HTML
-
Bacteriophages (phages) are viruses that infect and proliferate in bacteria. They play important roles inregulating the structure of microbial communities, metabolism and global biogeochemical cycles (Danovaro et al., 2008; Rohwer and Thurber, 2009; Suttle, 2005).Cold-active bacteriophages are capable of infection and reproduction at temperatures ≤ 4 ℃ (Rex et al., 2006). Anumber of cold-active bacteriophages have been isolated, including six phage isolates from Baltic Sea ice infecting Shewanella spp. and Flavobacterium spp. (Luhtanen et al., 2014; Sencilo et al., 2014). Forty phage-resistantisolates from 49 Flavobacterium psychrophilum strainswere isolated in Chile, Denmark and the USA (Castillo et al., 2012; Jian et al., 2013; Stenholm et al., 2008).Bacteriophage 9A infecting Colwellia psychrerythraea strain 34H was isolated from an Arctic nepheloidlayer (Colangelo-Lillis and Deming, 2013; Wells and Deming, 2006). Bacteriophages vB_EcoM-VR5, vB_EcoM-VR7 and vB_EcoM-VR20 showing an unusuallow-temperature plating profile were isolated fromLithuanian aquatic environments (Kaliniene et al., 2010).Three different phage-host systems were isolated fromArctic sea ice and the ocean north-west of Svalbard (Borriss et al., 2003).
Bacillus species are aerobic, Gram-positive, sporeformers and rod-shaped. They are ubiquitous in aquatic environments, soil and dried foods. A number of specific Bacillus species are used in molecular production and food fermentation. Although Bacillus is a very diversegenus with more than 100 species, only the Bacillus cereusgroup of species is associated with non-opportunisticinfection of mammals (Zwick et al., 2012). Cases of B.cereus associated with immunocompromised patients areincreasing (Ceuppens et al., 2013).
Although many Bacillus spp. phages have been isolated, B. cereus phages, especially cold-active B. cereusphages, have not received much attention, althoughsome temperate phages of B. cereus have been characterized (Kong et al., 2012; Lee et al., 2012). Bacteriophagesare divided into two categories, lytic and lysogenic. Inthe former case, bacterial cells are lyzed after replicationof the virion in the lytic cycle. Thus, lytic phages havebeen used in the control of B. cereus in mashed potatoes (Lee et al., 2011). In contrast, host cells infected bylysogenic phages do not immediately lyze.
The isolation and characterization of cold-active bacteriophagesfrom low temperature environments contributesto the understanding of cold-adaptation mechanisms and co-evolution of bacteriophages and their hosts.Glaciers are a unique ecosystem. Mingyong Glacieris located in the Meili Snow Mountains, part of theHengduan Mountains in Yunnan Province, China, and shows high biodiversity (Tang and Zheng, 1990). In thisstudy, a novel cold-active bacteriophage, VMY22, specific for B. cereus strain MYB41-22, was isolatedfrom Mingyong Glacier and the biological features ofthis bacteriophage were characterized.
-
Culture medium: Peptone-Yeast power-Glucose-Vitamins (PYGV) liquid medium (pH 6.0) was prepared asdescribed (Atlas, 2004). Fifteen grams of agar or 4 g ofagarose were added to liquid medium to prepare solidPYGV medium or semi-solid PYGV medium, respectively.
Bacterial strain: Water samples were obtained fromMingyong Glacier (E98°81′18.4″, N28°47′33.0″, 4–6℃, pH 5.4–6.0) located in Yunnan province, China.Samples were spread on PYGV solid plates and incubatedfor 2–3 d at 15 ℃. Bacteria l strains were isolated and purified by streaking on solid plates. Isolates wereidentified by 16S rRNA gene sequencing. PCR universalprimers used in this study were as follows: 27F (5′-AGAGTTTGATCCTGGCTCAG-3′), and 1492R (5′-GGTTACCTTGTTACGACTT-3′) (Lin et al., 2011; Liu et al., 2006). The PCR product of 1.5 kb was ligated intopMD -18T vector (Takara, Dalian, China) and the ligationproduct was transformed into Escherichia coli DH5αcompetent cells. Positive clones were sequenced byShanghai Sangon Biotech (Shanghai, China). The morphologyof B. cereus strain MYB41-22 was investigatedby scanning electron microscope.
A phylogenetic tree was constructed by neighbor-joiningusing Molecular Evolutionary Genetic Analysissoftware (MEGA 6.0) with 1000-fold bootstrap support.The 16S rRNA gene sequences of Bacillus strains and an outgroup of the type strain E. coli ATCC 11775T wereanalyzed.
-
B. cereus strain MYB41-22 was used as the host toisolate, propagate and characterize phage. Water samples (50 mL) collected from Mingyong Glacier were pre-incubatedwith host cells in order to enrich the phages. Afterincubation at 15 ℃ for 10 d, the culture was centrifuged (12000 × g, 15 min, 4 ℃, Beckman Avanti J-25, USA) and 0.22 μm filters (Millipore Corp., Bedford) were usedto filter the supernatant.
Phage was isolated by successive single-plaque isolationusing the double-layer agar method (Adams, 1959).Bacteriophage stock (100 μL) was mixed with MYB41-22 host cell culture (200 μL; OD600 = 0.3–0.6) and incubatedat 15 ℃ for 10 min, and then mixed with 4 mLof semi-solid PYGV medium and poured onto a PYGVsolid plate (Shen et al., 2012). Plaques were enumeratedafter the plates had been incubated at 15 ℃ overnight.
-
The method of single plaque isolation was used topurify phages (Xiang et al., 2005). All of the centrifugationwas carried out at 4 ℃. Cells were removed fromculture containing 1 × 109 PFU/mL phage particles bycentrifugation (12000 × g, 30 min, Beckman AvantiJ-25). The supernatant was supplemented with DNase I and RNase A (Sigma-Aldrich, 1 μg/mL) and incubatedfor 30 min at 37 ℃. 10% (w/v) PEG 8000 and 1 mol/LNaCl were added to the supernatant in order to precipitatephage particles. The phage particles were pelletedby centrifugation (11000 × g, 15 min) after incubation onice overnight, then resuspended in Suspension Medium (SM) buffer (50 mmol/L Tris-HCl, 100 mmol/L NaCl, 10mmol/L MgSO4, 0.01% gelatin), followed by extractionwith chloroform and centrifugation (11000 × g, 10 min).Solid CsCl (0.45 g/mL) was added to the supernatantwhich was then subjected to ultracentrifugation (280000 ×g, 24 h, SW41 rotor, Beckman LE-80K, USA). The band of phage was collected and dialyzed against SM bufferwithout gelatin.
-
Phage morphology was examined by transmissionelectron microscope (TEM). Approximately 1×109 PFU/mL phage was dropped into 200 mesh copper grids and stained with 2% (w/v) uranyl acetate. Morphology analysiswas conducted with a JEM-1230 TEM at 80 kV (JEOL, Tokyo, Japan).
-
MOI was defined as the ratio of phage particles topotential bacterial host cells (Lu et al., 2003). MYB41-22host cells (3 × 108 CFU/mL) were infected with VMY22 atdifferent ratios (0.001, 0.01, 0.1, 1, 10 and 100 PFU/CFU).The mixture was centrifuged (11000 × g, 10 min) to removethe pellet after incubation for 4 h at 15 ℃. A 0.22μm filter was used to filter the supernatant and the phagetiter was determined.
-
Phage adsorption experiments were carried out as follows:MYB41-22 host cells (3 × 109 CFU/mL) in PYGVwere infected with phage at an MOI of 0.1 and incubatedat 15 ℃. Aliquots of 100 μL were taken 0, 2, 4, 6, 8, 10, 15 and 20 min after infection, diluted in 0.9 mL coldPYGV, and centrifuged (12000 × g, 5 min, 15 ℃). Thesupernatants containing the unabsorbed phages were thentitrated using the double-layer agar method.
-
Fresh MYB41-22 host cells (1 mL, 3 × 109 CFU/mL) were mixed with phage at an MOI of 0.1 and adsorbedfor 30 min at 15 ℃ (Haq et al., 2012). The mixture wascentrifuged (13000 × g, 30 s, 15 ℃), and the pellets weresuspended in 5 mL of fresh medium before shaking at 15℃. Samples were taken at 5-min intervals and the phagetiter was measured by the double-layer agar method.Plates were incubated at 15 ℃ overnight to allow the detectionof plaques. Assays were repeated three times.
-
To examine the thermolability of the cold-activephage, VMY22 (2 × 109 PFU/mL) was incubated at differenttemperatures (40, 50, 60 and 70 ℃) for 1 h. Thesurvival rates of samples were determined at 10 min intervals.
-
In order to keep the pH stable, 10 μL phage stock (1× 109 PFU/mL) was mixed with 990 μL PYGV brothmedium, and incubated at room temperature for 1 h (Haq et al., 2012). The pH values of the mixture were adjustedfrom 3.0 to 11.0 with different buffers, including 50mmol/L citrate buffer (pH 3.0–5.0), phosphate buffer (pH6.0–8.0), Tris-HCl buffer (pH 9.0) and carbonate buffer (pH 10.0–11.0). The survival rates of samples were determined.
-
The sensitivity of phage to organic solvents and detergentswas studied. Phages were treated with 15% chloroformfor 10 min, 1 mg/mL proteinase K for 2 h at 37 ℃, Triton X-100 (0.3% w/v) for 10 min and SDS (0.3% w/v) for 30 min at room temperature, respectively. The survivingphage number was determined. The plaques wereexamined after incubation at 15 ℃ overnight.
-
Twenty millimolar EDTA, 10% SDS and 50 μg/mL proteinase K were added to a CsCl-purified phagesuspension and incubated at 56 ℃ for 3 h. Phenol/chloroform and ethanol were used to extract and precipitateDNA. The air-dried DNA pellet was dissolved in TEbuffer (10 mmol/L Tris-HCl, 5 mmol/L EDTA, pH 8.0) and digested with EcoR Ⅰ, BamH Ⅰ, Hind Ⅲ and Pst Ⅰ (TaKaRa, Dalian, China), respectively.
-
CsCl-purified phage particles mixed with loadingbuffer were boiled for 5 min, and then subjected to 12%SDS-PAGE. Coomassie brilliant blue R-250 (Bio-Rad, USA) was used to stain the protein bands.
Sampling and bacterial isolation
Bacteriophage isolation
Purification of phage particles
Electron microscopy
Determination of optimal multiplicity of infection (MOI)
Adsorption experiments
One-step growth curve of cold-active phage
Thermolability of cold-active phage
The pH sensitivity of cold-active phage
Sensitivity to organic solvent and detergents
Extraction of phage DNA and restriction endonuclease digestion
Protein analysis
-
The nucleotide sequence of 16S rRNA gene of strainMYB41-22 has been deposited in the NCBI with accessionnumber KC430113. Based on morphology, physiology and 16S rRNA gene phylogenetic tree analysis (Figure 1), MYB41-22 was identified as B. cereus MYB41-22.It was rod-shaped, spore-forming and belonged to theGram-positive class of bacteria (Figure 2). This psychrophilicbacterium could grow between 4–42 ℃, with optimumgrowth between 15 and 20 ℃.
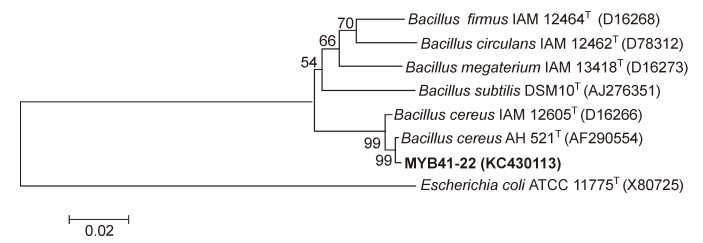
Figure 1. Phylogenetic analysis of strain MYB41-22 based on 16S rRNA gene sequences available from the NCBI GenBank database.
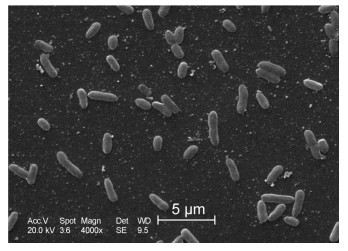

Figure 2. Scanning electron micrograph of strain MYB41-22. The scale bar marked automatically represents 5 μm.
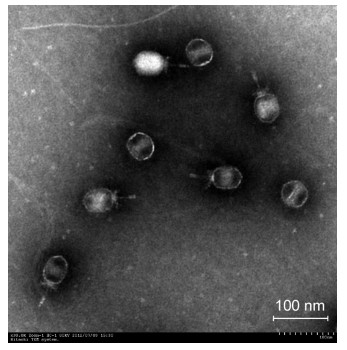
A phage infecting B. cereus MYB41-22 was isolated and named VMY22. Clear plaques of 2 mm were observedafter incubation of phage with bacteria at 4 ℃ for36 h on a double-layer plate. Cold-active bacteriophagescan infect and reproduce at temperatures ≤ 4 ℃, so it wastherefore recognized as a typical cold-active phage. TEMexamination revealed that VMY22 has an icosahedralhead wh ich is 31.9 nm in diameter and 59.2 nm in length, and a tail that is 43.2 nm in length (Figure 3), a typicalmorphology characteristic of the family Podoviridae.
-
B. cereus MYB41-22 strains could produce clearplaques after infection with VMY22 when incubated at 4–37 ℃, and showed maximal phage production at 15–20 ℃. Table 1 shows that the optimal MOI of VMY22was 0.1. Figure 4A shows that VMY22 could lyze almost100% of MYB41-22 cells in 15 min. A one-step growthcurve of VMY22 on MYB41-22 showed that the latentperiod was 70 min and the burst period was 70 min, with a burst size of about 78 bacteriophage particles per infectedcell (Figure 4B).
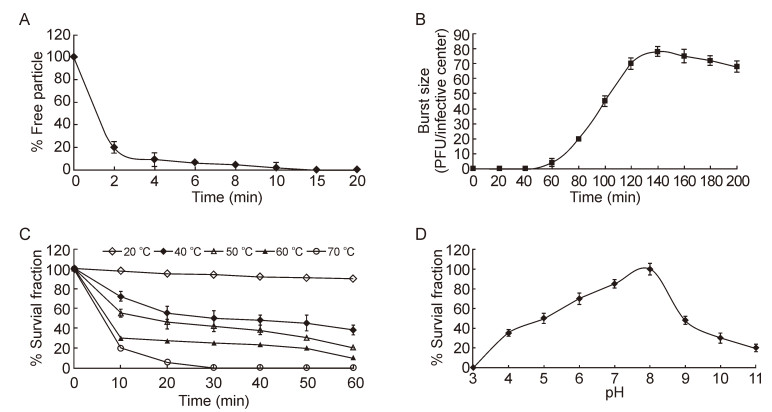
Figure 4. Characterization of VMY22. (A) The adsorption of phage VMY22 to host cells. (B) One-step growth curve of phage VMY22 on B. cereus MYB41-22. Values are the mean of three determinations. (C) Thermostability of VMY22 at 20 ℃ (open diamond), 40 ℃ (closed diamond), 50 ℃ (open triangle), 60 ℃ (closed triangle) and 70 ℃ (open circle). Values are the mean of three determinations. (D) pH sensitivity of VMY22. Values are the mean of three determinations.

Table 1. The optimal multiplicity of infection of B. cereus MYB41-22 by VMY22
-
Thermolability was the most salient physical featureof cold-active phage VMY22 (Figure 4C) (Wells and Deming, 2006). The VMY22 could tolerant more than60 min at 20 ℃ with minimal losses, and decreased rapidlywhen the temperature exceeded 60 ℃. VMY22 wasshown to be stable between pH 5.0 and 9.0, with maximumsurvival at pH 8.0 (Figure 4D).
-
Sensitivity to organic solvent and detergents were tested, and VMY22 was found insensitive to chloroform; itretained more than 80% infection activity after exposureto 15% chloroform. However, VMY22 infectivity wascompletely destroyed by treatment with protease K or by incubation with SDS or Triton X-100. The insensitivityto chloroform suggested that the capsid of VMY22 didnot contain lipids (Wells and Deming, 2006).
-
VMY22 DNA was extracted and digested with restrictionendonucleases EcoR Ⅰ, BamH Ⅰ, Hind Ⅲ and Pst Ⅰ (Figure 5A). The results showed that VMY22 wasdsDNA with an estimated size of 18–20 kb. Purifiedphage particles were analyzed by SDS-PAGE (Figure 5B). Three main protein bands were observed, with molecularmasses 22 kDa, 52 kDa and 57 kDa. Based on thegenome sequence of VMY22, these could correspond tolower collar protein, Podovirus_Gp16 superfamily protein and tail protein, respectively (data not shown). Themost abundant band was the 52 kDa protein, which maybe the major coat protein of cold-active phage VMY22.
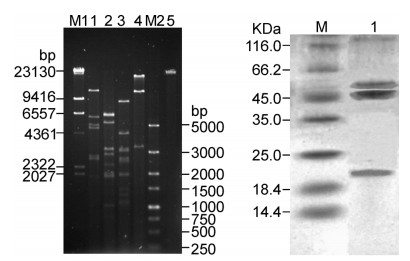
Figure 5. Analysis of VMY22 DNA and protein. (A) Restriction endonuclease digestion patterns of VMY22. M1: DNA marker; 1: VMY22 DNA digested with EcoR Ⅰ; 2: VMY22 DNA digested with BamH Ⅰ; 3: VMY22 DNA digested with Hind Ⅲ; 4: VMY22 DNA digested with Pst Ⅰ; M2: DNA marker; 5: Purified VMY22 DNA. (B) SDSPAGE of purified VMY22. M: protein marker; 1: Purified VMY22.
Isolation and morphology
Phage production
Thermolability and pH sensitivity
Sensitivity to chloroform and detergents
Analysis of phage DNA and protein
-
Cold-active phages have been isolated from low temperatureenvironments, such as seawater, food, sewage and sea ice (Anesio and Bellas, 2011; Lopez-Bueno et al., 2009; Rohwer and Thurber, 2009; Sawstrom et al., 2008; Sime-Ngando and Colomber, 2009). To the best ofour knowledge, this is the first report on the isolation and characterization of a B. cereus lytic cold-active phage from water samples from the Mingyong Glacier.
Of approximately 5, 600 known viruses, 136 Myoviridae, 203 Siphoviridae, 31 Podoviridae, 370 tailed bacteriophage and eight polyhedral, filamentous, and pleomorphic (PFP) infecting Bacillus have been observed (Ackermann, 2007). While numerous Bacillus phages have been isolated, very few cold-active B. cereus phages have been characterizedin detail. With the exception of phages AP50, AP50-04, AP50-11, AP50-23, AP50-26, AP50-27 and Bam35 belonging to the family Tectiviridae (Ackermann et al., 1994), all other Bacillus phages belong to thethree tailed phage families, Myoviridae, Siphoviridae and Podoviridae. Most B. cereus phages belong toMyoviridae, for example FWLBc1 (Lee et al., 2011), FWLBc2 (Lee et al., 2011), Bc431v3 (El-Arabi et al., 2013), BCP1-1 (Bandara et al., 2012), and BCP8-2 (Bandara et al., 2012), while B. cereus phage TP21-Lbelongs to the Siphoviridae (Klumpp et al., 2010). The B.cereus phage VMY22 isolated from Mingyong Glacier inthis study belongs to the Podoviridae.
The burst size of cold-active phage VMY22 was about78 PFU/cell at 15 ℃, lower than phage FWLBc1 (Lee et al., 2011) (322 PFU/cell at 37 ℃), FWLBc2 (300 PFU/cell, at 37 ℃) (Lee et al., 2011), and Bc431v3 (318 PFU/cell, at 30 ℃) (El-Arabi et al., 2013). Due to being acold-active phage, the burst size of VMY22 was lowerthan that of other B. cereus phages. In other cold-activephages, reduced burst sizes and protracted latent periodsat low temperatures than high temperatures have likewisebeen reported (Sillankorva et al., 2004; Wells and Deming, 2006), possibly due to bacterial phenotypicresponses to the transition to low temperature (Wells and Deming, 2006).
All of the Bacillus phages possess dsDNA. Genomeanalysis indicated that the genome size of phage VMY22was approximately 18–20 kb, much smaller than TP21-L (37.5 kb) (Klumpp et al., 2010), FWLBc1 (90 kb) (Lee et al., 2011), FWLBc2 (90 kb) (Lee et al., 2011), Bc431v3 (158 kb) (El-Arabi et al., 2013), BCP1-1 (150kb) (Bandara et al., 2012) and BCP8-2 (150 kb) (Bandara et al., 2012).
B. cereus is ubiquitous and is responsible for a minorityof food-borne illnesses. Taking advantage of its abilityto compete with Salmonella and Campylobacter, someharmless B. cereus in which animals and /or industriesthis technique is used to reduce Salmonella in the intestinesof livestock, thereby improving the growth of theanimals, as well as contributing to food safety.
Bacterial contamination can be found in low-temperatureenvironments such as fridges. Bacteriophages coulddecrease the number of bacterial infections and may be candidate antimicrobial agents against bacterial contamination.Cold-active bacteriophages possess the abilityto control pathogens in low-temperature aquaculture and phage therapy has received recent renewed interest (Castillo et al., 2012). As VMY22 is a cold-active bacteriophagewith a relatively low reproduction temperature, its characterization and the relationship bet weenMYB41-22 and B. cereus phage deserve further study.
-
This research was supported by the National NaturalScience Foundation of China (31160121) and the YunnanProvincial Education Fund project (2013Z138) and fundedby the Open Research Fund Program of the State KeyLaboratory of Virology of China (2013002).
-
All the authors declare that they have no conflicts ofinterest. This article does not contain any studies withhuman or animal subjects performed by any of the authors.
-
Ji XL, Tang B, and Wei YL designed the experiments.Ji XL, Zhang CJ, and Fang Y carried out the experiments.Ji XL, Zhang Q, Lin LB, and Wei YL analyzedthe data. Ji XL and Wei YL wrote the paper. All authorsread and approved the final manuscript.














 DownLoad:
DownLoad: