Medical Virology in Malaysia
2009, 24(2): 81 doi: 10.1007/s12250-009-3014-7
Received: 05 December 2008
Accepted: 10 February 2009
Virology is a branch of biological science dealing with the study of viruses, and medical virology focuses on the study and control of diseases due to viruses that is of medical importance. The development of medical virology in Malaysia has its beginning in the Institute for Medical Research (IMR), following the establishment of the Division of Medical Zoology and Virus Research in the institute on 23 March 1953. The second institution in the country to establish diagnostic and research work in medical virology was Department of Medical Microbiology, Faculty of Medicine, University Malaya. This was followed by University Kebangsaan Malaysia, University Sains Malaysia and University of Sarawak Malaysia. The National Public Health Laboratory (NPHL) is the latest institution to establish a laboratory in 2003 for virus isolation and services to support country surveillance and outbreak investigation of infectious diseases due to viruses. In the field of medical virology, Malaysia contributed substantially in the areas of virus diagnostic services, development and research ranging from survey and documentation on the existence and prevalence of viruses causing diseases in Malaysia, clinical presentation and epidemiological features of virus diseases, evaluation of new diagnostic tests to pathogenesis of viral diseases. Malaysia contributed to the discoveries of at least 12 new viruses in the world. ASEAN plus Three (China, Japan, Republic of Korea) Emerging Infectious Programme was established to overcome the challenges and impact of emerging and re-emerging infectious diseases in this region. Malaysia as the co-ordinator of the laboratory component of the programme, contributed to strengthen the regional laboratory capability, capacity, laboratory-based surveillance and networking. The future of medical virology in Malaysia in terms of integration of diagnostic, reference and research to support the country’s need will be enhanced and strengthened with the on-going development of the National Centre for Disease Control and Prevention (CDC Malaysia) which also incorporates a futuristic Special Diagnostic and Reference Laboratory
Bats and Viruses: a Brief Review
2009, 24(2): 93 doi: 10.1007/s12250-009-3032-5
Received: 10 February 2009
Accepted: 16 February 2009
Bats, probably the most abundant, diverse and geographically dispersed vertebrates on earth, have recently been shown to be the reservoir hosts of a number of emerging viruses responsible for severe human and livestock disease outbreaks. Flying foxes have been demonstrated to be the natural reservoir for Hendra and Nipah viruses. Evidence supporting the possibility of bats as potential reservoirs for SARS coronavirus (SARS-CoV) and Ebola virus has also been reported. The recent discovery of these viruses and other viruses occurring naturally in the bat population provides a unique insight into a diverse pool of potentially emergent and pathogenic viruses. The factors which influence the ability of zoonotic viruses to effectively cross the species barrier from bats to other animal populations are poorly understood. A brief review is provided here on the recently emerged bat viruses and on current and future strategies for research in this area
Novel Coronaviruses and Astroviruses in Bats
2009, 24(2): 100 doi: 10.1007/s12250-009-3031-6
Received: 20 January 2009
Accepted: 10 February 2009
Zoonotic transmissions of emerging pathogens from wildlife to human have shaped the history of mankind. These events have also highlighted our poor understanding of microorganisms circulated in wild animals. Coronaviruses and astroviruses, which can be found from a wide range of mammals, were recently detected in bats. Strikingly, these bat viruses are genetically highly diverse and these interesting findings might help to better understand the evolution and ecology of these viruses. The discoveries of these novel bats viruses not only suggested that bats are important hosts for these virus families, but also reiterated the role of bats as a reservoir of viruses that might pose a zoonotic threat to human health.
Hendra Virus Re-visited
2009, 24(2): 105 doi: 10.1007/S12250-009-3034-3
Received: 16 February 2009
Accepted: 26 February 2009
Hendra virus, a novel member of the family Paramyxovirus that has emerged from bats in Australia, causes fatal disease in livestock and humans. Eleven spillover events have been identified since the first description of the virus in 1994, resulting in a total of 37 equine cases and six human cases. All human cases have been attributed to exposure to infected horses; there is no evidence of bat-to-human or human-to-human transmission. Low infectivity and a high case fatality rate are features of Hendra virus infection in both horses and humans. The temporal pattern of spillover events suggests seasonal factors (plausibly be environmental, biological or ecological) as the proximate triggers for spillover. Minimisation of the future occurrence and impact of Hendra virus infections requires an understanding of the ecology of flying foxes, of virus infection dynamics in flying foxes, and of the factors that promote spillover. Management strategies seek to minimize the opportunity for effective contact between bats and horses, and limit potential horse-to-horse and horse-to-human transmission. Incomplete knowledge of the ecology of the virus, of the proximate factors associated with spillover, and the inherent difficulties of effectively managing wild populations, preclude a management approach targeted at bats.
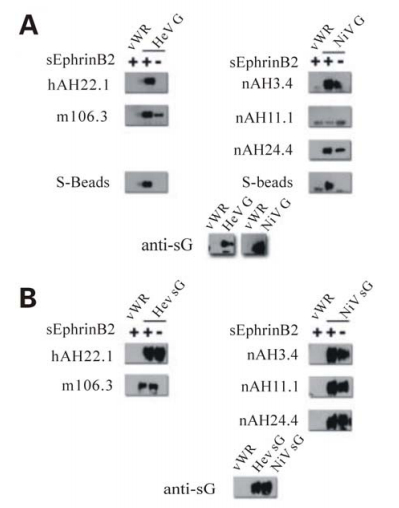
The Mechanism of Henipavirus Fusion: Examining the Relationships between the Attachment and Fusion Glycoproteins
2009, 24(2): 110 doi: 10.1007/s12250-009-3027-2
Received: 09 January 2009
Accepted: 16 January 2009
The henipaviruses, represented by Nipah virus and Hendra virus, are emerging zoonotic viral pathogens responsible for repeated outbreaks associated with high morbidity and mortality in Australia, Southeast Asia, India and Bangladesh. These viruses enter host cells via a class I viral fusion mechanism mediated by their attachment and fusion envelope glycoproteins; efficient membrane fusion requires both these glycoproteins in conjunction with specific virus receptors present on susceptible host cells. The henipavirus attachment glycoprotein interacts with a cellular B class ephrin protein receptor triggering conformational alterations leading to the activation of the viral fusion (F) glycoprotein. The analysis of monoclonal antibody (mAb) reactivity with G has revealed measurable alterations in the antigenic structure of the glycoprotein following its binding interaction with receptor. These observations only appear to occur with full-length native G glycoprotein, which is a tetrameric oligomer, and not with soluble forms of G (sG), which are disulfide-linked dimers. Single amino acid mutations in a heptad repeat-like structure within the stalk domain of G can disrupt its association with F and subsequent membrane fusion promotion activity. Notably, these mutants of G also appear to confer a post- receptor bound conformation implicating the stalk domain as an important element in the G glycoprotein’s structure and functional relationship with F. Together, these observations suggest fusion is dependent on a specific interaction between the F and G glycoproteins of the henipaviruses. Further, receptor binding induces measurable changes in the G glycoprotein that appear to be greatest in respect to the interactions between the pairs of dimers comprising its native tetrameric structure. These receptor-induced conformational changes may be associated with the G glycoprotein’s promotion of the fusion activity of F
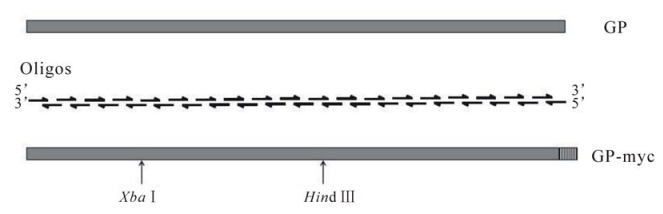
The Role of the Charged Residues of the GP2 Helical Regions in Ebola Entry
2009, 24(2): 121 doi: 10.1007/s12250-009-3015-6
Received: 05 December 2008
Accepted: 12 December 2008
The glycoprotein (GP) of Ebola is the sole structural protein that forms the spikes on the viral envelope. The GP contains two subunits, GP1 and GP2, linked by a disulfide bond, which are responsible for receptor binding and membrane fusion, respectively. In this study, the full length of GP gene of Ebola Zaire species, 2028 base pairs in length, was synthesized using 38 overlapping oligonucleotides by multiple rounds of polymerase chain reaction (PCR). The synthesized GP gene was shown to be efficiently expressed in mammalian cells. Furthermore, an efficient HIV-based pseudotyping system was developed using the synthetic GP gene, providing a safe approach to dissecting the entry mechanism of Ebola viruses. Using this pseudotyping system and mutational analysis, the role of the charged residues in the GP2 helical regions was examined. It was found that substitutions of the most charged residues in the regions did not adversely affect GP expression, processing, or viral incorporation, however, most of the mutations greatly impaired the ability of GP to mediate efficient viral infection. These results demonstrate that these charged residues of GP2 play an important role in GP-mediated Ebola entry into its host cells. We propose that these charged residues are involved in forming the intermediate conformation(s) of GP in membrane fusion and Ebola entry.
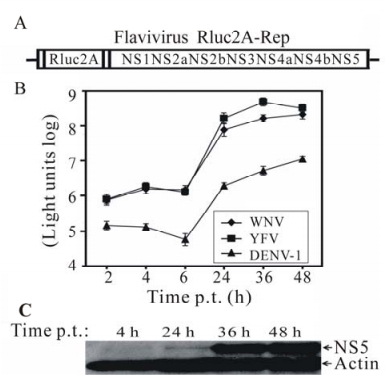
A Virus-type Specific Serological Diagnosis of Flavivirus Infection Using Virus-like Particles
2009, 24(2): 136 doi: 10.1007/s12250-009-3023-6
Received: 08 January 2009
Accepted: 14 January 2009
Many flaviviruses are emerging and reemerging pathogens, such as West Nile virus (WNV), dengue virus (DENV), yellow fever virus (YFV), and Japanese encephalitis virus. Serological assay is the dominant method for diagnosis of flavivirus infections in human. Because antibodies generated during flavivirus infections cross-react with other flavivirus members, plaque reduction neutralization test (PRNT) is the only available assay to determine the infecting flavivirus type. Since PRNT requires culturing raw viruses, it must be performed in biosafety level-3 or level-4 containment for many flaviviruses, and takes more than ten days to complete. To overcome these problems, we have developed flavivirus viral-like particles (VLPs) that could be used to replace raw viruses in the neutralization assay. The VLPs were prepared by trans packaging a luciferase-reporting replicon with viral structural proteins. This novel assay involves three simple steps: (i) VLPs from a panel of flaviviruses are incubated with flavivirus-infected sera at 37℃ for 1 h; (ii)the neutralized VLPs are used to infect Vero cells; and (iii) the infected cells are measured for luciferase activities at 22 h post-infection. The virus type whose VLP is most efficiently neutralized by the serum specimen (as quantified by the luciferase activities) is the etiologic agent. As a proof-of-concept, we show that a WNV-infected mouse serum neutralized the WNV VLP more efficiently and selectively than the DENV and YFV VLPs. Our results demonstrate that the VLP neutralization assay maintains the “gold standard” of the classic PRNT; importantly, it shortens the assay time from >10 days to <1 day, and can be performed in biosafety level-2 facility
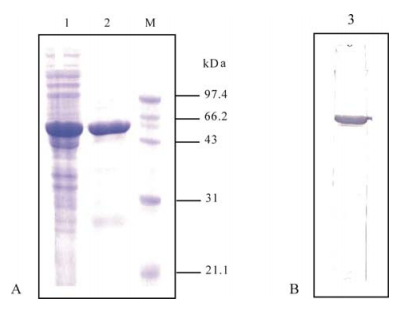
Indirect Enzyme-Linked Immunosorbent Assay Based on the Nucleocapsid Protein of SARS-like Coronaviruses
2009, 24(2): 146 doi: 10.1007/s12250-009-3037-0
Received: 24 February 2009
Accepted: 28 February 2009
The nucleocapsid protein (N) is a major structural protein of coronaviruses. The N protein of bat SARS-like coronavirus (SL-CoV) has a high similarity with that of SARS-CoV. In this study, the SL-CoV N protein was expressed in Escherichia coli, purified and used as antigen. An Indirect Enzyme-Linked Immunosorbent Assay (indirect ELISA) was developed for detection of SARS- or SL-CoV infections in bat populations. The detection of 573 bat sera with this indirect ELISA demonstrated that SL-CoVs consistently circulate in Rhinilophus species, further supporting the proposal that bats are natural reservoirs of SL-CoVs. This method uses 1-2 μl of serum sample and can be used for preliminary screening of infections by SARS- or SL-CoV with a small amount of serum sample