HTML
-
Influenza A viruses are classified into subtypes based on two surface proteins: hemagglutinin (HA) and neuraminidase (NA). There are 18 HA subtypes and 11 NA subtypes (H1–H18 and N1–N11, respectively), expressing various subtype combinations and circulating in nature (Latorre-Margalef et al. 2014; Song et al2014; Wu et al. 2014). In April 2009, a novel swine-origin influenza A H1N1 (S-OIV) virus, classified as influenza A (H1N1) pdm09 or pdmH1N1, first emerged in Mexico and quickly replaced the traditional seasonal H1N1 to become the dominant H1N1 subtype (Novel Swine-Origin Influenza Novel Swine-Origin Influenza AVIT et al. 2009). To date, the influenza A (H1N1) pdm09 virus has been circulating in the world for more than one decade and has become a global threat to public health.
Vaccination reduces the risk of flu illness when most circulating flu viruses are well-matched to the vaccine. However, antigenic drift, which continually happens over time as the virus replicates, might decrease the vaccine's effectiveness. Most flu shots are designed to target HA surface proteins/antigens of influenza virus. Like other subtypes, pdm09 HA plays a pivotal role in binding host cell receptors and releasing the viral RNA into the cell (Beer et al. 2018). It contains a receptor-binding site (RBS) in a globular head domain which is composed of three domains, the 130-loop (residues 134–138), the 190-helix (residues 188–195), and the 220-loop (residues 221–228), and targets sialic acid residues on host cells (Skehel and Wiley 2000; Mair et al 2014; Tzarum et al 2015). Four main canonical epitopes, Ca (including Ca1 and Ca2), Cb, Sa, and Sb, on the globular HA head have been recognized by the host immune system. To prevent them from being blocked by antibodies elicited due to vaccination and natural infection, changes in HA protein associated with antigenic drift happen continually over time as the virus replicates (Caton et al 1982; Martinez et al 2009). Circulating influenza viruses evade neutralization in their human hosts by acquiring escape mutations. Nevertheless, it might result in the change of structure and biological functions of HA and makes mutant virus easier for reinfecting humans if escape mutations residues overlap the RBS region.
HA K166Q (H3 numbering without signal peptide), is believed to be the first antigenic drift site identified by an epitope mapping approach and became dominant in the 2013–2014 influenza season (Yasuhara et al 2017). Christopher et al. identified seven distinct amino acid substitutions in the HA segment under the selective pressure of a human monoclonal antibody (EM4C04), six of which significantly reduced binding activity to EM4C04 (K123N, D131E, K133T, G134S, K157N, and G158E) and one of which, S186P, increased the binding affinity of HA to the receptor (O'Donnell et al 2012). According to the GISAID Flu database, S186P did not become dominant until 2018.
An escape mutation N159K was identified as being responsible for the antigenic drift in the epitope Sa in the ferret (Guarnaccia et al 2013). Rudneva et al. reported five amino acid substitutions in the epitopes Sa (N129S, K156E, G158E, and N159D) and Sb (D190N) (Rudneva et al 2012). Chen et al. showed A144E and K145E substitutions in the epitope Ca2, G158E in the epitope Sa, G173E in the epitope Ca1, and S188N and Q192E mutations in the epitope Sb by the epitope mapping strategy (Chen et al 2014).
Here we report an influenza A(H1N1)pdm09-infected clinical isolate which contained four substitutions in the epitope Sb (S186P, T188I, D190A, and Q192E). We analyzed the evolution of this antigenic drift motif, characterized its pathogenicity, and evaluated its transmissibility based on the vaccine strains recommended by WHO in the northern and southern hemisphere in the 2020–2021 influenza season.
-
On December 14, 2019, a 46-year-old male patient was enrolled in the Guangzhou No.8 People's Hospital (Guangdong, China) with fever (38.8 ℃) and respiratory symptoms. Bronchoalveolar lavage fluid was analyzed in a broad spectrum of microorganisms by metagenomic next-generation sequencing (mNGS) and confirmed to be positive for influenza A (H1N1) pdm09 virus.
-
Two clinical isolates, A/Guangdong/GLW/2018 (GLW/18) and A/Guangdong/LCF/2019 (GLW/19), were used in this study. The former viral RNA was extracted from patients' swab and the latter one from bronchoalveolar lavage fluid, respectively. Both cDNAs were generated by SuperScriptTM Ⅲ Reverse Transcriptase (Invitrogen) (Eisfeld et al 2014). Full genome sequences of both strains were determined by the Sanger Sequencing Method and assembled by the Vector NTI suite. Genome sequences were deposited in the GISAID's EpiFluTM Database: A/Guangdong/GLW/2018, EPI_ISL_335963; A/Guangdong/LCF/2019, EPI_ISL_412027.
-
Two strains, GLW/18 and LCF/19 were selected as the backbone for constructing different reassortant viruses to characterize and compare the 186–192 motif's pathogenicity on the HA segment. According to the manufacturer's instructions, viral RNAs of both isolates were extracted using a QIAamp Viral RNA Mini Kit (Qiagen, Valencia, CA). Full-length viral cDNA segments were amplified using reverse transcription-PCR with segment-specific primers as described (Hoffmann et al 2001), and cloned into the pHW2000-derived vector using a ligation-independent cloning method (protocol is available upon request). GLW/18-S186P and GLW/18-S186P-T188I mutagenesis plasmids were constructed using the QuikChange Ⅱ site-directed mutagenesis kit (Agilent). Genotypes of reassortant viruses used in this study are listed in Table 1.
H3 numbering HA1 HA2 HI Epitope Sb region 186 188 190 192 120 129 260 154 A/California/4/2009* > 20,480 GLW/18 S T D Q A N N R 2560 LCF/19 P I A E T D D G 320 GLW/18-LCF-HA P I A E T D D G 160 GLW/18-LCF-HA-chimera P I A E A N N R 320 LCF/19-GLW-HA S T D Q A N N R 2560 LCF/19-GLW-HA-chimera S T D Q T D D G 2560 GLW/18-HA-S186P P T D Q A N N R 640 GLW/18-HA-S186P-T188I P I D Q A N N R 160 *A/California/4/2009 RG version is used as a standard strain. Table 1. Genotypes and HI titers of the GLW/18, LCF/19 recombinant viruses and reassorted mutants.
Plasmids were sequenced to confirm that no mutations were introduced during the cloning procedure. Recombinant viruses and their parental prototype viruses were generated by transfecting eight plasmids into mixed MDCK and HEK293T cells, as described previously (Hoffmann et al 2000), using a TransIT LT-1 kit (Mirus, Madison, WI). All influenza viruses used in this study were rescued using the reverse genetics (RG) approach. RG viruses were sequence-confirmed and propagated in MDCK cells at 37 ℃ for two days (P1) using the standard protocol (Song et al 2014).
-
Viral titration was used for the determination of their growth curves. A standard growth kinetics approach was applied in this study. Briefly, MDCK cells were inoculated with each RG virus at MOI = 0.001 (multiplicity of infection) and incubated at 37 ℃ supplied with 5% CO2 in the minimal essential medium (MEM, GIBCO) containing TPCK-treated trypsin (GIBCO). Virus culture media were harvested at 12, 24, 48, and 72 h post-infection (h.p.i.), and the viruses were titrated in MDCK cells by plaque assay. The viral growth property was measured in MDCK cells with three independent wells as previously described (Wang et al 2019).
-
Viruses were serially diluted tenfold and absorbed onto confluent MDCK cells in 6-well plates and then incubated at 37 ℃ for 1 h. The supernatant was removed, and the cells were washed with PBS and then overlaid with 1% MEM agarose containing 2 μg/mL L-(tosylamide-2-phenyl) ethyl chloromethyl ketone (TPCK)-treated trypsin. The plates were incubated at 37 ℃ for 48 h and then fixed with 4% formaldehyde in 1× PBS buffer for at least 2 h. Plaques were visualized by staining with 1% crystal violet in 20% alcohol solution.
-
HI tests with recombinant viruses were conducted as described (Wu et al 2008). Briefly, recovered patient sera collected from the GLW/18-infected patient were treated with a receptor-destroying enzyme (RDE, Denka Seiken Co., Tokyo, Japan). Control antibodies and sera were then twofold serially diluted in 96-well plates. Subsequently, 8 HA units of the virus were added to each well and incubated at room temperature for 1 h. Then, 50 μL of 0.5% Turkey erythrocytes (Lampire Biological Laboratories, PA) was added to the serum/virus mixture followed by incubation for 30 min. The HI titer is defined as the reciprocal of the highest dilution of sera that completely inhibits hemagglutination. The HI test was started at a 1:20 dilution for anti-GLW/18 serum.
-
HA segment sequences of pdm09 viruses were downloaded from the GISAID flu database. The phylogenetic tree was constructed using MEGA 7.0 by the Neighbor-Joining (NJ) method with 1000 bootstrap replicates.
-
GraphPad Prism ver. 8.02 for Windows (GraphPad Software, USA) was used to analyze the data. Two-way ANOVA was performed. Probability values of less than 0.01 (P < 0.05) were considered statistically significant. No samples were excluded from the analysis.
Case
Virus Isolation
Generation of Recombinant Viruses
Growth Kinetics
Plaque Assay
Hemagglutination Inhibition (HI) Assay
Bioinformatics
Statistical Analysis
-
The patient was a 46-year-old man with hypertension, diabetes, chronic heart failure, viral hepatitis (type B), and gallbladder polyps who lived in Guangdong Province. Radiograph imaging of the chest was recorded during the disease progression (Fig. 1).
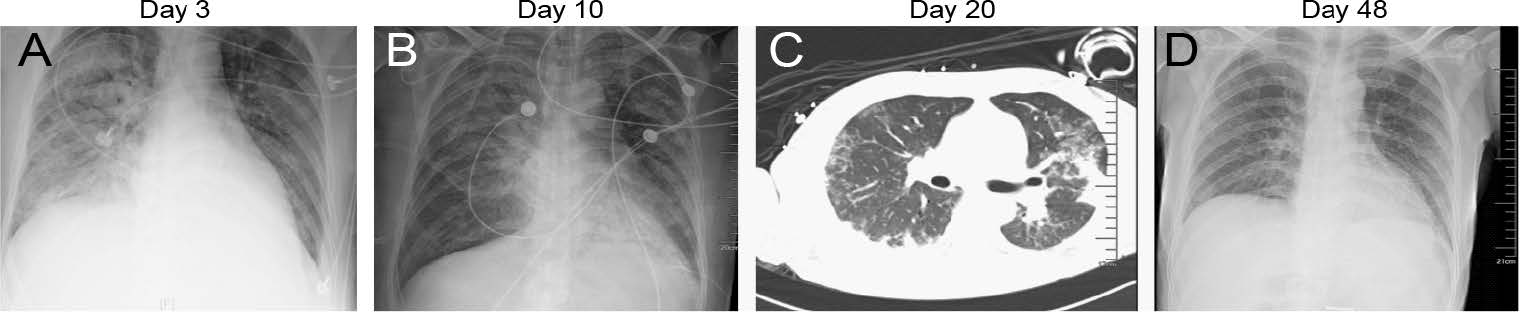
Figure 1. Representative radiographic findings of influenza A (H1N1) pdm09 virus. Clinical course of a 46-year-old man with hypertension, diabetes, chronic heart failure, viral hepatitis (type B), and hypoproteinemia infected with influenza A (H1N1) pdm09 virus, Guangdong, China, in 2019. A Radiograph imaging of the chest taken showed large and patchy shadows of both lungs with blurred edges, mainly the right lung, and partial blurring on the right diaphragm on the 3rd day after hospitalization; B Image taken on the 10th day, showing improvement of both lungs; C computed tomographic (CT) scan of the chest showed bilateral ground-glass opacities at the periphery of both lungs at day 20; D Image taken in the convalescent period of the infection on day 48, the X-ray film showed that the lung lesions were basically absorbed.
From December 14 to 23, 2019, the patient was admitted to the Guangzhou No.8 People's Hospital. On day 1, the patient started coughing, had a fever (38.8 ℃), dizziness, chills, and fatigue symptoms. On the same day, shortness of breath developed, and blood oxygen saturation was approximately 91%. The patient was diagnosed with severe pneumonia, acute respiratory distress syndrome (ARDS), and type II respiratory failure. On day 2, H1N1 was detected in his bronchoalveolar lavage fluid; therefore, the patient was treated with Tamiflu (oseltamivir phosphate, 150 mg/d). On day 3, a chest X-ray showed multiple large pieces of shadows in bilateral lungs and partial blurring on the right diaphragm. On day 10, December 24, the patient was transferred to the First Affiliated Hospital of Guangzhou Medical University. The patient's upper respiratory tract symptom onset accompanied by fever rapidly progressed to respiratory failure and multi-visceral failure. The pandemic H1N1 nucleic acid of influenza A virus was still detected as positive, and the patient was treated with meropenem, voriconazole, and oseltamivir. On day 32, the patient had no symptoms such as cough, shortness of breath, dizziness, chills, fatigue, and body temperature (37.5 ℃). Influenza A H1N1 virus test was negative. On day 39, the patient's condition was stable, and he was transferred from the intensive care department to the general ward. On day 63, the patient was recovered and was discharged (February 15, 2020). However, serum antibodies against anti-GLW/18 were not at a high concentration in vivo (HI = 40), suggesting that 2019 pdm09 H1N1 might undergo antigenic change.
-
We analyzed 41,987 full-length HA segment sequences of influenza A H1N1 pdm09 viruses downloaded from the GISAID database (cut-off date: August 24, 2020), 40708, which correspond to 97% of the sequences, were mainly classified into five genotypes based on the 186–192 motif. The earliest HA genotype containing a CA4-like motif (STSADQQ) was gradually replaced by a GLW-like motif (STTADQQ). The appearance of HA S188T and S186P in the epitope Sb also occurred in the coming year of the 2009 swine flu outbreak, although the substitutions became dominant in 2016 and 2019, respectively. A threonine-to-isoleucine substitution at HA 188 emerged in 2017, and its isolation rate reached a higher level in 2019 and the isolate with the PTIAAQE motif is circulating presently (Fig. 2A). The outbreak of the coronavirus SARS-CoV-2 led to reduced influenza surveillance and reporting activities in many countries. Additionally, travel restrictions and social-distancing measures in several countries resulted in decreased influenza activity. Overall, fewer viruses were available for characterization after August 2020.

Figure 2. Evolution of hemagglutinin protein of influenza A(H1N1)pdm09 virus. A Distribution of the hemagglutinin 186–192 motif; B The phylogenetic tree of hemagglutinin protein of influenza A(H1N1)pdm09 virus. The phylogenetic trees were generated using MEGA 7.0 by the Neighbor-Joining (NJ) method.
Interestingly, as a part of the epitope Sb, the 190 helix appeared in two amino acid substitutions D190A and Q192E in July 2019 and quickly became dominant in the pdm09 isolates, suggesting that such isolates (LCF-like motif PTIAAQE) adapted better to the population than the isolates containing the PTIADQQ motif and the PTTADQQ motif (Fig. 2B).
-
To examine the effect of viral replication, eight reverse genetics (RG) versions of pdm09 recombinant viruses (shown in Table 1) were infected in MDCK cells at an MOI of 0.001. We first compared the growth curves of two wild types, and our results showed that LCF/19 replicated faster than GLW/18 at 12, 24, and 48 h.p.i. (P < 0.01 at 24 h.p.i. and P < 0.05 at 48 h.p.i.) (Fig. 3A). To exclude factors from other segments, GLW/18-LCF-HA and LCF/19-GLW-HA recombinant viruses were generated.

Figure 3. Growth curves of the GLW/18, LCF/19 recombinant viruses and reassorted mutants. The growth kinetics of the wild-type GLW/18 RG virus and the indicated mutant viruses in MDCK cells were compared. Cell culture supernatants of MDCK cells infected at an MOI of 0.001 were collected at 12, 24, 48, and 72 h.p.i.. Virus titers are presented as the mean ± SD (n = 3). **P < 0.01, *P < 0.05 (one-way ANOVA).
The mean value obtained from triplicates showed that the LCF-HA segment grew faster than the latter with GLW-HA (no statistically significant difference) (Fig. 3A). Furthermore, to confirm that the 186–192 substitutive motif increased virus replication, RG versions of GLW/18-LCF and GLW/18-LCF HA chimeric segments were rescued. Both RG viruses grew faster than GLW/18 WT (P < 0.05 at 24 and 48 h.p.i.) (Fig. 3A). Overall, RG recombinant viruses containing the four substitutions on the epitope Sb replicated faster than those with GLW/18 HA genotypes in MDCK cells. Combined with the clinical data, mutagenesis that happened on the epitope Sb of the HA segment might increase the pdm09 viral pathogenicity in vivo.
To examine natural isolates containing the PTTADQQ motif and the PTIADQQ motif, two RG viruses, GLW/18-HA-S186P and GLW/18-HA-S186P-T188I, were generated to determine their growth kinetics curves. The GLW/18-derived recombinant viruses' growth curves were similar. Both demonstrated higher propagation velocities than GLW/18 WT at 24 and 48 h.p.i., respectively, indicating that the proline residue at HA 186 is a critical amino acid to decide the virus propagation (P < 0.05) (Fig. 3B).
-
To understand the molecular basis of the antigenic variation of pdm09 viruses since 2017, an HI test was performed (Table 1). Serum collected from GLW/18-infected patients was subjected to an HI test against GLW/18, whose HI titer was 2560. RG viruses containing the 186–192 motif STTADQQ of HA segments in the epitope Sb demonstrated at least eight times higher than those having the PTIAAQE. Of these mutations, HA S186P and T188I are critical in changing the binding activity to the sera. It seems that D190A and Q192E did not affect HI titers significantly.
-
The quadrivalent or trivalent influenza vaccine strains recommended by WHO in the 2020-2021 northern hemisphere influenza season were A/Guangdong-Maonan/SWL1536/2019 A/Hawaii/70/2019, both of which contain PTIAAQE mutation motif; these vaccines could prevent people from infection and severe outcomes caused by influenza viruses. (https://www.who.int/influenza/vaccines/virus/recommendations/en/). However, the vaccine strains recommended in the southern hemisphere between 2020 and 2021 do not include such mutation strains (A/Brisbane/02/2018 for use in 2020; A/Victoria/2570/2019 and A/Wisconsin/588/2019 for use in 2021) (Table 2). The GISAID database showed a few cases infected with pdm09 viruses carrying the HA segment's PTIAAQE mutation. Due to the difference in the selection of vaccine strains, people vaccinated in the southern hemisphere could still suffer a serious infection if infected with this antigenic drift-resulting pdm09 virus.
Year Vaccine type Strain name HA Epitope Sb region 186 188 190 192 Northern hemisphere 2020–2021 Egg-, cell- or recombinant-based A/Guangdong-Maonan/SWL1536/2019 P I A E A/Hawaii/70/2019 P I A E Southern hemisphere 2021 Cell- or recombinant- based A/Wisconsin/588/2019 P I D Q 2021 Egg-based A/Victoria/2570/2019 P I D Q 2020 A/Brisbane/02/2018 P T D Q Table 2. Genotypes of vaccine strains recommended by WHO in the southern and northern hemispheres in 2020–2021.
Taken together, a novel antigenic drift in the epitope Sb of the pdm09 virus was revealed by bioinformatic analysis and serological study. Currently, pdm09 viruses circulating in the northern hemisphere containing this mutated motif adapted well to human populations and are becoming dominant strains in the coming influenza season. Vaccine strains recommended for use in the northern hemisphere could protect against this escape mutant well. However, they may offer only partial protection against influenza infection if it quickly becomes a dominant strain in the southern hemisphere in winter 2020.
Clinical Specimen
Molecular Evolution of the 186–192 motif of the pdm09 HA Segment
HA S186P Enhances H1N1 Virus Propagation in Madin-Darby Canine Kidney (MDCK) Cells
The Escape Mutation in the Epitope Sb Decreased the Binding Activity Against GLW/18 Serum
The Vaccine Recommended for the Southern Hemisphere does not Contain the PTIAAQE Mutation
-
Influenza A virus is a serious global health threat that imposes an enormous economic and health burden, potentially causing periodic catastrophic pandemics (Kosik and Yewdell 2019). Among humans, the annual epidemic influenza illness is caused by two types, influenza A(H1N1) and influenza A(H3N2), which have been common influenza viral subtypes circulating among humans since the late 1970s (Cox and Subbarao 2000). In April 2009, a swine-origin A(H1N1) virus (pdm09), taking over the previously circulating seasonal A(H1N1) virus, caused the first influenza A virus pandemic of the twenty-first century (Novel Swine-Origin Influenza AVIT et al. 2009), and continues to cocirculate with the seasonal A(H3N2) virus. Influenza A(H3N2) appeared in the late 1960s and circulated among humans for over 50 years; therefore, two surface glycoproteins, hemagglutinin and neuraminidase, were evolved various mutations to evade the host immune system. Unlike influenza A(H3N2), the history of influenza A (H1N1) pdm09 is much shorter than that of influenza A(H3N2) circulating in the human population. Therefore, its surface proteins keep more conserved regions compared to its earliest isolate.
The two H1N1 pdm09 strains used in this study were GLW/18, isolated from a 7-year-old boy, and LCF/19, collected from a 46-year-old man with many diseases. Children, the elderly, and people with weakened immune systems are susceptible to being infected by the influenza A (H1N1) pdm09 virus.
To our knowledge, the first epitope mutation (HA K166Q) was identified in the clinical isolates in 2017 that showed superior growth to that of the wild-type virus (Yasuhara et al 2017). Both GLW/18 and LCF/19 clinical isolates also carry this escape mutation, suggesting that the HA K166Q is dominant in the epidemic strains. H1N1 pdm09 isolates containing HA S186P and T188I appeared in 2010 and 2017, respectively, but gradually became dominant in 2018. Moreover, double substitutions D190A and Q192E on the 190-helix domain occurred in Mid-2019, and its isolation is high, suggesting that those two residues, A190 and E192, at the 190-helix might have a much more stable structure than D190 and Q192. Therefore, the virus containing the escape mutant PTIAAQE (186–192) in the epitope Sb has become a stable strain in the population, indicating that it is easily circulating in the coming winter seasonal flu epidemics the northern hemisphere. A190V egg-adapted substitution occurred in WHO recommended vaccines, A/Guangdong-Maonan/SWL1536/2019 and A/Hawaii/70/2019, after serial passages in eggs. It is unclear whether this substitution will become dominant soon. The accumulated antigenic drift that finally causes antigenic shift might represent a pressing public health challenge.
Due to the coronavirus (COVID-19) pandemic, few flu data were uploaded to the GISAID Flu database after August 2020. We are concerned that the people living in the southern hemisphere could not obtain full protection if immunized with a vaccine lacking this mutation.
COVID-19 continues its spread globally and has now passed 50 million confirmed cases in 190 countries. Both viruses have similar transmission characteristics and common clinical manifestations but different treatments. Co-infection cases have been reported in different research groups (Konala et al 2020; Ozaras et al 2020; Singh et al 2020). It is unclear whether patients infected have a worse prognosis than those in whom SARS-CoV-2 is the only detected pathogen. Since vaccine additives can enhance immune flexibility, flu vaccine immunized-people might produce immune responses against COVID-19 if infected with SARS-CoV-2.
-
This work was supported by the National Natural Science Foundation of China (Grant No. 82074311), the General Project of Guangzhou Medical University (Grant No.SKLRD-MS- 201908), and the Yunnan Provincial Science and Technology Department (Grant No. 202005AF150043). We would like to thank Robert Webster of St Jude Children Research Hospital, USA, for providing the pHW2000 vector; Dr. Kai Huang and Dr. Pui Wang for constructive comments.
-
LX and YC performed the research and analyzed the results. BC, LB, YL, ZZ, WG, QC, and YL performed virus isolation, analyzed the clinical data. HC, XD and XW co-directed the study. KQ and WS directed the study, analyzed, interpreted results and wrote the manuscript.
-
The authors declare no conflict of interest.
-
The studies involving human participants were reviewed and approved by the Ethics Committee of The First Affiliated Hospital of Guangzhou Medical University (File No. 2014–50 approved on June 2, 2015). The patients/participants provided their written informed consent to participate in this study.














 DownLoad:
DownLoad: