-
Foamy viruses (FVs) comprise the only genus in the Spumaretrovirinae subfamily of the Retroviridae and have been isolated from many mammalian species including human, non-human primates (chimpanzee, baboon), bovine, equine and feline species [1, 4, 6, 15, 16, 25]. FVs have three hallmark retroviral genes, gag, pol and env. Meanwhile, there are at least two accessory genes, named tas and bet, located between env and the 3′ long terminal repeat (LTR) [23]. However, different from ordinary retrovirus, such as human immunodeficiency virus (HIV) and human T-cell leukemia virus (HTLV), FVs have a separate pol mRNA, an infectious DNA genome and two promoters in their genome, the LTR and the internal promoter (IP) [24]. Tas, the product of tas, can activate both the LTR and IP promoters in trans as the pivotal regulator of viral gene expression during FVs′ replication [5, 10]. All these special features of FVs set them apart from the other known retroviruses and make them attractive targets for study in order to understand more details about the FVs evolutionary advantages and to investigate their suitability as transfer vectors. Strain 3026 of bovine foamy virus (BFV3026) was isolated by our laboratory in 1995 [13].The strain shows typical features of FVs and has been investigated with the aim of constructing a gene transfer vector.
FV infection seems to lead to life-long latency in almost all hosts since these viruses are nonpathogenic in their natural hosts or in experimentally infected animals [7, 11, 12]. However, their infection can induce cellular syncytia formation only in susceptive cells in vitro, while in some other cells they cannot [17, 22]. Therefore, the traditional titration methods based on cytopathic effect (CPE) may have some limitations in titrating foamy viruses. Indicator cell lines have been developed for the rapid and efficient detection and quantification of viral infections. Since the viral transactivator, Tas, can activate the promoter of a reporter gene, the viral infection can be determined through the expression of the viral protein. This technique has been applied widely for the diagnosis of various viruses, such as HIV, herpes simplex virus (HSV), primate foamy virus (PFV) and feline foamy virus (FeFV) [19, 23]. To date the chloramphenicol acetyltransferase (CAT) gene, the β-galactosidase gene, the enhanced green fluorescent protein (EGFP) gene and the luciferase gene have been used as reporter genes.
In this study, we established the BFV indicator cell line-BFVL, in which the firefly luciferase gene was employed as a reporter gene, to detect the presence of infectious BFV based on activation of the LTR to express firefly luciferase. The BFVLTR-driven luciferase plasmid was transfected into baby hamster kidney cells (BHK21) and the stable transfected cells were screened and isolated. This cell line was determined to be stable, sensitive and specific in detecting and quantitating the active BFV infection.
HTML
-
Baby hamster kidney cell line (BHK-21), Canine thymus cell line (Cf2Th) and BFV indicator cell line (BICL), which harbored EGFP reporter gene and was established by our laboratory previously [14], were maintained in Dulbecco's Modified Eagle Medium (DMEM, Gibco) supplemented with 10 % fetal bovine serum (FBS) and 2 mmol/L glutamine, 50 U/mL penicillin and 50 μg/mL streptomycin sulphate. All the cells were cultured at 37℃ in 5 % CO2.
The Bovine foamy virus (BFV 3026 strain) was isolated from peripheral lymphoid cells of infected bovines by our laboratory in an earlier study [13]. The viruses were propagated in Cf2Th cells. When typical syncytia were formed, ten 10-cm dishes of cells were gathered and resuspended in 10 mL freeze medium (10% dimethyl sulfoxide, 20% fetal bovine serum, 70% DMEM) and then stored as stock viruses at -70℃. The Bovine immunodeficiency virus (BIV R29 strain) was provided by Dr. Charles Wood (University of Nebraska, Lincoln) and cultured in Cf2Th. The preparation of BIV virus stocks was similar to BFV. Bovine herpes virus-1 (BHV-1) was stored in our laboratory and cultured in MDBK cells. When typical syncytia were observed, the medium was collected and centrifuged at 3000 × g for 10 min. Then the supernatant was filtered through a 0.45 μm filter membrane and stored at -70 ℃ as BHV-1 virus stock. PFV was stored in our laboratory and cultured in BHK-21 cells. The preparation of PFV virus stocks was similar to BHV-1.
-
The BFV3026 LTR sequences (positions -7 to 1012) were amplified by polymerase chain reaction (PCR) using the BFV3026 infectious clone as a template, with the following primer sets: forward, 5′-TCCGGTACCGAGAGGGTGTGGTGGGAAG-3′; reverse, 5′-GAACTCGAGTCTCTCACGGGCGCAGC-3′, with the following parameters: 94 ℃ for 5 min, then 30 cycles with a denaturation step at 94 ℃ for 1 min, annealing at 55 ℃ for 1 min, and elongation at 72 ℃ for 1 min, with a final extension at 72 ℃ for 7 min. The 5′-and 3′-primers were engineered with Kpn I and Xho I (restriction sites are underlined in the primers sequences). The PCR fragments were subsequently digested and directionally cloned into the corresponding sites of the pGL3-Basic vector (Promega). The construct was referred to as pGL3-BFVLTR, an intermediate plasmid. The intermediated plasmid was digested with Not I and Sal I. The digested fragment was directionally cloned into the corresponding sites of the pCMV-tag2B vector (Stratagene). The construct was referred to as the last BFV reporter plasmid, pCMVtag2B-BFVLTR.
-
To establish stable cell lines, BHK cells plated at a density of 105 cells/well in 12-well plates were transfected by standard polyethylenimine (PEI) (Polysciences Inc, USA) procedure with 1 μg pCMVtag2B-BFVLTR. At 24 h post-transfection, cells were subjected to neomycin (G418, Gibco) selection (600 μg/mL) for 2 weeks until colonies appeared. Stable clones with G418 resistance were obtained by trypsinization of colonies, followed by plating at limiting dilutions onto a 96-well plate. Ten single clones were then expanded and analysed for luciferase expression upon BFV3026 infection. One pCMVtag2B-BFVLTR-containing clone designated BFVL was selected for further study, on the basis of low background expression levels and a constant increase in luciferase expression along with increasing virus titers.
-
The indicator cells were plated in 12-well plates (105/ well) and cultured at 37℃ for 20 h. The cells were then incubated with a serial dilution of BFV3026 stocks at 37℃ for 48 h. The luciferase activity was measured by using a commercial assay system (Promega, USA). The cells were rinsed with PBS and lysed with lysis buffer (Promega, USA). The cell lysates were then clarified from the insoluble materials by centrifugation at 12000 rpm for 3 min. The clarified lysate (5 μL) was mixed with the luciferin reagent (50 μL) and luciferase activity was measured with a chemiluminescence measurement machine (Promega, USA), the readout was count per second. The relative light unit was determined automatically.
-
Analysis of Bovine Tas (BTas) and Gag expression was performed as described previously [26, 24]. Briefly, the indicator cells were transfected with pcDNA3.1 (+) -BTas clones. Then after 48 h, the cells were rinsed twice in PBS and then lysed. Equal amounts of cell lysates were separated by 12 % polyacrylamide gel electrophoresis (SDS-PAGE) and analyzed by Western blotting with murine anti-Tas polyclonal sera (prepared by our lab). The indicator cells were also infected by BFV3026 and 48 h following infection the cells were prepared as described above. The virus titer was indicated by capsid protein Gag expression analyzed with murine anti-Gag polyclonal sera (prepared by our lab). Goat anti-mouse Ig peroxidase-conjugated secondary antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA).
-
The Cf2Th or BFVL cells in 96-well plates (104cells / well) were infected with 10-fold serial dilutions of viruses and incubated for 48 h at 37℃ in a humidified atmosphere of 5 % CO2 in air. The infected cells were observed for syncytia formation or lysed to detect luciferase expression, respectively. In the CPE-based assay, the well containing one syncytia formation in Cf2Th cells represented the reference positive well used for calculating the BFV′s titers. Meanwhile, the well in which the luciferase activity ratio relative to the control was more than 5-fold was regarded as the positive well in the BFVL-based assay. The virus titers were expressed by TCID50, calculated with the Reed-Muench method [9].
Cells, culture conditions and virus
Plasmids
Transfection and establishment of stably trans-fected cell lines
Luciferase assay
Western blotting
Titration of BFV using the Cf2Th and BFVL cells
-
To generate reporter cell lines that can be used for screening of BFV infection more simply, rapidly and sensitively, the BHK cells which were permissive for BFV replication were chosen. The cells were trans-fected with pCMVtag2B-BFVLTR, which contained the luciferase reporter gene driven by the BFV3026 promoter (positions -7 to 1012). This plasmid harbors the neomycin-resistance gene that confers G418 resistance advantage to transfected cell lines. Ten neomycin-resistant clones were then isolated by limiting dilutions and analyzed for basal and BFV-induced luciferase expression. One clone designated BFVL was selected for further study, on the basis of low background expression levels and a constant increase in luciferase expression along with increasing virus titers. As shown in Fig. 1A, luciferase expression in BFVL cells was induced in a dose-dependent manner with BTas 2-fold increase since BTas could drive it through the LTR region. We have shown that BFVL cells could respond to BTas activation and the luciferase induction in BFVL cells is proportional to the amount of BTas ranged from 50 ng to 800 ng, as detected by Western blot analysis of the level of protein BTas (Fig. 1A, bottom panel). To determine the response ability of BFVL cells to BFV infection, 2 × 105 BFVL cells were incubated with 2-fold diluted BFV3026 stock viruses ranging between 3125 to 1 × 105TCID50. At 48 h post infection, luciferase expression was analyzed. We found it increased with the increasing virus titer as detected by Western blot analysis of the level of representative viral capsid protein Gag (Fig. 1B). These results demonstrated that BFVL cell lines stably harboring the luciferase reporter system were generated.

Figure 1. Characteristics of BFVL cells. A: BTas induces luciferase expression in BFVL cells in a dose-dependent manner. BFVL cells were transfected with mock or indicated amount of p3.1 (+) -BTas. After 48 h the relative luciferase activities were analyzed. The relative expression of BTas in the transfected cells was monitored by WB assay (bottom). B: BFV3026 infection stimulates luciferase expression in BFVL cells in a dose-dependent manner. BFVL cells were mock-infected or infected with BFV3026-infected Cf2Th cells at different TCID50 as shown. After 48 h, cell lysates were prepared and assayed for quantitative luciferase expression. BFV3026-infected Cf2Th were also analyzed by Western Blot assay using Gag antibody (bottom). The data shown in each column are the mean ± SD (error bars) of three independent experiments.
-
To verify the specificity of luciferase activation on BFV infection, the BFVL cells were infected with other viruses including Bovine Immunodeficiency Virus (BIV), Bovine Herpes Virus (BHV) and PFV. As shown in Fig. 2A, 0.1 MOI of BFV could increase the luciferase activity by about 23-fold, while other viruses including 0.1 MOI of BIV, BHV and HFV, only induce 1.8-fold, 1.2-fold and 1.5-fold increases respectively. The result indicates that BFV could induce luciferase activity increased significantly compared with BIV, BHV and PFV. In order to exclude the possibility that different viruses have different appropriate titers on viral infection, we infected BFVL cells with 2-fold diluted BFV and BIV virus stocks. We found BIV with higher titers still could not induce significantly higher levels of luciferase expression (Fig. 2B). These findings show that the BFVLTR promoter in the BFVL cell line is specific for BFV.
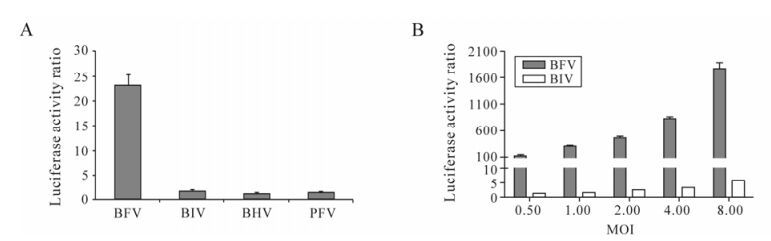
Figure 2. Specificity of BFVL cells for the detection of BFV infection. A: BFVL cells were infected with 0.1 MOI of BFV3026-infected Cf2Th cells and other laboratory strains, such as BIVR29, BHV, PFV. At 48 h post infection, cell lysates were assayed for quantitative luciferase expression. B: BFVL cells were infected with two-fold diluted BFV3026 and BIVR29 virus stocks. Each virus titer (MOI) is shown. At 48 h post infection, cell lysates were assayed for quantitative luciferase expression. Results shown represent the mean ± SD (error bars) of three independent experiments.
-
To gain an understanding of luciferase expression kinetics on BFV infection, BFVL cells were infected with BFV3026 at different MOI (0.01, 0.1, 1). The luciferase expression as a result of BFV3026 infection was evaluated at different time points post infection (12, 24, 36, 48, 60, 72, 96 h). As shown in Fig. 3, the level of luciferase activity in BFVL cells with different MOI BFV3026 infecting showed time-dependency and peaked at 84 h post infection. Although the luciferase signal gradually increased before 84 h post infection, the luciferase signal of BFVL cells induced by BFV3026 at a MOI of 1 and 0.1 starting from 24 h post infection, while the MOI of 0.01 was started from 60 h post infection. When BFVL cells were infected with BFV3026 at a MOI of 1, the luciferase expression showed a 3-fold increase relative to the control at 12 h post infection, however, it showed a 45-fold increase at 24 h post infection. Thus, within this initial period, there is a significant but non linear increase in the luciferase signal. The linear range of this kinetic curve was from 24 h to 84 h post infection. However, the linear ranges of the kinetic curves were from 12 h to 84 h and from 60 h to 84 h when BFVL cells were infected with BFV3026 at a MOI of 0.1 and 0.01, respectively. We also found the minimal detectable concentration of BFV was 0.01 MOI. Taken together, these results suggest that BFV3026 infection at the MOI of 0.1 could induce appropriate luciferase expression. Moreover, the time to detect the level of luciferase expression is at least 24 h and no later than 84 h. Although the changes in luciferase activity of BFVL peaked at 84 h. post infection, it was possible to differentiate infected and uninfected cells at 48 h post infection.

Figure 3. Luciferase activity in BFVL cells infected with BFV at different MOI. BFVL cells were mock-infected or infected with BFV3026 at MOI 0.01 (▼), MOI 0.1 (▲), and MOI 1 (■). The luciferase expressions were determined at 12, 24, 36, 48, 60, 72, 84 and 96 hours post infection. Results shown represent the mean ± SD (error bars) of three independent experiments.
-
To rapidly detect and quantitate BFV infectious titers, we determined the relationship between BFV concentration and luciferase activity. BFVL cells were infected with serial diluted BFV3026 virus stocks. The level of luciferase expression was analyzed 48 h post infection and a standard curve was established by plotting luciferase activity ratio relative to the control versus their corresponding virus MOI. The linear ranges of the virus concentrations to induce luciferase expression were from 0.06 to 8 MOI. The linearity of this curve (R2= 0.99) underlines the usefulness of BFVL-based assay for rapid quantification of BFV infectivity titers (Fig. 4).
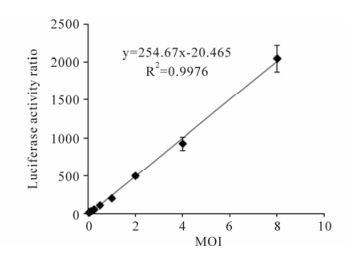
Figure 4. Quantification of BFV infection with BFVL-based assay. BFVL cells were inoculated with BFV3026 stock viruses in series diluted MOI (8, 4, 2, 1, 0.5, 0.25, 0.125, 0.0625 MOI) and luciferase expressions were measured at 48 h after the infection. A standard curve was established by plotting the luciferase activity ratio versus their corresponding virus titers. Results shown represent the mean ± SD (error bars) of three independent experiments.
-
We also examined the sensitivity of BFVL cells in comparison to the conventional CPE-based assay and another BFV indicator cell line BICL-based assay containing the EGFP reporter gene [14]. Cf2Th, BICL, and BFVL cells were simultaneously infected with different titers of BFV3026 virus stocks. At 48 h post infection, the infectious titers of the BFV virus stocks were determined based on the CPE-based assay, the BICL-based assay and the BFVL-based assay respectively. As shown in Fig. 5, the BFVL-based assay was about 10, 000-fold more sensitive than the CPE-based assay, and about 1, 000-fold more sensitive than the BICL-based assay by TCID50, regardless of the inoculated virus titers. This result suggests that the BFVL-based assay is the most sensitive assay for the detection of infectious BFV among the available assays.
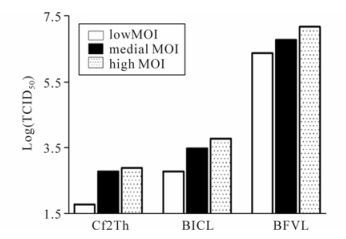
Figure 5. Comparison of the sensitivity of the BFVL-based assay with CPE-based and BICL-based assays for the detection of BFV infection. The titration of BFV3026 was examined by the conventional CPE-based assay by end-point dilution, BICL-based assay and BFVL-based assay. The virus stocks of 5 μL, 10 μL and 20 μL represent that virus titers that are low, medial and high respectively. Cf2Th, BICL, and BFVL were inoculated with different titers viruses. Forty-eight hours later, the virus titers were determined by the end-point method, BICL-based and BFVL based assays.
Generation of BFVL reporter cell lines
Specificity of BFVL cells for the detection of BFV infection
Luciferase expression kinetics in BFVL cells on BFV infection
The correlation of BFV titers and the luciferase activity in BFVL cells
Comparison of the sensitivity of the BFVL-based assay with the CPE-based and BICL-based assays
-
It is considered that FVs are nonpathogenic in naturally or experimentally infected animals [18]. Therefore, FVs are being exploited as retrovirus vectors for gene therapy since they have no disease association [7, 8, 20]. However, it is still a possibility that FVs may induce pathology in immunosuppressed hosts and so an integrated evaluation of the risks of FVs-based gene transfer is still necessary before its application. An easy and sensitive quantitative assay is required to further study the biological properties of FVs. The conventional CPE-based or syncytia formation assay by end-point dilution is limited because it cannot detect the FVs infection without syncytia formation. Moreover, the conventional CPE-based assay is time consuming, labor intensive and relatively insensitive. Thus, indicator cell line methods to monitor viral infection by exploiting the connection between the trans-activation of the viral LTR promoter and the viral transactivators have been developed. Among these developed indicator cell lines, luciferase-based indicator cell lines have several outstanding advantages compared with cell lines with other reporter genes and they have been used for high-throughput screening of anti-viral inhibitors.
In this study, we have constructed an indicator cell line, designated BFVL, for BFV titration that employs the firefly luciferase gene as the reporter gene on the downstream of the inducible BFVLTR promoter. We have demonstrated that BFVL cells could specifically respond to BTas activation but not to trans-activators of other viruses, such as Tas (data not shown). The luciferase induction in BFVL cells was specific to BFV infection but not to other viruses. The results of this study also showed that the BFVL-based assay was about 10, 000 times more sensitive than the CPE-based assay. Although our group has established an indicator cell line, designated BICL, which employs the EGFP gene as the reporter gene [14], this cell line was inconvenient for quantitation of the BFV infection. Meanwhile, the BFVL-based assay was 1, 000 times more sensitive than the BICL-based assay and could quantitate BFV infection rapidly and easily. In summary, the BFVL indicator cell line provides a rapid, easy, sensitive, accurate and quantitative method for monitoring and investigating BFV infection in vitro.
Viruses are obligate intracellular parasites and depend on cellular machinery for their efficient replication. BFV is able to activate the NF-κB pathway to enhance viral transcription like a group of viruses such as HIV-1 and HSV-1 [21, 27, 28]. However, in another aspect of FV-host interactions, IFP35 as a host cellular factor exerts an antiviral function by interfering with the transcription of viral genes via interaction with viral regulatory proteins. Thus, BFVL-based assay is indispensable and essential for determining activators and inhibitors on BFV infection since the molecular and immunological underpinnings of FV-host interactions are just beginning to be understood.
Despite the recent advances in the field, several issues remain to be clarified. The integration pattern along the human genome remains unclear. If such site specificity is determined, this will be beneficial for investigating the reason for the lack of tumorigenesis following long-term infection by FVs and streng-thening their potential use as gene transfer tools [2]. Furthermore, it would be interesting to elucidate the modes of superinfection of BFV with other viruses such as BIV and BHV. In our system, the BFVLTR promoter in the BFVL cell line was specific for BFV since other viruses could only induce low luciferase activities. However, inoculation with high titers of BIV resulted in slightly increased luciferase activities compared with the low titer. This result indicates that host cellular pathways may play a pivotal role in mediating the superinfection of BFV with other viruses, such as NF-κB pathway, since many viruses are able to influence the regulation of NF-κB in multiple and sophisticated ways [3, 21, 28].











 DownLoad:
DownLoad: