is needed to elucidate the mechanistic specificity for each fusion protein.
The exact role(s) of the charged residues in the helical regions of Ebola GP2 in protein folding is not known yet. Further studies by an integral approach of molecular, biochemical, and structural analyses on GP are required to reveal how these charged residues are involved in maintaining the correct conformation(s) in the different steps during membrane fusion and Ebola entry. It is still possible that some of these residues play a role in maintaining the native conformation of GP such as stabilizing the trimeric state of GP, or interacting with GP1 which is highly unlikely based on the results in this report. However, we favor a hypothesis that these residues, at least some of them, are directly involved in forming the intermediate conformation(s) of GP2 during the transition from the native conformation to the fusion-active conformation in membrane fusion. Although the second site substitutions used in this study were unable to compensate the defect of one mutant (R578D) in the helical regions of GP2 (see Figure 6), we think it is likely that the oppositely charged residues in the N-helical and C-helical regions of GP2 interact with each other or with the charged residues outside of these regions by salt bridges. Analysis of these mutant proteins may allow us to capture the intermediate or transition conformations of GP2 in membrane fusion and Ebola entry.













 DownLoad:
DownLoad: