-
Cyanophages are viruses that infect cyanobacteria, and some cyanophages reportedly infect different genera of cyanobacteria (Gao E B, et al., 2009; Liu X Y, et al., 2006; Safferman R S, et al. 1963; Yoshida T, et al., 2006). The isolation and cultivation of additional cyanophages that infect different cyanobacteria is needed in order to gain a better understanding of the relationships between cyanophages and their hosts. Although there is evidence that viruses are an important feature of shallow freshwater lake microbial ecosystems and display remarkable morphological diversity, little is known about the precise control mechanisms of freshwater cyanophages that affect the growth and decline of bloom-forming cyanobacteria (Liu Y M, et al., 2006; Yoshida T, et al., 2006; Yoshida M, et al., 2008; Zhang Q Y et al., 2008 and 2012).
M. aeruginosa is one of the most common bloom-forming cyanobacteria found in eutrophic freshwaters (Wood S A, et al., 2012) and usually the dominant cyanobacterial genus in eutrophic lakes, but cultivable cyanophages that infect this genus are scarce (Tan X, et al., 2009; Yoshida T, et al., 2006). In China, Microcystis blooms occur in a large proportion of the aquatic environments, including lakes, reservoirs, and ponds, and often cause serious water management problems (Chen J, et al., 2009; Wu Z X, et al., 2007). Many studies focus on the physical and chemical factors that affect the ecology of Microcystis blooms, but controlling these algal blooms is difficult during warm months (Davis T W, et al., 2009; Oliver R L, et al., 2000; Yoshida T, et al., 2006; Sengco M R, 2009; Yang F, et al., 2013). Some biological factors, such as algicidal bacteria and cyanophages, have been suggested as possible regulators of cyanobacterial blooms (Brussaard C P, 2004; Yang F, et al., 2013; Zhang Q Y, et al., 2009).
The establishment of laboratory phage-host systems would be invaluable to the study of the interactions between cyanophages and their bloom-forming hosts and could help develop methods for controlling Microcystis blooms. In this study, we describe the isolation, cultivation, and partial characterization of a freshwater cyanophage that specifically infects the bloom-forming cyanobacterium M. aeruginosa, which is found in Lake Dianchi, a eutrophic lake in southwestern China. This cyanophage was tentatively named MaMV-DC (Microcystis aeruginosa myovirus from Lake Dianchi) based on its host, morphological features, and location where it can be isolated.
HTML
-
Four cyanobacterial species (21 strains) were used to isolate the MaMV-DC cyanophage and determine its host range (Table 1). Strains were obtained from the Freshwater Algae Culture Bank, Institute of Hydrobiology, Academy of Sciences, Wuhan, China. Each cyanobacterial strain used in this research was clonally grown in BG-11 medium (Gao E B, et al., 2009) at 25 ℃ under a 14-hour light/10-hour dark cycle using 35 μmol photons m-2 s-1 with cool white fluorescent illumination.
Species Strain Susceptibility Microcystic aeruginosa FACHB-524 a + FACHB-526 - FACHB-905 a - FACHB-915 a - FACHB-1179 a - FACHB-1217 - FACHB-1291 a - HAB1801 a - HAB0334 - Anabaena spirioides HAB1211 - FACHB-709 - FACHB-1198 - FACHB-1199 - FACHB-1219 - FACHB-1246 - HAB0502 - HAB0508 - PCC7120 - Anabaena oumiana HAB0984 - Synechococcus sp. FACHB-1061 - Botryococcus sp. FACHB-1108 - aStrains used during the first isolation process. +, sensitive; -, not sensitive. Table 1. Cyanobacterial strains and their susceptibilities to MaMV-DC.
-
Freshwater samples were collected from Lake Dianchi, the sixth largest freshwater lake in China, in southwest Kunming City. Centrifugation and filtration were performed at our laboratory to remove zooplankton, phytoplankton, bacteria, and cellular debris from the sample. The sample was centrifuged at 8000 g for 10 minutes at 4 ℃ and filtered through a 0.45-μm pore filter (Millipore, Bedford, MA, USA). Aliquots of the filtrate (200 μL) were added to 24-well plates containing 1 mL of exponentially growing cultures of M. aeruginosa strains; fresh BG-11 medium was used as the negative control. Samples were incubated at 25 ℃ under the previously described lighting conditions. Cultures were visually examined daily for visible changes in cyanobacterial morphology using light microscopy.
The cultured M. aeruginosa FACHB-524 cells were treated with successive plaque purification cycles in order to develop the clonal and lytic MaMV-DC cyanophage. Briefly, the supernatant of the lysed cells was serially diluted with BG-11 medium and each 20 μL dilution of supernatant was mixed with 0.5 mL of the exponentially growing M. aeruginosa FACHB-524 cultures (approximately 9.5×108 cells mL-1). Following incubation at 25 ℃ for 1 hour, 2.5 mL of top agar (0.4% molten BG-11 agar medium) containing the supernatant dilution and M. aeruginosa cells was spread onto 1% BG-11 agar plates. M. aeruginosa FACHB-524 cultures without the cyanophage suspension served as the negative control. All plates were incubated under the previously described conditions and monitored daily. After plaque formation, isolated plaques were selected and used to infect fresh cyanobacterial cultures. A new round of cyanophage infection and plaque isolation was conducted, as described above, until homogeneous plaques appeared.
-
The host range of MaMV-DC was determined by adding 0.2 mL aliquots of the MaMV-DC suspension to 0.8 mL of exponentially growing cyanobacterial cultures (Table 1). Cyanobacterial cultures without the MaMV-DC suspension served as the negative control. All cultures were monitored daily for host cells lysis using light microscopy. Cyanobacterial strains that did not lyse after 14 days of incubation were defined as insusceptible hosts for MaMV-DC.
-
One-step growth experiments were performed to evaluate the development and burst size of MaMV-DC. Ten mL of the MaMV-DC suspension (cyanophage titer 6×108 infectious units) was added to 100 mL of exponentially growing M. aeruginosa FACHB-524 cultures (approximately 107 cell mL-1). Cultures were incubated for 6 days in an illuminating incubator. After inoculation, the density of the cyanobacterial cells was determined daily by direct counting using a hemocytometer (Shanghai Medical Optical Instrument Plant, Shanghai, China), and the MaMV-DC titer was estimated using the most-probable-number technique, as previously described for the PaV-LD cyanophage (Gao E B, et al., 2009).
The lysis curves for MaMV-DC were established as described above. M. aeruginosa FACHB-524 cultures were monitored daily by measuring the optical density of the wavelength at 680 nm (OD680) after inoculation with the MaMV-DC suspension or an autoclaved MaMV-DC suspension.
-
Cyanophage particles from M. aeruginosa FACHB-524 cultures were purified by sucrose density gradient centrifugation (20% -60%). The cyanophage lysates were treated with chloroform and centrifuged to remove cell debris, and the cyanophage particles were concentrated using polyethylene glycol 8000 (9.3% [wt/vol]). The concentrates were centrifuged on a discontinuous sucrose density gradient at 24, 000 rpm for 40 minutes (Beckman Optima L-90 K, SW 41 swing-out rotor; Beckman, Fullerton, CA). The resulting bands in the 30% -60% range were separately collected and washed twice with sterile phosphate-buffered saline (PBS). The purified cyanophage particles were resuspended in sterile PBS and transferred to an Eppendorf tube for negative stain electron microscopy and genomic DNA extraction.
-
The purified cyanophage particles were negatively stained with 2% uranyl acetate (w/vol) and observed at 80 kV using a JEM-1230 transmission electron microscope (JEOL, Tokyo, Japan). Cyanophage DNA was prepared for restriction endonuclease cleavage analysis according to the method described by Gao et al (2012). Briefly, the purified cyanophage particles were treated with sodium dodecyl sulfate-proteinase K digestion, and genomic DNA was subjected to phenol-chloroform extraction and ammonium acetate-isopropanol precipitation.
-
To assess the sensitivity of the MaMV-DC nucleic acids to restriction enzymes, the extracted genomic DNA was prepared for restriction enzyme treatment. The nucleic acid was incubated overnight with two restriction enzymes—Pst Ⅰ (12 U/μL; TaKaRa, Kyoto, Japan) and Sac Ⅰ (12 U/μL; TaKaRa)—at 37 ℃ according to the manufacturer's recommendations. The restriction enzyme-digested products were electrophoresed in 1% agarose (wt/vol) and visualized using a gel analysis system (Syngene G: Box; Cambridge, UK) after staining with ethidium bromide (Yoshida T, et al., 2006).
The purified MaMV-DC particles were mixed with an equal volume of 2×SDS-PAGE loading buffer (100 mmol/L Tris-HCl [pH 6.8], 4% SDS, 0.2% bromphenol blue, 2% 2-mercaptoethanol, 20% glycerol), boiled for 5 minutes, and loaded onto 15% SDS-polyacrylamide gels. The proteins were separated at a constant voltage (120 V) and visualized using the Coomassie brilliant blue staining method.
Cyanobacterial strains and growth conditions
Isolation and cultivation of cyanophages
Host range test
Growth experiments
Cyanophage purification
Electron microscopy and genomic DNA extraction.
Restriction enzyme treatment and protein structure analysis
-
The host range of MaMV-DC was tested on 21 strains of cyanobacteria (4 cyanobacterial species). MaMV-DC only infected and lysed M. aeruginosa FACHB-524, but did not infect 8 other strains of M. aeruginosa or the other tested cyanobacterial strains (Table 1). The results show that MaMV-DC demonstrates a relatively narrow host range.
-
Host cell lysis was observed in M. aeruginosa FACHB-524 cultures within 1 week during the first isolating process. The initial green color of the healthy cultures (Fig. 1A) visibly changed to light yellow (Fig. 1B). The abundance of normal M. aeruginosa cells (Fig. 1C) was markedly influenced by MaMV-DC infection, and the round cyanobacterial cells were destroyed and their number sharply reduced within 3 days after infection (Fig. 1D). After three rounds of isolation and propagation, homogeneous plaques with a diameter of approximately 0.5 cm formed on the lawns of the M. aeruginosa FACHB-524 cultures at 6 days after infection (Fig. 2B), instead of the variety of plaques that formed in the non-monoclonal phage suspension (Fig. 2A). These findings indicate that the clonal plaqueisolated virus, MaMV-DC, is an infectious agent.
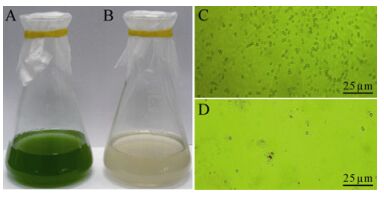

Figure 1. Micro-and macrographs of Microcystis aeruginosa FACHB-524 cultures. A: Macrograph of a normal culture. B: Macrograph of a MaMV-DC infected culture. C: Micrograph of a normal culture. D: Micrograph of a MaMV-DC infected culture. Scale bar=25 μm.
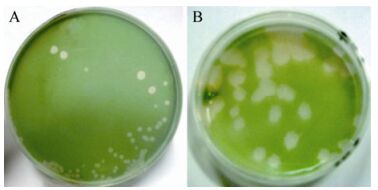
Figure 2. MaMV-DC formed plaques on the lawns of M. aeruginosa FACHB-524 at 6 days postinfection. A: Plaques in the second passage were formed by non-monoclonal phages. B: Plaques in the third passage were formed by clonal MaMV-DC.
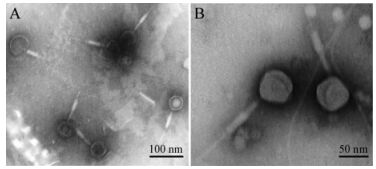
Transmission electron microscopic images of negatively stained MaMV-DC show that the phage-like particles have a tadpole-shaped icosahedral head (approximately 70 nm in diameter) and contractile tail (approximately 160 nm in length; Fig. 3). The morphological features of MaMV-DC are similar to those of T4-like phage Ma-LMM01, which only infects a toxic strain of M. aeruginosa and also has an isometric head (86 nm) and contractile tail (90 -209 nm) (Yoshida T, et al., 2006). Based on these morphological features, this cyanophage was classified in the Myoviridae family.
-
The growth kinetics of MaMV-DC were estimated using one-step growth experiments. The cell density of M. aeruginosa FACHB-524 began to decrease and the MaMV-DC titer began to increase at 1-day postinfection, indicating that MaMV-DC has a latent period of approximately 24 -48 hours. Within 6 days postinoculation, the virus titer increased to 1.02×109 from 6.01×106 infectious units mL-1, while the density of host cells decreased from 1.28 × 107 to 7 × 104 cells mL-1. Based on the increase in cyanophage abundance and the concurrent decline in host cell density, about 80 infectious units were estimated to have been released by each host cell infected with MaMV-DC (Fig. 4A). The lysis curves illustrate that host cell lysis occurred within 2-4 days after inoculation, while the healthy control cultures kept growing exponentially (Fig. 4B).
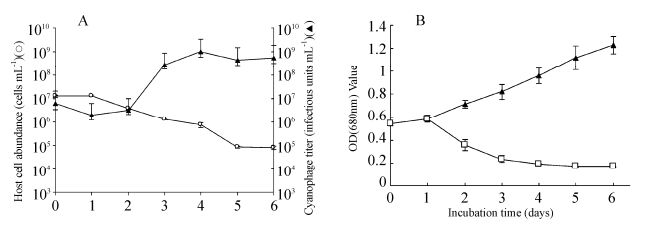
Figure 4. Changes in the number of.M. aeruginosacells and MaMV-DC infectious units. A) One-step growth curve of MaMV-DC. The two datasets represent the abundance of.M. aeruginosa FACHB-524 cells (O) and the cyanophage titer (▲). Each curve represents the average results from three experiments, and error bars indicate standard deviations. B) Lysis curves for MaMV-DC. Cyanobacterial cultures incubated with MaMV-DC (□) or inactivated MaMV-DC (▲) were monitored using a spectrophotometer (680-nm optical density). Each curve represents the average of three experiments, and error bars indicate the standard deviations.
The lysis curves illustrate that host cell lysis occurred within 2 -4 days after inoculation, while the healthy control cultures kept growing exponentially (Fig. 4B).
-
Four major structural protein bands were separated from the purified MaMV-DC particles using SDS-PAGE, demonstrating molecular weights of approximately 84, 47, 38, and 26 kD, respectively (Fig. 5A).
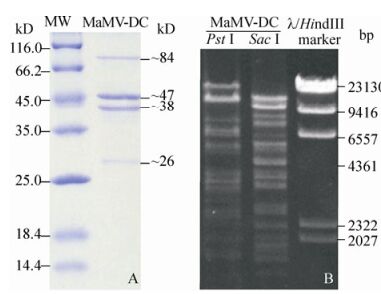
Figure 5. MaMV-DC structural proteins and genomic DNA that were digested by restriction endonucleases. A) Major MaMV-DC structural proteins (MaMV-DC) and molecular standards (MW). B) MaMV-DC DNA fragments digested using two restriction endonucleases (Pst Ⅰ and Sac Ⅰ). The λ/Hind Ⅲ marker was used as the molecular weight standard.
MaMV-DC genomic DNA was examined using restriction endonuclease analysis. The MaMV-DC genome was sensitive to the restriction enzymes Pst Ⅰ and Sac Ⅰ, suggesting that it may be poorly methylated. According to the endonuclease digestion profile, at least 15 and 14 bands were visualized using Pst Ⅰ and Sac Ⅰ, respectively, and their molecular weights ranged from 2 -24 kb (Fig. 5B).
Host specificity
Isolation, cultivation, and morphology of MaMV-DC
Growth and lysis characteristics
Protein patterns and restriction endonucleases digestion profile of MaMV-DC
-
In this paper, we present data on the isolation and characterization of a freshwater cyanophage that specifically infects the bloom-forming cyanobacteria M. aeruginosa. The diagnosis of MaMV-DC infection was based on cyanophage isolation, host range testing, cyanobacterial cell infection, electron microscopic observation, growth experiments, restriction endonuclease digestion, and protein structure analysis. A host-phage system for bloom-forming cyanobacteria was established and is expected to contribute to the isolation of other cyanophages from natural freshwaters. The results also pave the way for the future utilization of this phage as a biological agent to control cyanobacterial blooms.
Some freshwater cyanophages demonstrate a narrow host range, infecting only the cyanobacterial strains with which they are isolated. The results of our host range tests suggest that MaMV-DC has a fairly narrow host range as well. Only M. aeruginosa FACHB-524 was infected and lysed by MaMV-DC, while 20 other cyanobacterial strains, including 8 strains of M. aeruginosa, were not sensitive to MaMV-DC, indicating that MaMV-DC is strain-specific rather than species-specific. The host range specificity of the tailless PaV-LD cyanophage is correlated with the geographical location of the isolated host (Gao E B, et al., 2009). In contrast, MaMV-DC infects M. aeruginosa FACHB-524 isolated from Lake Donghu, but does not infect M. aeruginosa FACHB-905 isolated from Lake Dianchi where the MaMV-DC-containing freshwater sample was collected (Wu Z X, et al., 2007). It is generally believed that host specificity reflects specific attachment sites on host cells, but the molecular mechanisms involved in the host specificity have rarely been experimentally studied. One type of host enrichment strategy may be selected by cyanophages, resulting in a narrow host range (Wichels A, et al., 2002). Further studies on additional cyanobacterial hosts and cyanophages are needed to determine the essential factors involved in the sensitivities of cyanobacterial cells and cyanophages.
Cyanophage morphology is the basis for classification (Safferman R, et al., 1983; Suttle C A, et al., 1993). Based on the morphological features of the tail, most cyanophages isolated from either fresh or marine water are categorized as Siphoviridae, Podoviridae, or Myoviridae. Recently, a tailless cyanophage was isolated and sequenced, which provided novel insights into cyanophage morphology and cyanophage-host interactions (Gao E B, et al., 2012). MaMV-DC has a contractile tail and an icosahedral head, which are the common characteristics of myoviruses (Ackermann H W, 2007). Therefore, MaMV-DC is considered a member of the Myoviridae family.
The MaMV-DC and Ma-LMM01 cyanophages demonstrate similar hosts, morphological features, and major structural protein patterns (Yoshida T, et al., 2006). They both lyse bloom-forming M. aeruginosa in freshwater lakes, have an icosahedral head and contractile tail, and contain four major polypeptides (molecular weights of approximately 84, 47, 38 and 26 kDa, respectively). However, there are differences between MaMV-DC and Ma-LMM01 in terms of the cleavage maps formed by restriction endonucleases. Furthermore, although both of their hosts are M. aeruginosa, it is interesting and meaningful that MaMV-DC infects a non-toxic strain and Ma-LMM01 infects a toxic strain. To our knowledge, all myoviruses isolated from M. aeruginosa infect and lyse toxic strains only (Yoshida et al. 2008). The effect of these cyanophages on the growth and dynamics of M. aeruginosa has been investigated, and an impact on the shifts of toxic and non-toxic populations instead of on the total abundance was suggested (Yoshida et al. 2008). MaMV-DC as the first myovirus infected non-toxic M. aeruginosa is expected to contribute to our better understanding the dynamics of toxic and non-toxic strains of M. aeruginosa during Microcystis blooms and make it possible to compare the infection mechanisms of these cyanophages at the genomic level. For this purpose, we are performing a complete genome sequence analysis of MaMV-DC.
-
This work was supported by the National Natural Science Foundation of China (grant nos. 31072239, 31270213), the Knowledge Innovation Program of the Chinese Academy of Sciences (grant no. KSCX2-EW-Z-3), and the State Key Laboratory of Freshwater Ecology & Biotechnology Program (grant no. 2011FBZ12).
-
Conceived and designed the experiments: QY Zhang; Performed the experiments: T Ou, SH Li and XY Liao; Analyzed the data: T Ou and QY Zhang. Contributed reagents/materials: SH Li and QY Zhang; Wrote the paper: T Ou and QY Zhang.













 DownLoad:
DownLoad: