-
The Herpesvirus family is divided into three sub-families: Alphaherpesvirinae, Betaherpesvirinae and Gammaherpesvirinae. Herpes simplex virus type Ⅰ (HSV-1) is a neurotropic alpha-herpesvirus that only naturally occurs in humans with no known animal reservoirs. HSV-1 is a DNA virus with rapid pro-liferation and high infectivity. It is a common human pathogen that occurs worldwide with 60% to 95% of infection in adults, and is responsible for a broad range of human infectious diseases such as gingiva-stomatitis, keratitis, cutaneous herpes, genital herpes and encephalitis[25].
HSV-1 encodes its own ribonucleotide reductase (RR), which reduces ribonucleoside diphosphates to the corresponding deoxyribonucleotides and is essential for the formation of substrates for DNA synthesis[30]. The HSV-1 RR is composed of two distinct homodimeric subunits. The large subunit (Mr140000), designated ICP6 [9, 10], is encoded by the UL39 gene; the small subunit (Mr38000) is encoded by the UL40 gene[20]. The RR of HSV-1 is composed of two subunits arranged in a tight complex of α2β2 structure, and association of the subunits is necessary for enzyme activity [8, 13, 21, 22]. HSV-1 can not utilize cellular ribonucleotide reductase and therefore is dependent upon its own reductase for replication. It has been demonstrated that the viral enzyme may be required for virus growth in non-dividing cells and for viral pathogenesis and reactivation from latency in infected hosts in animal models studies [3, 4, 12, 14, 32].
RNA interference (RNAi) is an innate cellular anti-virus mechanism. RNAi technology, by use of synthesized small interfering RNA (siRNA) duplexes, has been reported as an effective tool for treatment of infectious diseases by silencing viral genes to inhibit replication[1]. We have demonstrated previously that synthetic siRNA targeting UL39 gene silenced effectively and specifically UL39 mRNA expression and inhibited HSV-1 replication[25]. In this study, we examined the effects of siRNAs targeting the UL39 and UL40 genes on HSV-1 replication in Vero cells. The effect of siRNA on the expression of target genes was measured by real-time quantitative reverse tran-scription PCR.
HTML
-
The HSV-1 strain F (ATCC VR-733) was obtained from Hong Kong University. The Vero cell line (ATCC CCL 81), obtained from Wuhan Institute of Virology, Chinese Academy of Sciences, was maintained in Dulbecco's modified Eagle medium (DMEM) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 0.22% sodium bicarbonate (Sigma, St. Louis, USA), and 50 μg/mL gentamycin (Gibco). For experiments, the Vero cells were cultured in DMEM without serum and antibiotics. The HSV-1 strains were grown in the Vero cells at 37℃.
-
The siRNAs used in this study consisted of 21-nucleotide double-stranded RNAs, each strand of which contained a 19-nucleotide target sequence and a two-uracil (U) overhang at the 3' end. According to the design rules for RNAi[7], four fragments of UL40 gene from HSV-1strain F were predicted to have RNAi capacity. These fragments were used as the target siRNA (siRNA1, siRNA2, siRNA3, and siRNA4). The siN.C, not targeting any known gene, was used as negative control (GenePharma) (Table 1). The siRNA39 targeting the UL39 gene was designed as siRNA392 described previously[25]. All siRNAs were synthesized by Shanghai GenePharma Co., Ltd.

Table 1. Design of siRNA targeting the UL40 gene
-
Vero cells were grown in 24-well plates to 80% -90% confluence and then transfected with specific or control siRNA (80 nmol/L) using Lipofectamine 2000. After 4 h, the cells were infected with 25 PFU of HSV-1 strain F. Cells were overlaid with 1 mL of a 1:1 mixture of NaCMC and 2× Dulbecco's modified Eagle medium without serum to allow only cell-to-cell spread of virus. At 48-72 h post-infection, plates were fixed with 10% paraformaldehyde for 2 min and then stained with 1% crystal violet for 20 min to count the number of plaques per well and to photograph the monolayer. Overlapping plaques and plaques at the edge of the well were counted as a single plaque.
-
Vero cells were grown in 24-well plates to 80% -90% confluence and then transfected with specific or control siRNA (80 nmol/L) using Lipofectamine 2000. After 4 h, the cells were infected with 25 PFU of HSV-1 strain F. After 48 h, total RNA from the infected cells was extracted using Trizol (Invitrogen). RNAs were reverse transcribed to yield single-stranded cDNA.
Real-time PCR reactions were performed using SYBR Green Real-time PCR Master Mix (TAKARA, Japan) according to the manufacturer's instructions, by using the ABI Prism 7000 Sequence Detection System (Applied Biosystems). The following primers were used for amplification. UL40 sequence specific primers: Forward, 5′-CTTCCTCTTCGCTTTCCTGT CG-3′; Reverse, 5′-CGCTTCCAGCCAGTCCACCT T-3′. UL39 sequence specific primers: Forward, 5′-G GCTGCAATCGGCCCTGAAGTA-3′; Reverse, 5′-G GTGGTCGTAGAGGCGGTGGAA-3′. 18s RNA sequence specific primers: Forward, 5′-CCTGGATA CCGCAGCTAGGA-3′; Reverse, 5′-GCGGCGCAATACGAATGCCCC-3′.
Each real-time PCR reaction was performed for 40 cycles at 95℃ for 1 min, 95℃ for 15 sec, 58℃ for 15 sec and 72℃ for 40 sec. The relative expression of UL40 gene was normalized to the house-keeping gene 18s RNA and was calculated using the formula[2]: Rel Exp =2(-ΔΔCT) Each real-time PCR reaction was performed in triplicate.
-
Results were given as means ± S.D. and statistical significance was determined by the t test; P values of less than 0.05 were considered as statistically significant.
Viruses, bacterial strains and cells
Synthesis of siRNAs
Plaque assays
Real-time RT-PCR
Statistical analyses
-
HSV-1 produces visible plaques on serum-starved Vero cells at 48h post inoculation that develop central clearing as the virus spreads outward (Table 2). Vero cells were transfected with UL40-specific siRNAs (siRNA1, siRNA2, siRNA3 and siRNA4) or negative control siN.C and then infected with 25 PFU of HSV-1. Compared with virus control, transfection with siN.C or UL40-specific siRNA1 had no effect upon viral plaque formation following infection (P > 0.05). In contrast, HSV-1 infection of Vero cells that were transfected with UL40-specific siRNAs (siRNA2, siRNA3 and siRNA4) produced much less plaques (P < 0.01). The inhibition rates of siRNA1, siRNA2, siRNA3 and siRNA4 on HSV-1 plaque formation was 1.6%, 50%, 41.9% and 62.9%, respectively.

Table 2. Effect of siRNA on HSV-1 plaque formation
-
HSV-1 produces visible plaques on serum-starved Vero cells at 48h post inoculation that develop central clearing as the virus spread outward (Table 3). Vero cells were co-transfected with UL40-specific siRNAs (siRNA1, siRNA2, siRNA3 or siRNA4) and UL39-specific siRNA39 and then infected with 25 PFU of HSV-1. Compared with virus control, transfection with siN.C had no effect upon viral plaque formation following infection (P > 0.05). In contrast, HSV-1 infection of Vero cells that were co-transfected with UL40-specific siRNAs (siRNA1, siRNA2, siRNA3 or siRNA4) and siRNA39 produced much less plaques (P < 0.05 or P < 0.01).The inhibition rates of siRNA1+siRNA39, siRNA2+siRNA39, siRNA3+siRNA39 and siRNA4+siRNA39 on HSV-1 plaque formation was 22.3%, 49.2%, 62.3% and 73.8%, respectively.

Table 3. Effect of siRNAs on HSV-1 plaque formation
-
To determine whether the decrease in the number of HSV-1 plaques was due to siRNA silencing expression of the UL40 gene or UL39 gene, a real-time PCR analyses was performed as above. The results indicated UL40-specific siRNAs but siRNA1 remarkably inhibited the expression of the UL40 gene (Fig. 1). Compared with virus control, siRNA1、siRNA2、siRNA3 and siRNA4 reduced the expression of UL40 gene to levels that were 97.27%(P > 0.05)、30.17%(P < 0.01)、32.06% (P < 0.01) and 18.69% (P < 0.01) respectively less than those seen in cells treated with negative control siN.C. In UL40-specific siRNAs and UL39-specific siRNA39 co-transfected, UL40-specific siRNAs and UL39-specific siRNA39 remar-kably inhibited the expression of the RR genes (Fig. 2). Compared with virus control, siRNA1 and siRNA39 reduced the expression of UL40 gene and UL39 gene to levels that were 98%(P > 0.05) and 68.5% (P < 0.01) respectively; siRNA2 and siRNA39 reduced the expression of UL40 gene and UL39 gene to levels that were 36.52%(P < 0.01) and 32.38% (P < 0.01) respectively; siRNA3 and siRNA39 reduced the expression of UL40 gene and UL39 gene to levels that were 33.89%(P < 0.01) and 31.23% (P < 0.01) respectively; siRNA4 and siRNA39 reduced the expression of UL40 gene and UL39 gene to levels that were 19.43%(P < 0.01) and 28.69% (P < 0.01) respectively.
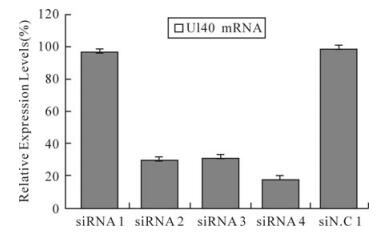
Figure 1. Relative expression levels of UL40 mRNA in HSV-1 infected Vero cells. The relative expression of UL40 mRNA in cells treated with different siRNAs compared with virus control (100%) was determined by real-time PCR. The result was normalized to the house-keeping gene 18s RNA and was calculated using the formula: Rel Exp = 2(-ΔΔCT). Data represent the means ± S.D. of three wells on same plate.
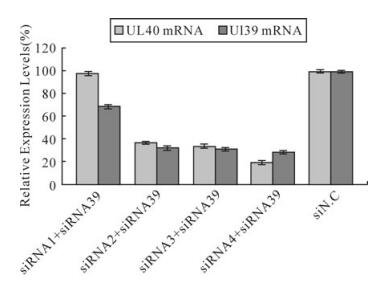
Figure 2. Relative expression levels of RR mRNA in HSV-1 infected Vero cells. The relative expression of RR mRNA (UL39 mRNA and UL40 mRNA)in cells treated with different siRNAs compared with virus control (100%) was determined by real-time PCR. The result was normalized to the house-keeping gene 18s RNA and was calculated using the formula: Rel Exp = 2(-ΔΔCT). Data represent the means ± S.D. of three wells on same plate.
Effect of UL40-specific siRNA on HSV-1 infection
Effect of UL40-specific siRNA and UL39-specific siRNA co-transfected on HSV-1 infection
Effect of siRNA on the expression of RR genes
-
Despite the impact of RNAi in probing gene function and in the development of novel therapeutics and antivirals, there have been only a few studies exploring the potential for RNAi approaches to HSV-1. These reports have focused primarily on the α and γ genes of HSV-1[18, 23, 32]. RR, as a β gene, is a well-recognized target for antiviral agents given the direct role it plays in regulating DNA replication[5, 26, 29] and its indirect role in regulating other enzymes in the DNA synthesis pathway via control of the nucleotide pool. In the present report, we showed that HSV-1 was susceptible to the RNAi pathway, both the siRNAs tested resulted in silencing of the HSV-1 RR gene and a reduction in viral plaque formation.
Although four siRNAs had sequences homologous to the HSV-1 UL40 mRNA, they differed in their efficacy to establish RR silencing. In several independent experiments, siRNA4 consistently inhibited UL40 expression more efficiently than the other siRNAs, indicating that the difference in efficacy can not be ascribed solely to differences in transfection efficiency. Variation in the silencing activity of several siRNAs targeting the same mRNA has also been reported previously [11, 16, 31]. The reasons for the differences in RNAi efficacy are not well understood, but has been postulated to be due to differences in the thermodynamic properties of the siRNAs[17], as well as secondary and tertiary RNA structure of the target site that disables recognition by the siRNA[16, 19, 27].
David et al [6]demonstrated that HSV-1 ribonucleotide reductase activity is not essential for growth in cultured cells and that the cellular enzyme, when active, can support virus growth. They found hrR3, which is incapable of inducing viral ribonucleotide reductase activity in untransformed cells, was capable of synthesizing substantial amounts of viral DNA in exponentially growing Vero cells (74% as compared with wild-type levels). However, viral DNA synthesis of the mutant was more compromised when it was grown in serum-starved Vero cells (29% as compared with wild-type levels in serum-starved Vero cells). No difference existed in the relative amounts of DNA synthesis when wild-type virus was grown under either condition. For our experiments, the Vero cells were cultured in DMEM without serum and antibiotics. Our results showed that siRNAs silencing RR genes could inhibit efficiently HSV-1 replication in serum-starved Vero cells. This is concordant with their finding.
The development of siRNA-mediated antiviral therapy may be of greatest benefit among populations infected with virus strains resistant to conventional antivirals and in cases of severe or recrudescent disease. In vivo activity of RNAi has been demon-strated for both viral and host genes of mice [15, 28]. These data suggest that siRNA can be developed as a parenteral drug. The in vitro system described will enable us to more easily pursue further studies of RR functions and provides a foundation for the development of in vivo applications of siRNA.













 DownLoad:
DownLoad: