-
Insect viruses are integral components of the ecosystem. They play important roles in regulating insect populations in agriculture and forestry in conjunction with other insect pathogens, such as bacteria and fungi as well as parasitoids, and predators. Among these insect viruses, the ascoviruses are a group first isolated in the late 70's and early 80's. They are double-stranded circular DNA viruses with a genome size of 115-180 kb (6, 11, 14, 37). Their delayed discovery is attributed to the fact that the infection symptoms by ascoviruses are not very pronounced in the field (7, 14, 21). In the laboratory, the symptoms of ascovirus-infected larvae include an extended larval stage of up to 2 months and difficlties in molting to cast the exuvium completely (10, 23). Since infected larvae do not develop into the pupal stage, ascoviruses usually cause a chronic but fatal disease to the host insects. Natural ascovirus infection in the field can range from 25% to 74% and often occurs in fields where chemical insecticides are restricted (8, 9, 21). The low occurrence of ascovirus in fields containing chemical pesticides is due to the low population of parasitoids affected by chemical insecticides, as ascoviruses are mainly transmitted by parasitoids during oviposition in the fields (20, 34). However, per os infection by ascovirus to lepidopteran larvae is possible and depends on the isolates (specific strains), which the infection rates can vary from 0 to 59% (21, 28). It seems the abundance of ascovirus is an environmental indicator; chemicalfree fields have a higher incidence of ascovirus infection, whereas fields with chemicals contain fewer or no ascoviruses. In chemical-free agricultural fields, ascoviruses have been isolated from economically important insect species such as Helicoverpa zea, Heliothis virescens, Spodoptera frugiperda, and S. exigua (8, 9, 21).
Since ascoviruses are naturally transmitted by parasitoids and replication of ascoviruses in the insect host often results in death of the parasitoids in the larvae, it is unlikely ascoviruses can be developed as biological pesticides. There are, however, several aspects of ascoviruses attracting the scientific community for research. These include its cytopathology, gene transcription, genome organization and evolution. In this brief review, we will discuss these biological features with emphasis on its evolution and phylogenetic relationship with other insect viruses.
HTML
-
Ascovirus classification takes into consideration the broad host ranges of ascoviruses and uses a combination of characters to differentiate species in the genus. These characters include virion morphology, presence of occlusion bodies, lack of DNA-DNA hybridization with other species at low stringency, restriction fragment length polymorphism (RFLP), host of isolation and experimental host range, association with specific hymenopteran parasitoids and DNA polymerase (DNAP) sequence, if available (15). Based on these criteria, four ascovirus species have been recognized by the International Committee on Taxonomy of Viruses (ICTV). These include Spodoptera frugiperda ascovirus 1a (SfAV-1a), Trichoplusia ni ascovirus 2a (TnAV-2a), Heliothis virescens ascovirus 3a (HvAV-3a) and Diadromus pulchellus ascovirus 4a (DpAV-4a). The number behind the viral species indicates the chronological order of discovery. The lower case letter following the number represents the genotype based on RFLP that is recognized for the accepted species. However, the species concept is subject to change once more new isolates are characterized and the described isolates are studied in more detail (15).
-
The first three ascovirus species (SfAV-1, TnAV-2 and HvAV-3) were discovered in lepidopteran insects in the US (8, 12, 20, 21). The fourth ascovirus (DpAV-4a) was reported from France (5, 6). There seems to be a world-wide distribution of ascoviruses. For example, ascoviruses have been reported from Indonesia and Australia (1, 10, 23). In each of the three lepidopteran ascovirus species, several variants were isolated from the same or different hosts in the same or different regions. For example, SfAV-1 has at least two variants, one from Florida and one from South Carolina; TnAV-2 has at least four variants, one from California and three from South Carolina; and HvAV-3 has at least fourteen variants, five from the U.S., three from Indonesia and six from Australia (9, 10, 14, 21, 23, 28).
Ascoviruses isolated from the same region are more closely related than those isolated from different regions. This might be caused by geographical separation during evolution as lateral genetic transfer could only occur within the local viruses or host insects. For instance, the restriction endonuclease (REN) profiles of all TnAV-2 variants from the U.S. are very similar and all U.S. HvAV-3 variants are also similar in REN profiles (9, 14). However, the HvAV-3 isolates from US and Indonesia look very different in terms of REN profiles even though DNA-DNA hybridization indicated they might be the same species (9). The three Indonesian ascoviruses from S. exigua showed similar REN profiles (23). The Australian HvAV-3e isolated from H. armigera in southeast Queensland is clustered with SeAV-5a from Indonesia based on phylogenetic analysis using the major capsid protein (MCP) and DNA polymerase (DNAP) sequences (9, 1). Both countries are in the Asian Pacific and geographically neighboring countries. However, these are the only a few cases suggesting this notion. Whether the genetic relatedness of these ascoviruses is truly correlated to geographic regions remains to be examined with more phylogenetic analyses.
-
HvAV-3 is the only one of the three lepidopteran ascovirus species isolated from regions outside of North America so far. It is also the species with the most variants reported and the most genetic diversity. It is the species with the highest variation in RFLP within the short period of time observed. For example, the four HvAV-3 (Hz84-271, Hv85-2, Hv87-3 and Hz87-5) isolates from 1984 to 1987 in Tifton, Georgia had minor PstⅠ REN profile difference by one or two DNA fragments (B. Federici, pers. comm.). However, TnAV-2b isolated in 1982 in South Carolina had only one REN fragment different to TnAV-2c isolated from the same location in 1997 (9). TnAV-2c and TnAV-2d collected in 2003 did not show PstⅠ REN profile changes to those isolated in 1997 in the same location in Blackville, South Carolina (9, 14, 23). This may therefore suggest that ascoviruses may have variable mutation rates or evolution rates, with HvAV-3 higher than TnAV-2 and SfAV-1. The diversification of these ascovirus species is probably caused by high mutation rates which drive the speciation and evolution of ascoviruses. The reasons why HvAV-3 has the most genetic diversity and is the only one isolated overseas of North America are unknown and have never been investigated. It seems all the HvAV-3 variants have the same genome size, about 180 kb, so that major gene acquisition through lateral gene transfer may not be a major contribution to the genome diversity. Plausible explanations for this high diversity might be that HvAV-3 has a higher error-rate DNAP or it does not carry an effective DNA damage repair system compared to those of other ascovirus species such as SfAV-1 and TnAV-2.
-
Ascoviruses cause disease in lepidopteran larvae in the families Noctuidae and Yponomeutidae (5, 20, 34). The unique feature of ascovirus-infected larvae is the presence of virion-containing vesicles in the hemolymph. The accumulation of large amounts of vesicles in the hemocoel makes makes hemolymph opaque white. These vesicles are formed by the partitioning of the host cells in which the ascoviruses replicate. Further investigation of the mechanism of cell cleavage by ascoviruses indicates that a viral-encoded caspase in SfAV-1a plays a role (3, 13). The shape of the virion is either allantoid or bacilliform depending on the species (13). However, both virion shapes are present in HvAV-3 (Fig. 1 A). Once the virions or virion-containing vesicles on the surface of the ovipositor of a wasp are injected into the hemocoel of a susceptible larva, the virions attach to susceptible cells such as fat body cells, and enter them probably by endocytosis (Fig. 1 B). In the cytoplasm, the viruses travel toward the nucleus, probably through an endocytic pathway, and induce a dramatic change to the nucleus. They first cause invagination of the nuclear envelope and later dismantle the nuclear envelope (10, 12, 13). Thus, the infected cell becomes nucleus-free (Fig. 1.C). Following the disruption of the nuclear envelope, the cell is partitioned to form vesicles and virion morphogenesis takes place concurrently (12, 13). However, vesicle formation is not typical in cell infection by a HvAV-3 (also called Spodoptera exigua ascovirus 5a, SeAV-5a) in vitro (Fig. 1.C). The mechanism of nuclear envelope dismantling is unknown, but this phenomenon seems to be analogous to the dissolution of the nuclear membrane in mitotic prometaphase and meiotic prophase Ⅰ. It is generally believed that phosphorylation of the lamins by the Cdc2 protein kinase results in the breakdown of the nuclear envelope (30). Several protein kinases have been identified in the ascovirus genome (2, 3, 37). The functions of these protein kinases played in the nuclear envelope broken down remain to be tested.
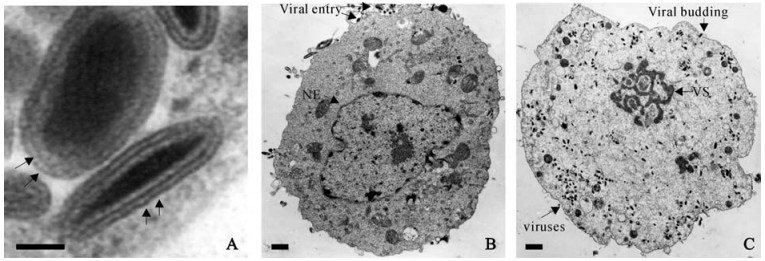
Figure 1. Transmission electron microscopy of SeAV-5a (HvAV-3) virion (A) and infecting SF-21 cells with nuclear envelope (NE) at 12 hr post infection or P.I. (B) and without a nucleus at 48 P.I. (C). Panel A shows the cross section of AeAV-5a virions in two different shapes, allantoid (upper) and bacilliform (lower). Arrows point to the double membrane envelopes. Panel B also shows viral entry and an intact nucleus. Panel C shows the viral replication as well with presence of virogenic stroma (VS) without viruses associated in a nucleus-free SF-21 cell. Bar markers represent 0.1 μm in A, 1 μm in B and C.
-
Among the four ascovirus species, the genomes of the three lepidopteran ascoviruses, SfAV-1, TnAV-2 and HvAV-3, have been sequenced recently. TnAV-2c was the first ascovirus that had the complete genome sequenced (37). Following that, SfAV-1a genome was sequenced and reported (3). The HvAV-3 genome has also been sequenced (2). GC contents of the ascovirus genome vary depending on species. TnAV-2c has the lowest GC content of 35.4% among the three sequenced genomes, SfAV-1a the highest of 49.2% and HvAV-3 has 45.8% (2, 3, 37). No introns were found in the three sequenced genomes. Genome parity analysis of the three ascovirus species suggested that HvAV-3 is more closely related to SfAV-1a than to TnAV-2c (2). This conclusion is supportive of the early DNA-DNA hybridization and a major capsid protein immunobloting studies which suggested the same relationship (9, 10). Homologous repeats were found in all the three genomes with HvAV-3 having 5 repeat regions, TnAV-2c having 2 and SfAV-1a having 4.5 (2, 3, 37). Genome sequences also revealed RNA polymerases in all the three ascovirus species. TnAV-2c contains four RNA polymerase subunits and a bacterial sigma factor (37). Whereas only three RNA polymerase subunits were found in the SfAV-1a and HvAV-3 without a sigma factor in their genomes (2, 3). It is still unknown if these RNA polymerases are responsible for all of the genes in the genomes or only some of the genes. Whether these viral enzymes are packed in the virions has not been investigated.
TnAV-2c transcripts are polyadenylated, as determined by 3' RACE studies; whereas the closed related iridoviruses have no polyA tails in their mRNA (unreported data, 27, 39). Unlike poxviruses, no polyA polymerase (PAP) has been found in the ascovirus genomes (2, 3, 37). This may suggest that ascovirus mRNAs are polyadenylated by the host PAP. Major polyadenylation of host transcripts takes place in the nucleus (38). Whether polyadenylation of ascovirus mRNA takes place before or after the breakdown of the nuclear envelope has not been scrutinized.
-
It is not certain how ascoviruses evolved, but within the family of Ascoviridae, genera or species may form a lineage which can be inferred from recognizable biological features among the four ascovirus species (Table 1). It is likely that the evolutionary history of the four ascoviruses proceeded from smaller to larger genomes, replication that is dependent on hymenopteran hosts to replication less reliant on hymenopteran hosts, a narrower to broader host range and different tissue tropism. According to this evolutionary trend, DpAV-4 represents the oldest ascoviral lineage, followed by SfAV-1 and then the more recently evolved HvAV-3/TnAV-2 viruses. The DpAV-4 ascovirus might have evolved from an iridovirus (32). The ascoviruses attack different tissues in the hemocoel of insects. SfAV-1a is about 157 kb, larger than the 116 kb-DpAV-4a genome (3, 6). TnAV-2 and HvAV-3 have genome sizes of about 180 kb, larger than that of SfAV-1a (2, 37). SfAV-1a infects only the Spodoptera species (21). TnAV-2 and HvAV-3 have a broad host range in lepidopteran insects. The tissue tropism depends on the ascovirus species. For instance, SfAV-1 and HvAV-3 attack primarily the fat body tissue cells whereas TnAV-2 replicates in other tissue cells in addition to fat body cells (15). DpAV-4 replicates only in the hymenopteran wasps. Viruses replicating in the hymenopteran hosts may represent more ancient viruses than those replicating in the lepidopteran hosts if the viruses co-evolved with their hosts (24). It is then also possible that DpAV-4 evolved earlier than iridoviruses which has species infecting lepidopteran hosts (26). Therefore it may be possible that SfAV-1 and lepidopteran iridoviruses shared a common ancestor with DpAV-4 and SfAV-1 gave rise to TnAV-2 and HvAV-3 (32). This is supported by a report that hymenopteran nucleopolyhedrovirus (NPV), with a smaller genome than the lepidopteran NPV, is considered to be a more ancient virus than the latter (25). There is also a suggestion that the larger lepidopteran NPV genome encodes some auxiliary genes that reduces viral dependence on host cellular machinery confers some selective or evolutionary advantage (25). Based on Darwin's theory of natural selection, ascoviruses increase their fitness by adapting to changes in host availability. By increasing their host range and expanding tissue tropism, ascovirus species increase their ability to parasitize a broader range of hosts and maximize progeny production in more tissues, thus overcoming potential host defenses. Expanding host range and widening tissue tropism might therefore be adaptive advantages during ascovirus speciation. It is still unknown if the extra 30 kb of TnAV-2 and HvAV-3 genomes contribute to the wider host spectrum and tissue tropism. More genetic studies on ascoviruses may reveal host range control genes which will advance our understanding of the general feature of host range control mechanisms.

Table 1. A summary of some biological characters of the four ascovirus species.
We applied the discrete character method, CLIQUE, in the PHYLIP 3.5 package (16) to construct the phylogenetic tree using the biological data from Table 1. The results showed that TnAV-2 and HvAV-3 shared a common ancestor and this common ancestor and SfAV-1 again shared another common ancestor which is a close relative of DpAV-4 (Fig. 2. A). Using single gene phylogenetic analysis with computer program PAUP 4.0 (33) on DNAP genes, a general evolutionary trend which is similar to the biological prediction can be inferred; that is, DpAV-4 evolved earlier than SfAV-1 which diverged into TnAV-2 and HvAV-3 (Fig. 2. B). However, TnAV-2c was predic ted evolving earlier than the rest of lepidopteran asco-viruses (9). Single gene analysis only presents gene evolution and it probably does not necessarily reflect viral genome evolution. Evolution of viruses needs to take into consideration the whole genome. When all the genomes of all four ascovirus species o have been sequenced, genome wide analyses might be performed to test if the DNAP evolution matches the genome evolution.

Figure 2. Phylogenetic analyses of ascoviruses. A: a phylogenetic tree inferred from biological characters of the four ascovirus species by PHYLIP without an outgroup assigned (16). B: a NJ phylogenetic tree inferred from amino acid sequences of DNA polymerase genes by PAUP. In B, Chilo iridescent virus (CIV) was used as an out group in the tree constructing. DpAV-4a, D. pulchellus ascovirus 4a, SfAV-1a, S. frugiperda ascovirus 1a, TnAV-2d, T. ni ascovirus 2d, HvAV-3c, SeAV-5a, S.exigua 5a also known as H. virescens ascovirus 3.
-
The four cytoplasmic DNA viruses, iridovirus, ascovirus, asfarvirus and poxvirus, show similarities and differences in their virion morphology. Morphological similarities were used to group viruses before molecular data were used for classification. Ascoviruses and poxviruses have double membrane envelopes with ovalshaped structures whereas iridoviruses and asfarvirus have icosahedral virions (Fig. 1A) (17, 40). There are similarities and differences in the biology of these viruses. Horizontal transmission of ascoviruses is mainly carried out by virus-contaminated ovipositors of parasitic wasps during oviposition on the caterpillars. Parasitic eggs of some wasps in the ascovirus-infected larvae normally could not develop into healthy adults (20, 34). There are also some insect iridoviruses such as an invertebrate iridescent virus (IIV) from S. frugiperda which are vectored by wasps. Replication of IIV is detrimental to the vectoring ichneumonid endoparasitoid Eiphosoma vitticolle (26). The poxvirus, which has the largest genome of animal viruses, seems to be more independent and infects insects through oral ingestion (18). Entomopoxviruses in fact are occluded in a proteinaceous structure called a spheroid which is believed to be an evolutionary adaptation for the virus to exist in the natural environment and propagate through ingestion.
The evolutionary origin of cytoplasmic DNA viruses has not been well studied. The insect specific baculovirus, especially the nucleopolyhedrovirus (NPV), replicate solely in the nucleus of infected cells (19). Viral DNA replication strategies among these cytoplasmic DNA viruses also show some evolutionary progress (Table 2). Iridovirus replicates first in the nucleus and continue in the cytoplasm (39). Ascovirus DNA replication has not been experimentally demonstrated yet but was reported in the nucleus based on the fact that virogenic stroma (VS) are often observed in the nuclei of infected cells (10, 12,). Virion morphogenesis, however, occurs in the cytoplasm after the nuclear envelope is ruptured. In the late stage of replication, ascoviruses partition the host cells and eventually the virions become encompassed within vesicles followed by cell disintegration (12). Poxviruses replicate DNA exclusively in the cytoplasm (17). It is likely that these viruses (irido-virus, ascovirus, asfarvirus and poxvirus) might have evolved from nuclear-replicating viruses such as NPV or granulovirus (GV) by becoming more independent of the nucleus for virus replication and more independent of wasp transmission, with the ascoviruses being the oldest evolutionary lineage, followed by the iridoviruses and the most recently evolved asfarviruses and entomopoxviruses. The relationship between the three viral families is supported by similar vesicle formation mechanisms in iridovirus and ascovirus and virion structure resemblance between ascovirus and poxvirus (36). Since asfarvirus and poxvirus share many genome similarities such as genome configuration (linear) and gene transcription strategies, it is possible that poxvirus had a reticulate evolution by recombination between ascovirus and asfarvirus which would make poxvirus a chimeric virus. Another possibility is that poxvirus evolved from asfarviruses, as poxviruses are genetically more similar to asfarvirus, and later acquired genes from ascoviruses that confered an ascovirus-like morphology. Phylogenetic relationships based on the DNAP gene from the poxviruses, ascovirus and asfarvirus showed entomopoxvirus and ascovirus are closely related and all the cytoplasmic viruses (poxvirus, ascovirus and asfarvirus) evolved from nuclear-replicating baculovirus (35).

Table 2. A summary of genetic and biological characters of baculovirus and the four cytoplasmic virus groups.
Phylogenetic analysis using major capsid protein (MCP), DNAP, thymidine kinase, ATPase Ⅲ and the presence of about 40 SfAV-1a protein homologues in CIV suggested that ascoviruses evolved from iridovirus (32). This was further supported by a genome wide comparison with other viruses, which showed 10% of SfAV-1a proteins are orthologs of Chilo iridescent virus (CIV) proteins and this strengthened the argument of evolution of ascovirus from iridovirus (3). However, there were no time scales placed in the phylogenetic analysis (32). Sharing a high percentage of homologous proteins between ascovirus and iridovirus only indicates they are closely related. Whether ascoviruses evolved from iridoviruses or iridoviruses evolved from ascoviruses is not certain.
A comparison of ascoviral and iridoviral proteins that are shared with baculoviral proteins provides some clues. The type species, CIV of iridoviruses shared 5 homologous proteins with baculovirus (22), whereas the ascoviruses, SfAV-1a and TnAV-2c shared 20 and 16 homologous proteins with baculoviruses, respectively (3, 37). The HvAV-3 shared 23 baculovirus repeat ORFs (bro) with baculoviruses (2). In addition, ascovirus and baculovirus virions share genome and virion structure similarity; both have circular genomes and the virions have an outer envelope in bacilliform (Table 2, Fig. 1A) (9). If viral evolution was a continuous process, maintaining certain morphological features, then more homologous proteins should be shared between closely related viruses. Therefore, it is highly likely that iridoviruses evolved from ascoviruses and the later evolved from baculoviruses because ascoviruses shared more homologous proteins with baculoviruses than did iridoviruses. In addition both ascoviruses and baculoviruses shared similar genome structure and morphological similarity. There are also suggestions that the African swine fever virus (ASFV, Asfarviridae) is the evolutionary link between the poxviruses and iridoviruses based on genome structure, gene transcription and DNA replication strategy (31). Stasiak et al. presented that the cluster of iridoviruses and ascoviruses was differentiated from Asfarviridae, which were previously a member of iridovirus (32). Phylogenetic analysis using DNAP that closely groups ascoviruses and poxviruses supports the hypothesis that ascoviruses and poxviruses are closely related even though evidence for direct evolution of ascoviruses to poxvirus is still weak. However, this study did not place iridovirus on the phylogenetic tree (35).
Using biological and genetic data set from Table 2, a phylogenetic tree with baculovirus serving as an outgroup was deduced among the four cytoplasmic DNA virus families by PHYLIP (Fig. 3A) (16). The tree predicts that the pox/asfar-virus shared a com mon ancestor with iridoviruses, and these three viruses again shared a common ancestor with ascoviruses. Molecular phylogenetic analyses using DNAP of the four cytoplasmic virus and baculovirus by neighbor joining method in PHYLIP, an unrooted tree was produced similar to what the biological dataset analysis predicted (Fig. 3B) (16). The molecular phylogenetic analyses using DNAP sequences supports the notion that all cytoplasmic DNA viruses originated from nuclear replicating baculovirus and that ascoviruses are more closely related to baculoviruses than iridoviruses. The prediction of using combined biological and molecular data has never been presented andanalyzed before. However, our predictions based on biological and molecular data match perfectly.
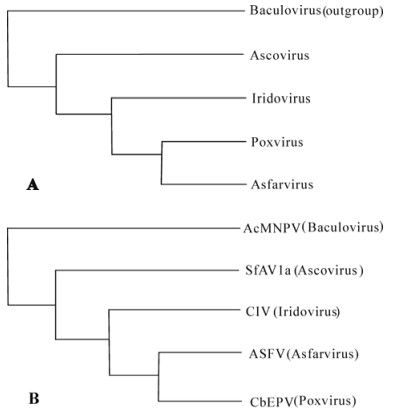
Figure 3. Phylogenetic analyses of ascoviruses. A: a phylogenetic tree inferred from biological characters of the four ascovirus species and nuclear replicating baculovirus by PHYLIP with baculovirus served an out group (16). B: a NJ phylogenetic tree inferred from amino acid sequences of DNA polymerase genes. AcMNPV, Autographa californica multinucleopolyhedrovirus; AfAV-1a S. frugiperda ascovirus 1a; CIV, Chilo iridescent virus; ASFV, African swine fever virus; CbEPV, Choristoneura biennis entomopoxvirus.
-
Understanding of the genetics and evolution of ascoviruses is still in its infancy. A clearer picture of the evolutionary relationship of ascoviruses with other insect viruses such as baculovirus, iridovirus and entomopoxvirus will be clearer with the complete genome sequences of all four ascovirus species, which is expected in 2007. It is expected that more research on gene transcription and cell infection strategies of acoviruses will pave the way to better understanding of how transcription strategies were maintained in closely related viral groups.













 DownLoad:
DownLoad: