HTML
-
Pathogenic Escherichia coli (E. coli) is an important human and animal pathogen (Jenkins 2018; Kaper et al. 2004). Pathogenic E. coli strains were categorized into many pathotypes. Six defined E.coli pathotypes can cause intestinal infections, including enterotoxigenic (ETEC), Shiga toxin-producing/enterohemorrhagic (STEC/EHEC), enteropathogenic (EPEC), enteroaggregative (EAEC), enteroinvasive (EIEC), and diffusely adherent (DAEC) (Kaper et al. 2004). By contrast, four defined E. coli pathotypes, i.e. avian pathogenic E. coli (APEC), uro- pathogenic E. coli (UPEC), neonatal meningitis causing or meningitis-associated (NMEC/MAEC) and sepsis E. coli (SEPEC), were able to cause extra intestinal infections (Moulin-Schouleur et al. 2007).
Avian pathogenic E. coli (APEC) is the primary cause of colibacillosis, characterized by acute fatal septicemia or sub-acute fibrinous pericarditis, airsacculitis, salpingitis, and peritonitis, in poultry (Lutful Kabir 2010). Antibiotics have been not only used as antibacterial agents for the treatment of bacterial infections, but also as prophylactic and growth-promoting feed additives in breeding industry for a long time. Consequently, the wide usage of antibiotics has inevitably led to the emergence of multidrug-resistant bacteria (Suresh et al. 2018). Recent study showed that sixty-eight percent of APEC isolates carried at least 5 out of 8 antimicrobial resistant genes from broiler chickens in Jordan (Ibrahim et al. 2019). In a study in China, Li et al. reported that 87 APEC isolates showed 100% multi-drug resistant phenotypes, particularly against ampicillin, kanamycin, ciprofloxacin, levofloxacin, streptomycin, gentamycin, ofloxacin, norfloxacin, and ceftriaxone (Li et al. 2015). In addition, Liu et al. reported that APEC strains demonstrated high pathogenicity (15/26) and multidrug-resistant abilities (26/26) (Liu et al. 2019). Nowadays, multi-drug resistant E. coli has become a significant problem in veterinary, food and medical fields (Diarra and Malouin 2014).
Due to the multi-drug resistance among these E. coli stains, antimicrobial treatment became difficult. Some E. coli vaccines, including whole inactivated vaccine and subunit vaccine, have also been used to control coliba- cillosis. However, the vaccines were usually only effective against single serotype of strains. In fact, it is difficult to develop effective vaccines to preven and control all the serotype pathogenic E. coli (Ebrahimi-Nik et al. 2018). Broadly protective vaccines against all or multi-pathotypes E. coli are still not available (Nesta and Pizza 2018).
Development of eco-friendly, effective and safe alternatives to antibiotics is imperative. In comparison to other treatment solutions, phages have multiple advantages, including ubiquity (Johnson et al. 2008), specificity, broad antimicrobial effects on several different serotypes strains and less interference to the normal flora (Clark and March 2006). Phages have been reconsidered as an alternative option against bacterial infections. For example, phages were used as alternatives to antibiotics for treating human infected with Pseudomonas aeruginosa and Acinetobacter baumannii (Chan et al. 2018; Schooley et al. 2017) and animals infected with E. coli (Chibani-Chennoufi et al. 2004). In addition, engineered mycobacteriophage cocktail have been used for the treatment of a patient infected with drug-resistant Mycobacterium abscessus (Dedrick et al. 2019).
However, narrow host-range and phage-resistant bacteria remain to be the main problems for phage therapy (Haq et al. 2012; Hermoso et al. 2007). Currently, two main approaches were used to solve the problems by developing a phage cocktail or generating genetic recombinant phages (Chan et al. 2013; Mahichi et al. 2009; Yu et al. 2018). For example, T4-like phages can recombine with a recombination plasmid containing the gene 37-38 segment from T2- like or T6-like phages (Tetart et al. 1998). These engineered phages thus acquired host range of the donor phages and can infect genetically distant host bacteria. In T4-like phages, the receptor-binding tip of long tail fiber determined the specificity and host range, which is encoded by the C-terminal domain of gene 37 (Bartual et al. 2010; Chen et al. 2017). Our previous research also showed that the recombinant T4-like phages can infect eight additional bacteria, suggesting the expansion of host range (Chen et al. 2017).
This study aims to customize chimeric T4-like phages for specific clinical pathogenic E. coli using a T4-like phages homologous recombination platform established by our laboratory (Chen et al. 2017). The basic biological characteristics and genetic stability of recombinant phage were then examined and compared with the parental donor phages. Our study suggested that the engineered phage WGqlae had a significant antimicrobial effect against clinical E. coli strains in planktonic and biofilm forms in vitro.
-
All 113 E. coli strains used in this study are listed in Supplementary Table S1. Among them, 104 E. coli strains isolated from the brains of ducks with septicemia and neurological clinical symptoms in Eastern China in previous study, were included (Chen et al. 2017). The remaining 9 strains were selected from the collection in our laboratory. These E. coli strains were used to determine the host range of phages. Phages WG01 and QL01 were used as the parental phages for homologous recombination, while E. coli strains DE017 and DE205B were used as host strains of the two phages for amplification, respectively (Chen et al. 2017).
Luria-Bertani (LB) liquid medium were used to cultivate bacterial strains. When necessary, 100 μg/mL ampi- cillin was added. LB 0.5% soft agar plates and LB 1.5% agar plates were used for phages plaque assay. Tryptic soy broth liquid medium (TSB, Merck, Germany) were used to culture E. coli biofilm. SM buffer (50 mmol/L Tris-HCl [pH 7.5], 100 mmol/L NaCl, 10 mmol/L MgSO4 and 0.01% gelatin) was used for the dilution and storage of the phages. Phage cultures were clarified by centrifugation at 5000 ×g, 4 ℃ for 10 min, then the supernatants were filtered through 0.22 μm filters (Merck Millipore, Germany) and stored at 4 ℃. Phosphate Buffer Saline (0.1 mol/L Na2HPO4, 0.15 mol/L NaCl2, pH 7.2) was used to wash and dilute E. coli strains.
-
We selected six diverse DNA fragments in gene 37 from QL01 as the recombination regions to replace the corresponding fragments in WG01 (Bartual et al. 2010; Chen et al. 2017). Six different fragments of gene 37 (i.e., QLae, QLbe, QLce, QLde, QLfe and QLm) were amplified (Fig. 1) and fused using the primers listed in Supplementary Table S2, with WG01 and QL01 phages as the DNA templates. Plasmids construction was performed as described previously (Yoichi et al. 2005). In brief, the PCR products were digested with BamH I and EcoR I, followed by insertion into plasmid vector pUC118 (Takara, Shiga, Japan), generating six different recombinant plasmids, pUC118ae, pUC118be, pUC118ce, pUC118de, pUC118fe and pUC118m.

Figure 1. Sketch map of gene 37 in QL01 and the results of the isolated chimera phages. Different color fragments from QL01 for WG01 recombination. Four types of recombinant phages, namely, WGqlae, WGqlbe, WGqlce and WGqlde, were isolated from DE205B, the host of QL01. The symbol X indicate that no chimeric phages were isolated with DE205B by infecting the plasmid-containing cell with phage WG01.
-
Homologous recombination of T4-like phages was performed as described previously (Chen et al. 2017; Yoichi et al. 2005) with minor modifications. Briefly, competent strain DE017 were transformed with the recombinant plasmid, pUC118ae, pUC118be, pUC118ce, pUC118de, pUC118fe and pUC118m, respectively. The DE017 transformant strains were cultured in 5 mL LB liquid medium with 50 ig/mL ampicillin at 37 ℃ until OD600 0.4, then the phage WG01 was added. After purification and fil- teration, recombinant phages were isolated using E. coli DE205B (susceptible to QL01, but resistant to WG01) as host bacteria by double-layer plate method (Adams 1959). Recombinant phages were purified by plaque assay. PCR and Sanger sequencing analysis of gene 37 from the recombinant phages was carried out using primers Fcheck/ Rcheck (Chen et al. 2017).
-
The recombinant phages, WGqlae, WGqlTbe, WGqlTce, WGqlde, and the parent phages WG01 and QL01 were prepared by double-layer plate method (Adams 1959). A total of 113 avian E. coli strains were used as the host bacteria to determine the host ranges (Supplementary Table S1) using spot test with minor modifications (Chen et al. 2016). In brief, bacterial cultures at exponential phase (100 μL) were spread on an LB agar plates. Each of six phage suspensions (10 μL at 109 PFU/mL) was spotted on the surface of the bacteria lawn. The plates were examined for lysis after 8 h incubation at 37 ℃.
-
Thermo and pH stability of phages was evaluated using a previously described protocol with minor modifications (Laemmli 1970). Phage suspensions (100 μL) were mixed with SM buffer (900 μL) in 1.5 mL sterile Eppendorf tubes, and were kept at different temperatures (i.e. 30-60 ℃) for 1 h. The survival rates of phages were determined by double-layer method as described above.
To determine the pH stability of phages, 100 μL phage suspension was added to 900 μL SM buffer with a gradient of pH values (range from 4 to 11), followed by incubation at 37 ℃ for 1 h. Similarly, double-layer plating method was used to examine the survival rates of phages.
-
Multiplicity of infection (MOI) was defined as the ratio of viral particles to potential host cells. Optimal MOIs of phage WG01, QL01 and WGqlae were determined by using standard protocol with minor modifications (Lu et al. 2003). Phages were diluted to 104-109 PFU/mL. Log phase host bacteria were adjusted to 1 × 108 CFU/mL (Table 1). Phages at different titer (100 μL, MOI = 0.0001, 0.001, 0.01, 0.1, 1, 10) and 100 μL 1 × 108 CFU/mL log phase host bacteria were added to 1.5 mL sterile Eppendorf tubes containing 800 μL LB medium, followed by incaution at 37 ℃ for 3.5 h with shaking (60 rpm). Bacteria cultures (phage-free) and phage cultures (cell-free) were used as control groups in this experiments. All assays were conducted in duplicate. The MOI that generated the highest phage titer within 3.5 h was considered as optimal MOI.
MOI Phages (PFU/mL) Bacteria (CFU/mL) Bacteriophage titer after 3.5 h (PFU/mL) Proliferation times WG01 QL01 WGqlae WG01 QL01 WGqlae 0.0001 1 × 103 1 × 107 2.82 × 105 3.27 × 105 1.77 × 106 282 327 1770 0.001 1 × 104 1 × 107 3.86 × 106 4.64 × 107 1.53 × 106 386 4640 153 0.01 1 × 105 1 × 107 5.2 × 106 2.47 × 107 1.31 × 107 52 247 131 0.1 1 × 106 1 × 107 3.57 × 107 4.58 × 107 2.21 × 107 35.7 45.8 22.1 1 1 × 107 1 × 107 2.87 × 107 3.26 × 107 3.29 × 107 2.87 3.26 3.29 10 1 × 108 1 × 107 2.4 × 107 2.03 × 107 2.91 × 107 0.24 0.20 0.29 Table 1. Comparison of MOIs between recombinant WGqlae and parental phage WG01 and QL01.
-
Latent time and burst size of phage WGqlae were determined by one-step growth curve as previous described (Pajunen et al. 2000). 10 mL Mid-exponential phase E. coli DE205B culture (2 × 108CFU/mL, OD600 = 0.4) were mixed with 10 mL 1 × 107 PFU/mL bacteriophage suspensions. Allow the phages to specifically adsorb to the host bacteria for 10 min at 37 ℃. Then, after removing unabsorbed phages by centrifuging at 5000 rpm for 10 min, the precipitated pellet was resuspended in 20 mL fresh LB medium, followed by incubation at 37 ℃. Samples were taken out at 10-min intervals for 100 min, and 1% chloroform (final concentration) was added to release the intracellular phage for phage titration using the doublelayer agar plate method.
-
To evaluate the genetic stability of recombinant phage WGqlae, the 1st, 10th and 20th generation WGqlae were collected. Gene 37 was amplified by primers Fcheck/ Rcheck (Chen et al. 2017). PCR products were sequenced by Suzhou genewiz biotechnology Co., Ltd. (Suzhou, China).
-
Lytic activity of phages WGqlae was evaluated by OD600nm values of the host bacteria every 2 h at the MOI of 0.1, 1 and 10 using 96-well microtiter plates. Each of phage group was comprised of 100 μL host bacteria (2 × 108 CFU/mL, DE205B or DE192) and 100 μL diluted phage lysate (2 × 107-2 × 109 PFU/mL), and the assay was conducted in quadruplicates. Control group was consisted of equal volume of host bacteria suspension and fresh LB liquid medium. The 96-well microtiter plates were incubated at 37 ℃ for 24 h, and the OD600 values were measured at 2 h intervals using an ELISA microplate reader (Biotek, VT, USA).
-
Quantitative detection of bacterial biofilm was performed as described previously with minor modifications (Stepa-novic et al. 2000, 2004). Briefly, E. coli DE192 and DE205B were incubated in TSB liquid medium containing 1%, 2% and 3% glucose at 37 ℃ for 24 h, 48 h, 72 h, 96 h and 120 h, respectively. Supernatants were then slowly decanted, and washed three times with 200 μL PBS. After air drying, each well was added 200 μL methanol to fix for 15 min. After discarding methanol and air drying, each well was added 200 μL 1% crystal violet and incubated for 5 min. After being washed three times with PBS, each well was added 100 μL acetic acid (33% v/v) to dissolve the crystal violet, and was placed in 37 ℃ incubator for 30 min. Subsequently, the OD595nm value was measured by a microplate reader.
-
Fresh host bacteria DE192 and DE205B (100 μL, 2 × 108 CFU/mL) were added to 96 well plates. Before culture, WGqlae group was added 100 μL phage WGqlae (2 × 108 PFU/mL); polymyxin B group was added 100 μL polymyxin B (4 mg/L); and WGqlae + Polymyxin B group was added 50 μL phages (4 × 108 PFU/mL) and 50 μL polymyxin B (8 mg/L, minimum inhibitory concentration was 2 mg/L for DE192 and DE205B). Control group were added equivoluminal TSB medium. Culture condition and quantitative detection of bacterial biofilm were performed as mentioned above.
-
Escherichia coli DE192 and DE205B were incubated to form biofilm in TSB liquid medium at 37 ℃ for 48 h and 72 h, respectively. Then, WGqlae group was added 200 μL phage WGqlae (1 × 108 PFU/mL); polymyxin B group was added 200 μL polymyxin B (1 × 108 PFU/mL), and WGqlae + Polymyxin B group was added 100 μL phages (2 × 108 PFU/mL) and 100 μL polymyxin B (8 mg/L). Control group was added equivoluminal TSB medium. Culture condition and quantitative detection of bacterial biofilm were performed as mentioned above.
-
All statistical analyses in this study were carried out with the GraphPad Prism 5 software package. Mean difference of control and treat groups of planktonic or biofilm assay were analyzed by student's t test. P value ≤ 0.05 was considered statistically significant.
Bacteria, Bacteriophages and Growth Conditions
Construction of the Plasmids with Diverse Recombination Regions
Homologous Recombination and Isolation of Recombinant Phages
Host Ranges of Recombinant Phages
Thermo and pH Stability of WG01 Derivatives
Optimal Multiplicity of Infection
One Step Growth Curve of Recombinant Phage WGqlae
Genetic Stability of Recombinant Phage WGqlae
Lytic Capacity of Phages WGqlae against E. coli in Planktonic Form
Quantification of Bacterial Biofilm
Inhibiting Efficacy of Phage WGqlae, Polymyxin B, and Their Combination on Biofilm Formation
Efficacy of Phage WGqlae, Polymyxin B and Their Combination on the Clearance of Formed Biofilm
Statistical Analysis
-
We randomly selected six different DNA fragments, QLae (709-2775 nt), QLbe (949-2775 nt), QLce (1189-2775 nt), QLde (1429-2775 nt), QLfe (1669-2775 nt) and QLm (1363-2383 nt) of gene 37 in QL01 to replace the corresponding fragments of WG01 using a gene manipulation experiment (Fig. 1).
Four recombinant phages (WGqlce, WGqlbe, WGqlae and WGqlde) were isolated from clear plaques on doublelayer plates with E. coli DE205B, one of host bacteria of phage QL01 (but not the host of WG01) (Fig. 2). Sequencing analysis of gene 37 confirmed the generation of chimeric phages WGqlce, WGqlbe, WGqlae and WGqlde. The gene 37 of WGqlce did not mutate in the process of homologous recombination. However, nucleic acid mutations occurred in gene 37 of WGqlae, WGqlTbe and WGqlTde, causing 14, 29 and 1 amino acid substitutions, respectively (Supplementary material S3).
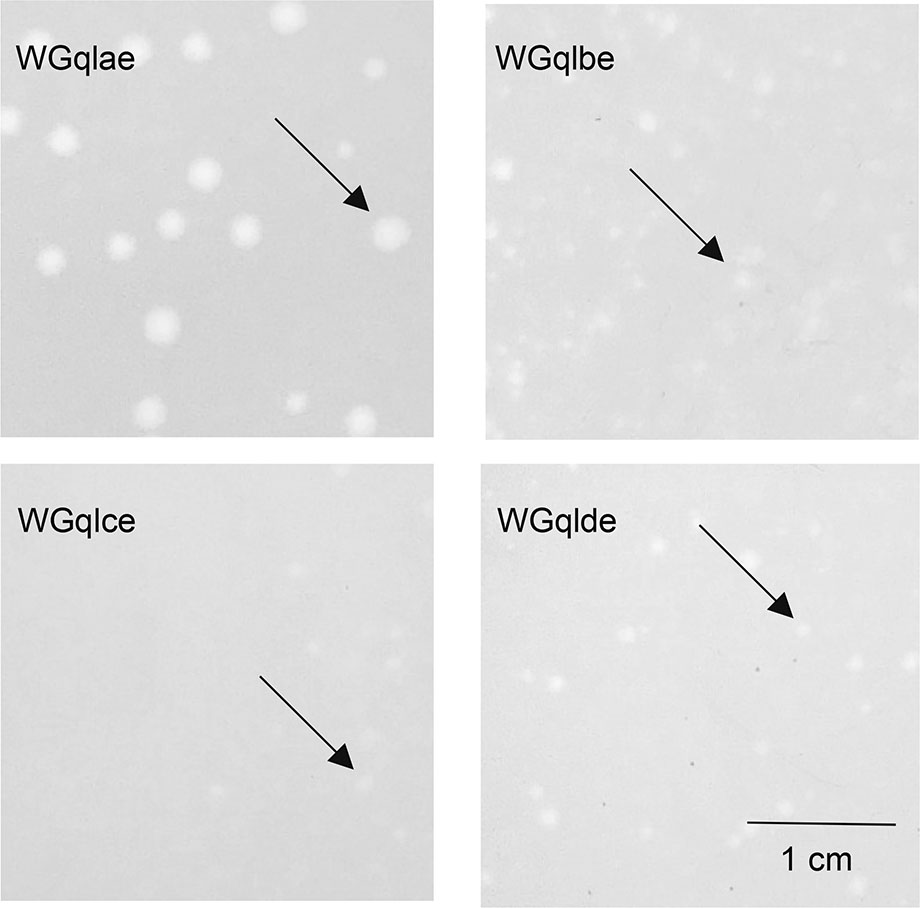

Figure 2. Plaque morphology of recombination phages. Phages WGqlae, WGqlbe, WGqlce and WGqlde were grown on E. coli DE205B in double agar LB plates for 6 h. WGqlce and WGqlbe formed clear and round plaques approximately 0.1 cm in diameter. WGqlae formed clear and round plaques approximately 0.3 cm in diameter. WGqlde formed clear and round plaques approximately 0.2 cm in diameter.
-
Host ranges of the phages WG01, QL01 and four WG01 derivatives were determined against 113 E. coli strains. Phages WG01 and QL01 lysed 20 (17.70%) and 50 (44.25%) of the 113 E. coli strains, respectively. While phages derivatives WGqlae (with AA 237-925 of QL01 gp37), WGqlbe (with AA 317-925 of QL01 gp37), WGqlce (with AA 397-925 of QL01 gp37) and WGqlde (with AA 477-925 of QL01 gp37) lysed 49 (43.36%), 52(46.02%), 52 (46.02%) and 52 (46.02%) of the 113 E. coli strains, respectively (Fig. 3, Supplementary Table S1). Among them, WGqlde, which was only replaced the downstream 1429-2772 bp in gene 37 of QL01, can lyse the most part of host strains of QL01. This result suggested that this domain of gp37 mainly determined the host range, which is consistent with the finding that C-terminal domain determined the receptor specificity. WGqlae, however, can infect four additional host bacteria, DE011, DE037, DE061 and DE077, in comparison to the parental phages, which suggested the expansion of host-range. Therefore, the recombination phage WGqlae was chosen to analyze the basic characteristic and antibacterial effect.

Figure 3. Host ranges analysis of recombinant and their parental phages. Phages derivatives WGqlae, Wgqlbe, Wgqlce and Wgqlde lysed 49, 52, 52 and 52 strains of the 113 E. coli strains, respectively (Only 64 strains were showed in this map, because other 47 strains were not susceptible to any of these phages). WGqlae can infect 4 additional host bacteria DE011, DE037, DE061 and DE077 in comparison to the parental phages.
-
Thermo stability test was carried out at pH 7 to investigate thermo stability of phages. The results showed that more than 60% of phage WGqlae remained active after incubation at 30 ℃, 40 ℃ and 45 ℃ for 1 h, which were similar to the results from WG01 and QL01. In contrast, less than 2% of three active phage particles were observed at 60 ℃ (Fig. 4A). The results showed that the thermo stability of the WG01 derivative WGqlae was nearly the same as donor phage QL01.
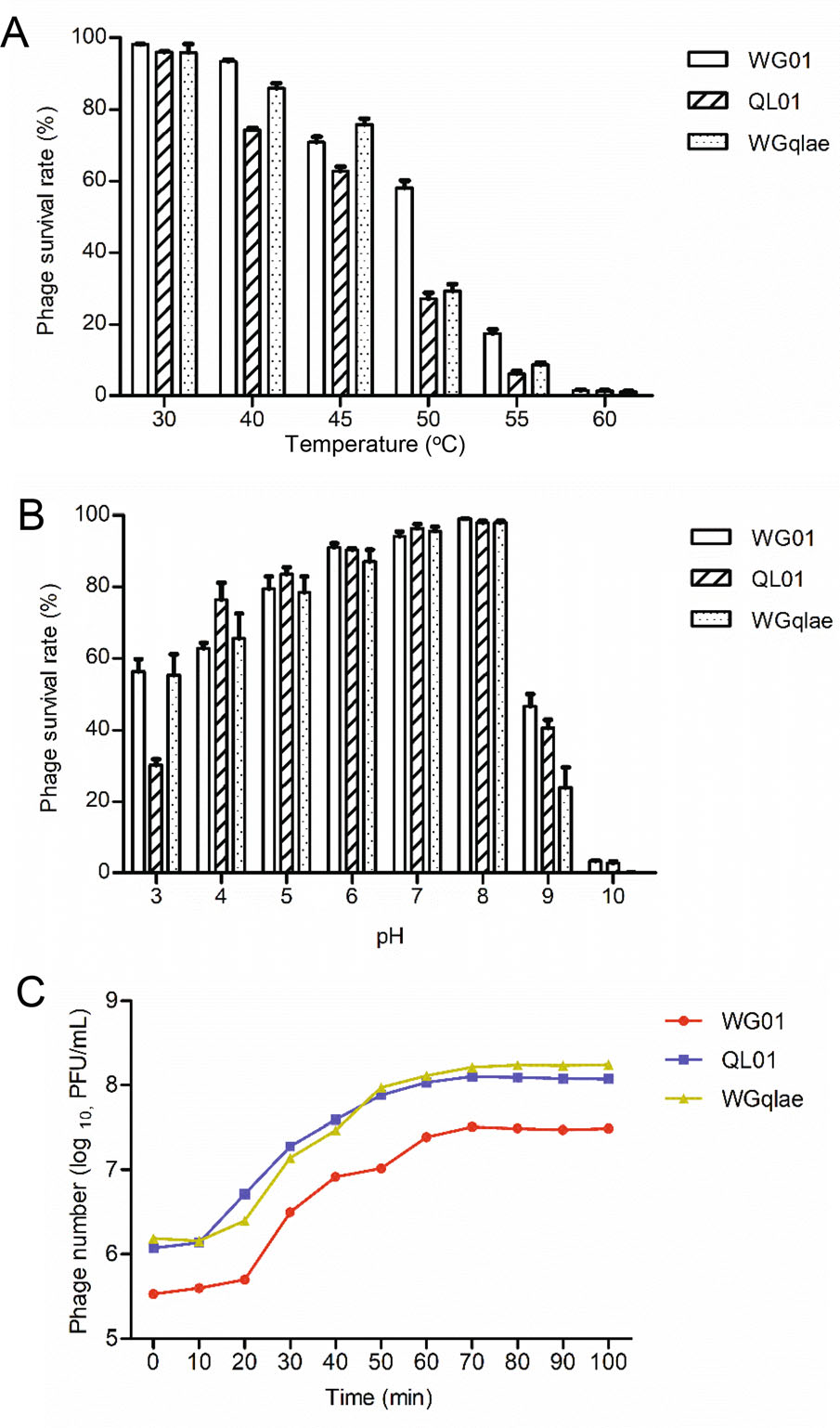
Figure 4. Biological properties of the recombinant phage WGqlae and its parental phages. A Thermo stability of recombinant phage WGqlae and the parental phages WG01 and QL01. B The pH stability of recombinant phage WGqlae and its parental phages WG01 and QL01. C One-step growth curve of the recombinant phage WGqlae and its parental phages WG01 and QL01. All the values are the means of 2 determinations.
The pH stability of phages was tested at 37 ℃ for 1 h. Phage WGqlae, WG01 and QL01 showed similar pH stability. In detail, they all showed the highest stability at pH 7, and they demonstrated similar survival rate (> 50%) at pH 5-9 (Fig. 4B). For all three phages, the phage titers decreased sharply at the acidic (pH = 4) or alkaline (pH = 10, 11) conditions.
-
MOI was defined as the ratio of viral particles to potential host cells. The optimal MOI of parental phage WG01 and QL01 were both 0.001. After incubation for 3.5 h, phage WG01 and QL01 multiplied 386 times and 4640 times, respectively. While the optimal MOI of phage WGqlae was 0.0001, and WGqlae increased by 1770 times after 3.5 h proliferation. The optimal MOI of recombinant phages WGqlae was lower than those of parental phage WG01 and QL01 (Table 2).
Time Control DE192 (OD595) DE205B (OD595) LB TSB 1% G-TSB 2% G-TSB 3% G-TSB LB TSB 1%G-TSB 2%G-TSB 3%G-TSB 24 h 0.29 0.49 0.53 0.53 0.53 0.45 0.63 0.41 0.46 0.38 0.34 48 h 0.34 1.06 3.81 2.02 1.95 1.58 1.76 1.57 1.50 1.18 1.34 72 h 0.32 1.43 2.38 2.37 2.38 1.59 1.37 1.86 1.14 0.90 1.04 96 h 0.31 0.63 0.82 1.06 1.02 1.05 0.92 0.67 0.67 0.68 0.61 120 h 0.33 0.78 1.16 1.47 1.24 1.25 1.21 0.89 0.74 0.78 0.88 Table 2. Optimal biofilm formation conditions of DE192 and DE205B.
-
One-step growth experiments were performed to compare latent time period and burst size of the phage WGqlae and the parental phages. One-step growth curve consisted of three phases-latent, log and stationary phases. Latent time period of the phage WGqlae was 10 min, and the burst size was about 110 phages/cell (Fig. 4C). In comtrast, phage WGqlae showed similar to phage QL01 in one-step growth experiments.
-
The gene 37 of phage WGqlae in 1st, 10th and 20th generation did not showed any mutations, suggesting that recombinant phage WGqlae was genetically stable.
-
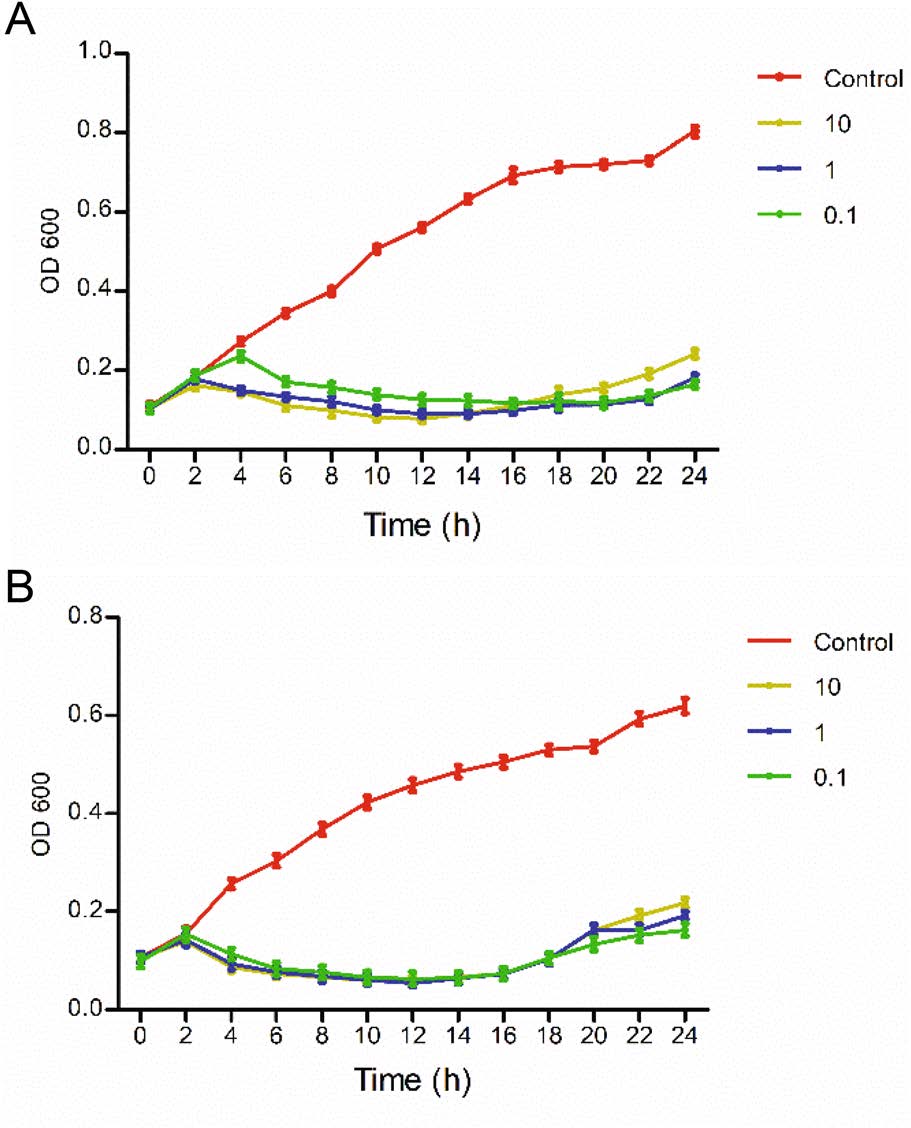
The OD600 values of the phage groups were significantly lower than that of the control group in 24 h, indicating that the derivative WGqlae inhibited bacteria proliferation. The OD600 values of the control group of both host bacteria DE192 and DE205B increased continuously in 24 h. All OD600 values of three phage groups (MOI 10, 1 and 0.1) began to decrease after a short increase in the first 2 h, and it did not increase until 16 h. In the first increased phase, the smaller MOI values were correlated with the more obvious increased trend, but in the second increased phage, the larger MOI values were associated with the more obvious decreased trend (Fig. 5).
-
Our results showed that optimal medium for biofilm formation of host bacteria DE192 and DE205B was TSB medium without glucose, and optimal culture time of DE192 and DE205B was 48 h and 72 h, respectively. Therefore, the optimal medium and conditions were subsequently used in the following biofilms test.
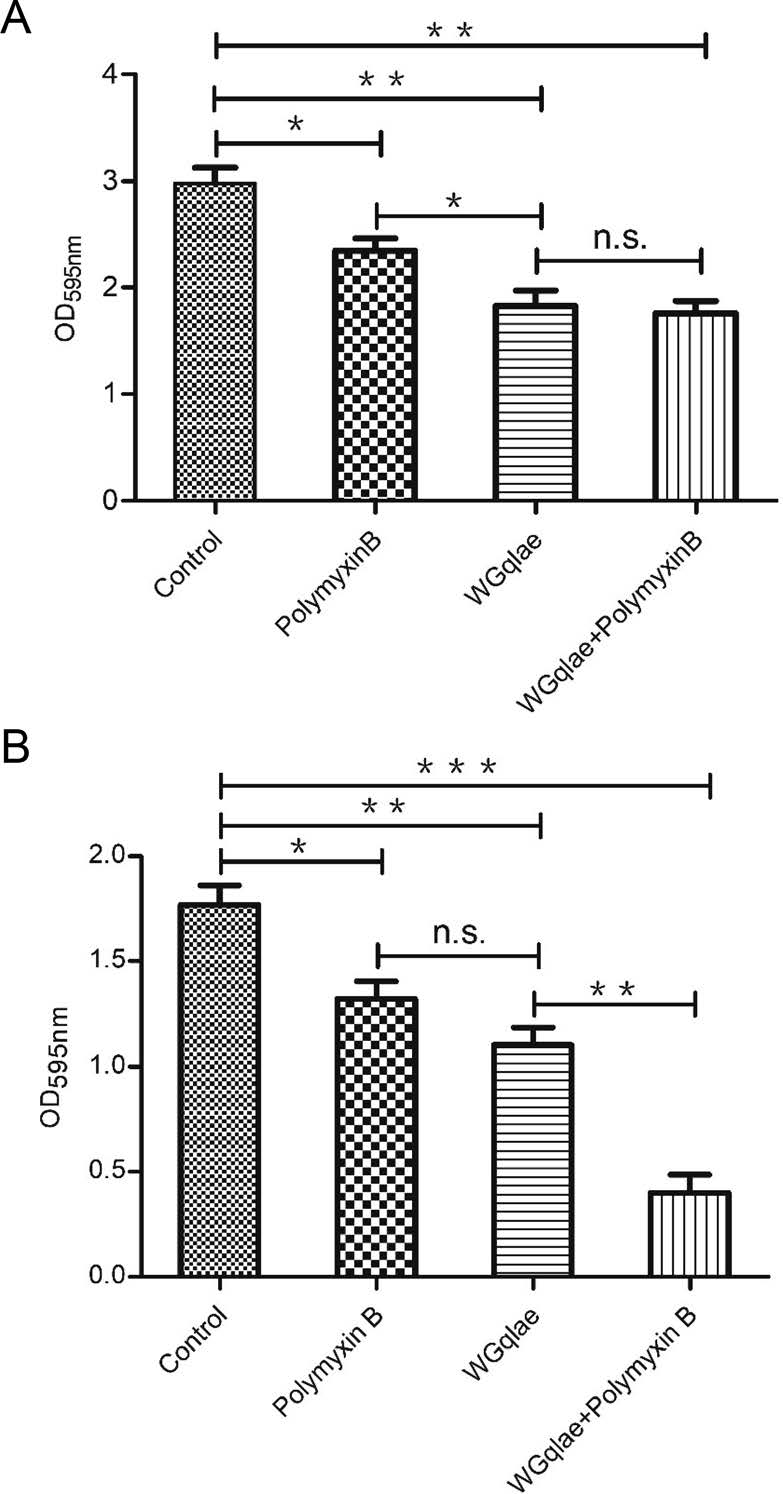
Recombinant phage WGqlae (P < 0.01), polymyxin B (P < 0.05) and the mixture of WGqlae + Polymyxin B (P < 0.01) inhibited the biofilm formation of host bacteria DE192 (Fig. 6A) and DE205B (Fig. 6B), significantly. When phage WGqlae was used alone, the biofilm of host bacteria DE192 and DE205B decreased by 38.59% and 38.20% respectively (P < 0.01), compared with the control group. As shown in Fig. 6A, phage WGqlae had better antibacterial effect than antibiotic Polymyxin B (P < 0.05). As shown in Fig. 6B, when the mixture of WGqlae + Polymyxin B was used, Polymyxin B significantly enhanced the ability of phage WGqlae to inhibit biofilm formation of strain DE205B (P < 0.01). Comparing with the control group, the biofilm formation of host bacteria DE192 and DE205B in WGqlae + Polymyxin B group was reduced by 40.94% (P < 0.01) and 77.53% (P < 0.001), respectively (Fig. 6).
-
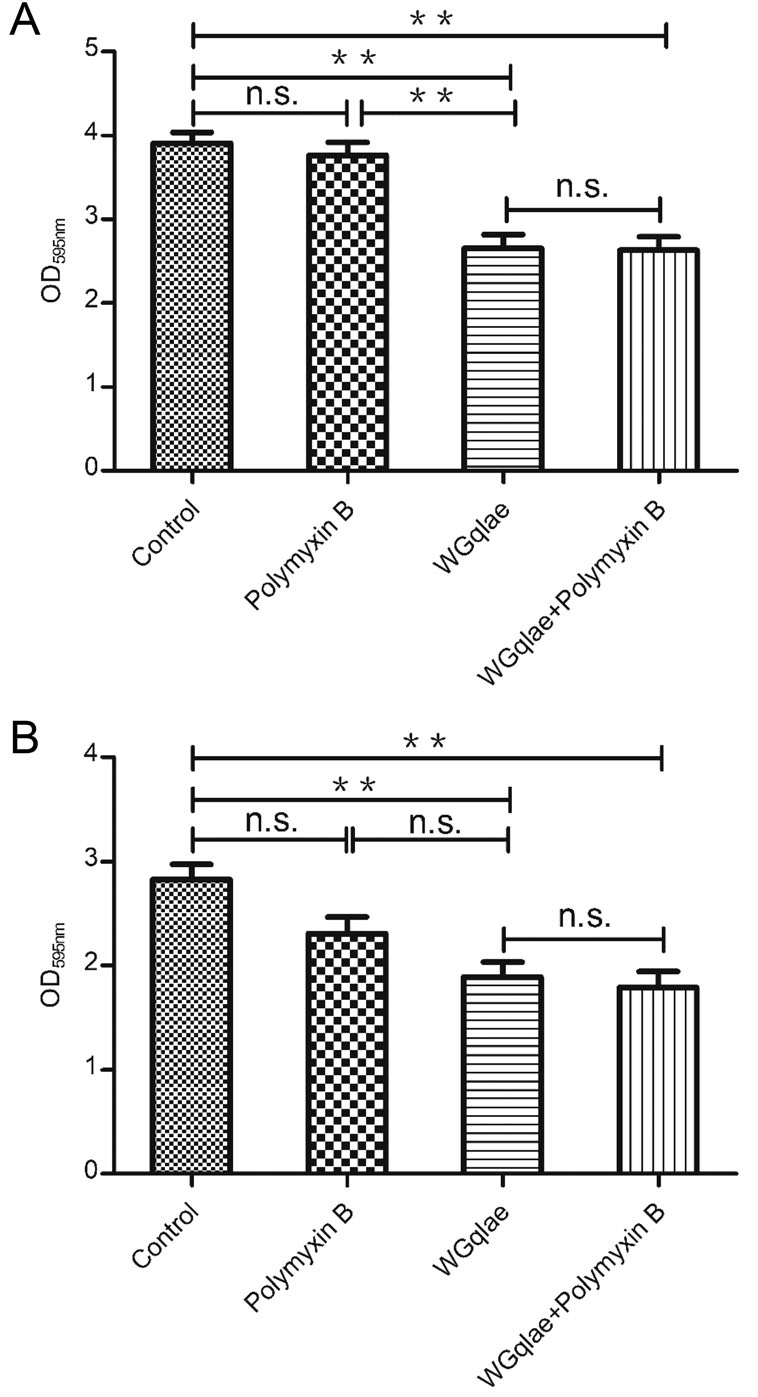
Comparing with the control group, the formed mature biofilm of E.coli DE192 and DE205B in phage WGqlae group decreased by 32.23% (Fig. 7A) and 33.22% (Fig. 7B), respectively (P < 0.01). No significant difference was observed between Polymyxin B group and the control group, suggesting Polymyxin B had no effect on established mature biofilm (both Fig. 7A, 7B). There was also no significant difference between WGqlae + Polymyxin B group and the only phage WGqlae group in removing the formed biofilm, which suggested that Polymyxin B can contribute to inhibiting biofilms formation but not to remove the formed mature biofilm of DE192 (Fig. 7A) and DE205B (Fig. 7B).
Recombination of Phage WG01 with Different Regions of Gene 37 of QL01
Host Range of WG01 Derivatives and Parental Phages
Comparison of Thermo and pH Stability of WG01 Derivatives WGqlae
Optimal Multiplicity of Infection
Burst Size and Latent Time Period
Genetic Stability of Recombinant Phage WGqlae
Efficacy of Phage WGqlae on Inhibiting E. coli in Planktonic State
Efficacy of Phage WGqlae, Polymyxin B and Their Combination to Inhibit Biofilms Formation
Efficacy of Phage WGqlae, Polymyxin B and Their Combination to Degrade Mature Biofilm
-
In this study, four chimeric phages (WGqlce, WGqlbe, WGqlae and WGqlde) with expanded host range were obtained by homologous recombination of gene 37. The four chimeras possessed the phage WG01 backbone with partial replacement of gene 37 from the phage QL01. To our surprise, derivatives WGqlae obtained nearly the entire host range from parental phage QL01 and an extra four host bacteria which was unsusceptible to the parental phages. The chimeric phages, WGqlae (with AA 237–925 of QL01 gp37), WGqlbe (with AA 317–925 of QL01 gp37), WGqlce (with AA 397–925 of QL01 gp37) and WGqlde (with AA 477–925 of QL01 gp37), had similar host ranges. Furthermore, our previous research showed that phage WG01 derivatives WQD1 obtained a large proportion of host range of QL01, even though only the AA 714-925 fragment of QL01 gp37 has been replaced (Chen et al. 2017). These results indicated that the main receptor binding region of QL01 was located in the C-terminal AA 714-925 of gp37, but not much to do with the N-terminal AA 1-713 region.
The distal tail fiber locus specifies the all-important adsorption tip of T4 phages and primarily determines the host range of the phages (Hashemolhosseini et al. 1994; Kellenberger et al. 1965; Tetart et al. 1998; Wais and Goldberg 1969). As early as 1996, Tetart et al. reported that T4 phage can recombine with gene 37-38 segment from another T-even phage, thus acquiring donor phage's host range (Tetart et al. 1996). Other studies reported that T4 phage can acquire the host range of donor phage T2- like, T6 phages, or even phage λ by recombination of the tail fiber gene (Tetart et al. 1998). However, in this study, phages WGqlfe and WGqlm were not succesfully isolated, which indicated that not all subsegments recombination can generate active engineered phage, probably because they changed the three-dimensional structure of gp37 or the defect of screening methods for chimeric phages. As it stands, the mechanism of the exact locus, subsegments module or key amino acids of domain (AA 714-925) in gp37 sequence determining receptor binding specificity remains unclear and will be an important project in future. Further study is required to address how the key domains or amino acid sites in gp37 sequence affect three dimensional structure and the function of gp37, as well as the host range.
Determining the environmental stability of phages is essential for biocontrol of pathogenic bacteria. Our study showed that the biological properties (thermo stability, pH stability and one-step growth curve) of the recombinant phage WGqlae was similar to the donor phage QL01. In addition, the gene 37 of chimeric phage WGqlae did not mutate after 20 generations, which suggested that the receptor binding specificity of the recombinant phage was genetically stable.
Biofilms are generally considered as an essential factor in the pathogenesis of various opportunistic bacterial and fungal infections. In addition, biofilms can colonize medical devices or wounds which were particularly hard to be removed as biofilms are inherently highly antibiotic resistant (Hughes and Webber 2017). Our results showed that the engineered phage WGqlae can inhibit E. coli DE192 and DE205B in planktonic state, prevent biofilm formation, and remove mature biofilm. This study further demonstrated that specific host-range engineered phages can be successfully used as antibacterial agents against E.coli in planktonic state and biofilm form. Some studies also indicated that phage-based antibacterial testing had significant effect against biofilm formation (Endersen et al. 2017; Kelly et al. 2012). Once bacteria have formed biofilm, they tended to be less susceptible to antibiotics (P > 0.05) (Fig. 7), causing a vital issue that antibiotics become ineffective. However, our study showed that phage WGqlae was highly efficient against formed mature biofilm (P < 0.01) (Fig. 7), which is superior to antibiotics in the treatment of biofilm producers. Furthermore, another advantage of phages was that they are protein-based entities totally devoid of metabolic machinery that can actively replicate only in the presence of viable host bacterial cells but have no affinity to eukaryotic cells (Rios et al. 2016). Consequently, the phage therapy has been proven to be safe and free from adverse side effects (Rios et al. 2016). Currently, several natural or engineered phage products have been being patented and as commercial antimicrobial agents (Kaistha and Umrao 2016).
In conclusion, our study demonstrated that recombined T4-like phage with expanded host range can inhibit planktonic pathogenic E. coli, prevent biofilm formation and remove mature biofilm. It is highlighted that we can engineeringly obtain phages containing the characteristics of wide host range and strong lytic activity, providing a promising tool for prevention and treatment of infections caused by pathogenic E. coli in the future. Further experiments should be performed to evaluate the safety and efficacy of engineered phages for the clinical usage.
-
This study was supported by Grants from the National Natural Science Foundation of China (U1803109), Key research and development plan of Jiangsu province (BE2019304), National Key R & D Program of China (2018YFC1602500), the Central University Basic Scientific Research Fund-Animal pathogenic bacteria (KYZ20 1846) and Jiangsu modern agriculture (waterfowl) industrial technology system disease prevention and control innovation team (JATS[2018]222).
-
WZ and HD conceived of the study. ML, DS, YL, YX, MC performed experiment, computational analysis. WZ and ML wrote the paper. ML and LC revised the final manuscript. All authors read and approved the final manuscript.
-
The authors declare that they have no conflict of interest.
-
This article does not contain any studies with human participants or animals performed by any of the authors.
















 DownLoad:
DownLoad: