HTML
-
More than 100 years past the catastrophic 1918 Spanish influenza pandemic, influenza viruses remain a constant global health threat. Three types of influenza viruses infect humans: type A (influenza A virus, IAV), type B (IBV) and type C (ICV). Two IAV subtypes (H3N2 and H1N1pdm09) and two IBV lineages (Yamagata and Victoria) co-circulate annually, causing seasonal epidemics and up to 650, 000 respiratory deaths globally, while ICV is detected less frequently and usually causes mild infections (Krammer et al. 2018; WHO 2018). In addition, antigenically novel IAVs generated by reassortment of the segmented genome can occasionally infect humans with high rates of morbidity and mortality, as well as potential pandemic risk (Krammer et al. 2018; WHO 2018).
Vaccines provide cost-effective protection against influenza. Currently, seasonal influenza virus vaccines predominantly induce antibody responses against the hemagglutinin (HA), one of the two major surface glycoproteins of the virus (Krammer 2019). However, the majority of vaccine-induced antibodies are directed against the highly plastic head region of HA and are strain-specific (Caton et al. 1982; Heaton et al. 2013). Amino acid residues on the surface of this immunodominant head region vary substantially among different strains and change continuously (referred to as antigenic drift), leading to new circulating virus strains (Wang and Palese 2011). Therefore, current influenza vaccines have to be reformulated each year based on surveillance and prediction, which is cumbersome, and their effectiveness is highly variable depending on the accurate forecasts of the circulating strains in each year (de Jong et al. 2000; Gerdil 2003). Moreover, the protection conferred by seasonal influenza vaccines does not cover emerging pandemic influenza virus strains.
These limitations of current vaccines emphasize the need to develop novel "universal" or "broadly protective" influenza vaccines. An ideal universal vaccine should cover all influenza A and influenza B viruses independent of antigenic drift or HA/neuraminidase (NA) subtype. In contrast, a broadly protective vaccine would cover a large subset of influenza viruses, for example, all human seasonal influenza virus strains. These two types of vaccines are generally referred to as "universal vaccines" (Memoli et al. 2019).
Most universal vaccines in development aim at inducing broadly protective antibody responses, while some others focus on inducing T cell responses alternatively (Nachbagauer and Krammer 2017). In either case, the conserved immunogens of the virus need to be targeted, and the mechanism of protection mainly depends on the immunogens of choice (Nachbagauer and Krammer 2017; Asthagiri Arunkumar et al. 2019). This review will discuss the conserved antigenic regions/epitopes of influenza viruses that can facilitate immunogen design and contribute to the development of universal influenza vaccines.
-
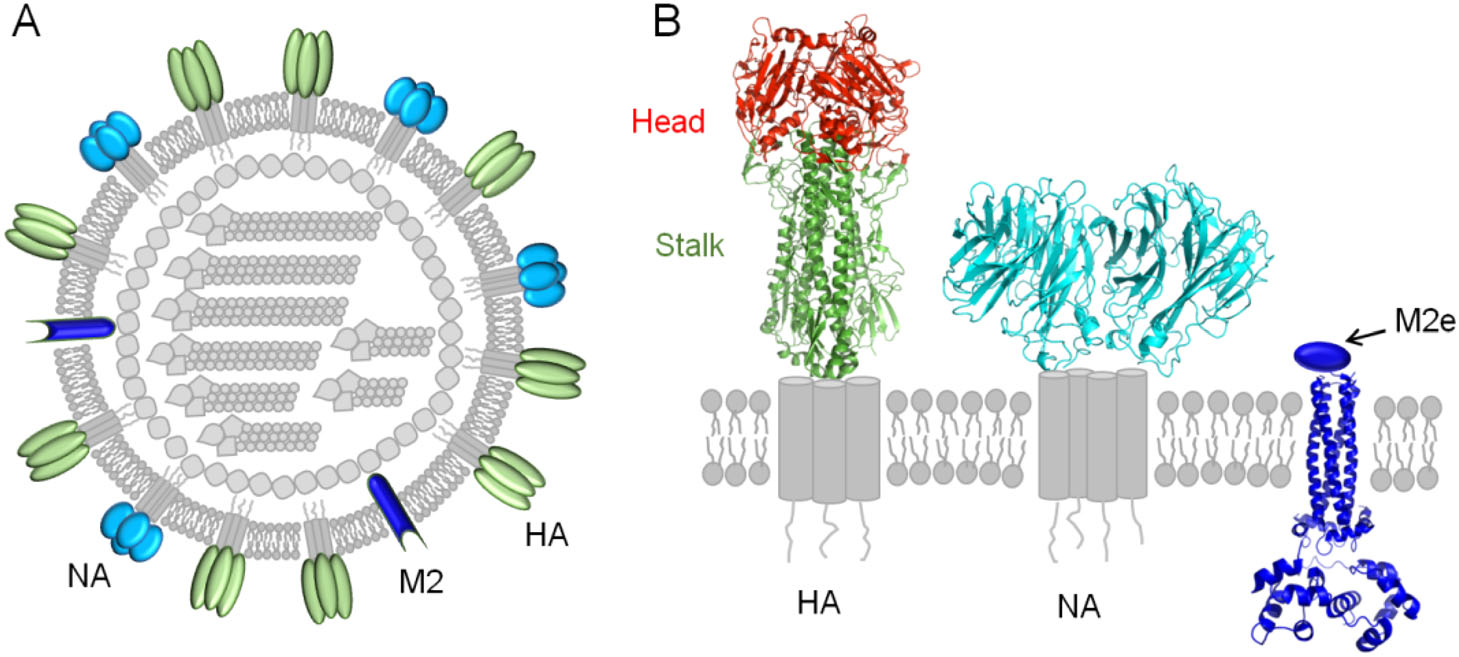
Induction of protective antibody responses against influenza viruses requires host recognition of the viral surface proteins, including HA, NA and matrix protein 2 (M2) (Fig. 1A). Thus the discovery of new broadly protective antibodies and related conserved epitopes can greatly accelerate design and implementation of universal vaccines (Deng and Wang 2018).
-
HA is assembled as homotrimers consisting of two domains, the membrane-distal globular head domain and the membrane-proximal helix-rich stalk domain (Fig. 1B). Based on the antigenicity, HA can be divided into 18 subtypes, which fall into 2 groups, including group 1 (H1, H2, H5, H6, H8, H9, H11, H12, H13, H16, H17, H18) and group 2 (H3, H4, H7, H10, H14, H15) (Du et al. 2019). As mentioned above, HA head domain is the major target of the antibody response following vaccination with seasonal influenza vaccines, however its high plasticity makes it easy for the virus to escape immune pressure (Heaton et al. 2013). Despite of this, monoclonal antibodies (mAbs) that cross-react between the head domains of different HA subtypes have been reported (Ekiert et al. 2012; Krause et al. 2012; Lee et al. 2012; Boonsathorn et al. 2014; Lee et al. 2014). These cross-reactive head antibodies seem rare, but they are interesting since they suggest the existence of cross-reactive epitopes in the head domains.
In contrast, HA stalk is the least variable region of HA, and great efforts have been focused on the development of universal influenza vaccines that depend on protective epitopes in this domain (Krammer and Palese 2015; Wu and Wilson 2018). Monoclonal antibodies against this domain can broadly neutralize different subtypes of group 1 (Okuno et al. 1993; Ekiert et al. 2009; Sui et al. 2009) or group 2 viruses (Ekiert et al. 2011) or even both (Corti et al. 2011). Interestingly, although some stalk-antibodies show lower or even none neutralizing potency in vitro, they can render robust protection against challenges with divergent influenza viruses in vivo (Dreyfus et al. 2012; DiLillo et al. 2014; He et al. 2015), most likely by Fc-mediated mechanisms like antibody dependent cell-mediated cytotoxicity (ADCC) and complement-dependent cytotoxicity (CDC) (Terajima et al. 2011; Jegaskanda et al. 2013a, b; Miller et al. 2013; DiLillo et al. 2014; Florek et al. 2014; Jegaskanda et al. 2014).
Glycosylation and conformational mobility (shape-shifting) of surface glycoproteins can help viruses to mask sensitive epitopes and evade the immune system. Interestingly, two latest studies revealed that glycan repositioning within either head or stem domains of HA can facilitate the elicitation of protective cross-reactive antibody responses. Huang et al. reported that removing glycosylation at residue 144 and deletion of lysine at position 147 can synergize to elicit broadly-reactive H1N1 influenza antibodies (Huang et al. 2019). A study by Boyoglu-Barnum et al. showed that introduction of the group 2 glycan at Asn38HA1 to a group 1 stem-nanoparticle can broaden antibody responses to cross-react with group 2 HAs (Boyoglu-Barnum et al. 2020). Although the precise sites that are vulnerable by glycan repositioning are not identified yet, they may provide novel target epitopes to achieve universal protection.
More recently, a novel conserved epitope at the HA head domain trimer interface was reported (Bajic et al. 2019; Bangaru et al. 2019; Watanabe et al. 2019). Primarily, the epitope is occluded at the contact surface between HA head domains, while the reversible HA "breathing" conformational dynamics can cause exposure of the interface. Antibodies targeting this cryptic epitope were shown to be protective against broad-spectrum subtypes of IAVs, suggesting the HA head interface is a new potential immunogen component for influenza universal vaccine design (Bajic et al. 2019; Bangaru et al. 2019; Watanabe et al. 2019; Wu and Gao 2019).
-
NA is the second most abundant glycoprotein on the surface of influenza A and B viruses, and plays an essential role during virus replication by facilitating release of progeny virions (Fig. 1B) (Du et al. 2019). It is well known that NA antibodies can protect against infection of influenza viruses (Schulman et al. 1968; Murphy et al. 1972; Monto and Kendal 1973; Doyle et al. 2013; Wan et al. 2013; Wilson et al. 2016; Wohlbold et al. 2017), by preventing virus budding and egress from infected cells, as well as ADCC and CDC (Wan et al. 2013; Krammer and Palese 2015; Wohlbold et al. 2017). NA consists of 11 subtypes (N1 to N11), and a series of "broadly" cross-reactive mAbs of NA have been isolated (Wan et al. 2013; Wilson et al. 2016; Wohlbold et al. 2017; Chen et al. 2018). Of note, most of these cross-reactive NA mAbs are limited to recognize heterologous virus strains of the same NA subtype, suggesting existence of conserved subtype-specific NA epitopes (Wan et al. 2013; Wilson et al. 2016; Wohlbold et al. 2017; Chen et al. 2018).
Interestingly, Doyle et al. reported a unique universally conserved sequence (amino acids 222-230, located in the vicinity of enzymatic active site) amongst all IAV NA subtypes, and a mAb targeting this specific epitope was demonstrated to inhibit all NA subtypes and protect mice from lethal doses of mouse-adapted H1N1 and H3N2 (Doyle et al. 2013). More recently, Stadlbauer et al. identified three broadly-protective mAbs targeting the NA active site that is highly conserved among all NA subtypes (Stadlbauer et al. 2019). These conserved NA epitopes, either subtype specific or universal, may be attractive immunological target for universal influenza vaccine design.
-
M2 of IAV (AM2) is a type III integral membrane protein, consisting of an N-terminal extracellular region (M2e), transmembrane region, and C-terminal cytoplasmic tail region (Fig. 1B) (Tobler et al. 1999). The M2e is a 24-residue peptide and is highly conserved among different IAV subtypes (Deng et al. 2015). Although the natural M2e has low immunogenicity and abundance, a M2e-specific and protective monoclonal antibody 14C2 has been isolated early (Zebedee and Lamb 1989; Treanor et al. 1990). Therefore this region has since been extensively evaluated as a promising immunogen for universal influenza vaccines (Farahmand et al. 2019; Yao et al. 2019).
BM2 of IBV is a functional homolog of AM2 but shares little sequence identity (Wanitchang et al. 2016; Mandala et al. 2019), and most mAbs targeting M2e are IAV-wide. Despite a mAb AS2 that recognizes the conservative N-terminus of M2 (amino acids 2-10) has been identified to possess neutralizing activity against both IAV and IBV in vitro (Liu et al. 2003), no evidence showed that immunity with current universal influenza vaccine candidates based on M2e of IAV can suppress IBV replication (Mezhenskaya et al. 2019).
Hemagglutinin
Neuraminidase
Matrix Protein 2
-
CD8+ T cells can detect and kill virus-infected cells by recognizing viral protein-derived peptides (epitopes) presented by major histocompatibility complex class I (MHC-I) on the cell surface (Fig. 2A). While CD8+ T cell-based immunity does not prevent infection, it can facilitate viral clearance and reduce the severity of disease (McMichael et al. 1983; Sridhar et al. 2013).
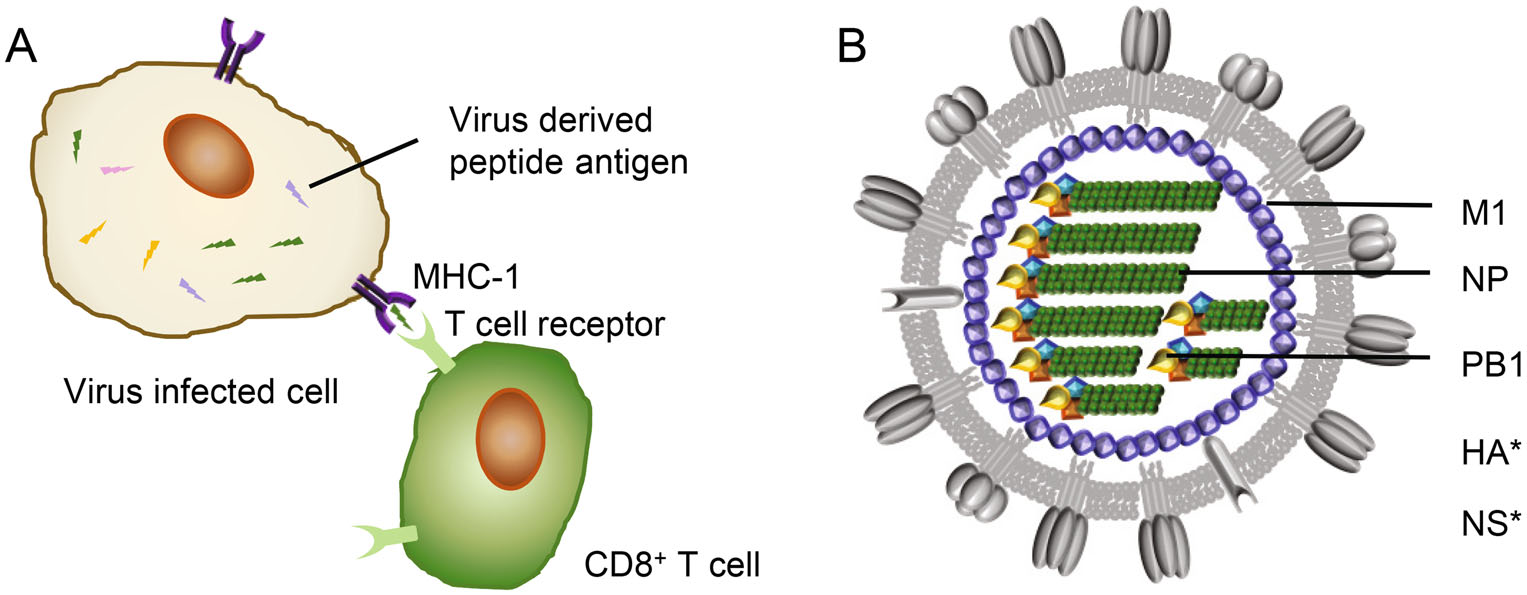
Figure 2. T-cell response and influenza viral internal proteins. A CD8+ T cell response. Virus-derived peptide antigens presented by major histocompatibility complex-I (MHC-1) at surface of virus infected cells can be recognized by CD8+ T cells through specific T cell receptors (TCRs), resulting in lysis of target cells. B A model of influenza viral particle with internal proteins. M1, Matrix protein 1; NP, nucleoprotein; PB1, polymerase basic-1. *, CD8+ T cell targets from IBV HA and NS (nonstructural protein 1) proteins also have been reported.
The current inactivated influenza vaccine formulation does not boost T cell response (Koutsakos et al. 2018). Nevertheless cytotoxic CD8+ T cells specific to the conserved viral epitopes have been evidenced to provide cross-protection across IAV, IBV and ICV (McMichael et al. 1983; Greenbaum et al. 2009; Gras et al. 2010; Sridhar et al. 2013; Quinones-Parra et al. 2014; Hayward et al. 2015; Wang et al. 2015, 2018; Koutsakos et al. 2019). Moreover, pre-existing T cell responses have been demonstrated to correlate with protection from influenza disease in humans by epidemiological studies (Epstein 2006; Hayward et al. 2015). The antigenic origin of these broadly cross-reactive epitopes is therefore of great interest in the development of CD8+ T cell-based universal influenza vaccines. It is believed that CD8+ T cell cross-reactivity is provided by the peptides derived from internal influenza proteins, mainly but not limited to nucleoprotein (NP) and matrix protein 1 (M1) (Fig. 2B) (Koutsakos et al. 2019).
In contrast to HA and NA, the internal influenza virus proteins are relatively well conserved and display limited antigenic diversity (Bui et al. 2007). A number of vaccine candidates have been developed to induce T cell responses, by intracellularly expressing NP and M1 using replication deficient viral vectors like Chimpanzee Adenovirus Oxford 1 (ChAdOx1) or Modified Vaccinia Ankara (MVA) (Berthoud et al. 2011; Lillie et al. 2012; Lambe et al. 2013; Antrobus et al. 2014a, b).
Interestingly, a recent study identified a universal PB1 epitope (PB1413), which is derived from a core motif (residues 406-422 of IAV-PB1 protein) present in the viral RNA-dependent polymerases (Koutsakos et al. 2019). PB1 is the most conserved protein across IAV and IBV, with about 60% amino acid identity. Vaccinating against the T cell epitope PB1413 might provide universal protection in humans. Besides, CD8+ T cell targets from IBV HA and NS1 proteins also have been identified and shown IBV-specific protection in mice, suggesting them as ideal targets for IBV-wide influenza vaccines (Koutsakos et al. 2019).
CD4+ T cell responses to influenza virus and their role for host protection against influenza infection have received increasing attention (Wilkinson et al. 2012). It has been well demonstrated that CD4+ memory T cells can help in directing a faster antibody response via cytokine secretion in response to mutated or immunologically novel viral antigens, leading to limited virus shedding and disease severity, possibly due to the direct lysis of infected epithelial cells (Poon et al. 2009; Wilkinson et al. 2012; Valkenburg et al. 2018). A vaccine that can induce CD8+ or CD4+ T responses against highly conserved regions of the influenza proteins may overcome the limitations of current season influenza vaccines (Sheikh et al. 2016). More importantly, universal T cell-inducing vaccines in combination with universal antibodies can confer increased protection against influenza (Asthagiri Arunkumar et al. 2019).
-
In theory, a universal influenza vaccine would work well if it contains a conserved epitope that remains the same from year to year and does not vary between strains. However, no such viral epitope capable of stimulating an immune response that stops most influenza viruses afflicting humans has been reported yet (Cohen 2018). Certainly there are main obstacles on the way to developing universal influenza vaccines, but different strategies can be used to overcome them.
-
Specific induction of broadly neutralizing antibodies can be achieved by targeting B cell recognition to conserved epitopes. However, the low immunogenicity of the conserved HA stalk and M2e makes antibody responses targeting these antigens constitute only a small fraction of the total anti-virus antibodies in nature infections (Sui et al. 2011). Various immunofocusing approaches have been therefore developed to overcome this problem.
The M2e is small in size and has low abundance in its native state, but its immunogenicity can be improved using tandem repeats and fusing them to highly immunogenic carriers (Neirynck et al. 1999; De Filette et al. 2008; Eliasson et al. 2008; Turley et al. 2011; Kim et al. 2013). Strategies directing the antibody response to HA stalk were also evaluated, such as shielding of the HA head epitopes by hyperglycosylation (Fig. 3A) (Lin et al. 2012; Eggink et al. 2014), and constructing headless mini-HA as protective immunogen by removing the head domain completely (Fig. 3B) (Steel et al. 2010). Notably, the latter strategy requires maintaining the conformation of the stalk domain in absence of the head domain, which is challenging. Nevertheless promise progress has been made to achieve the stable native structure of trimeric HA stalk, either in soluble form or presented on nanoparticles (Lu et al. 2014; Mallajosyula et al. 2014; Impagliazzo et al. 2015; Yassine et al. 2015).
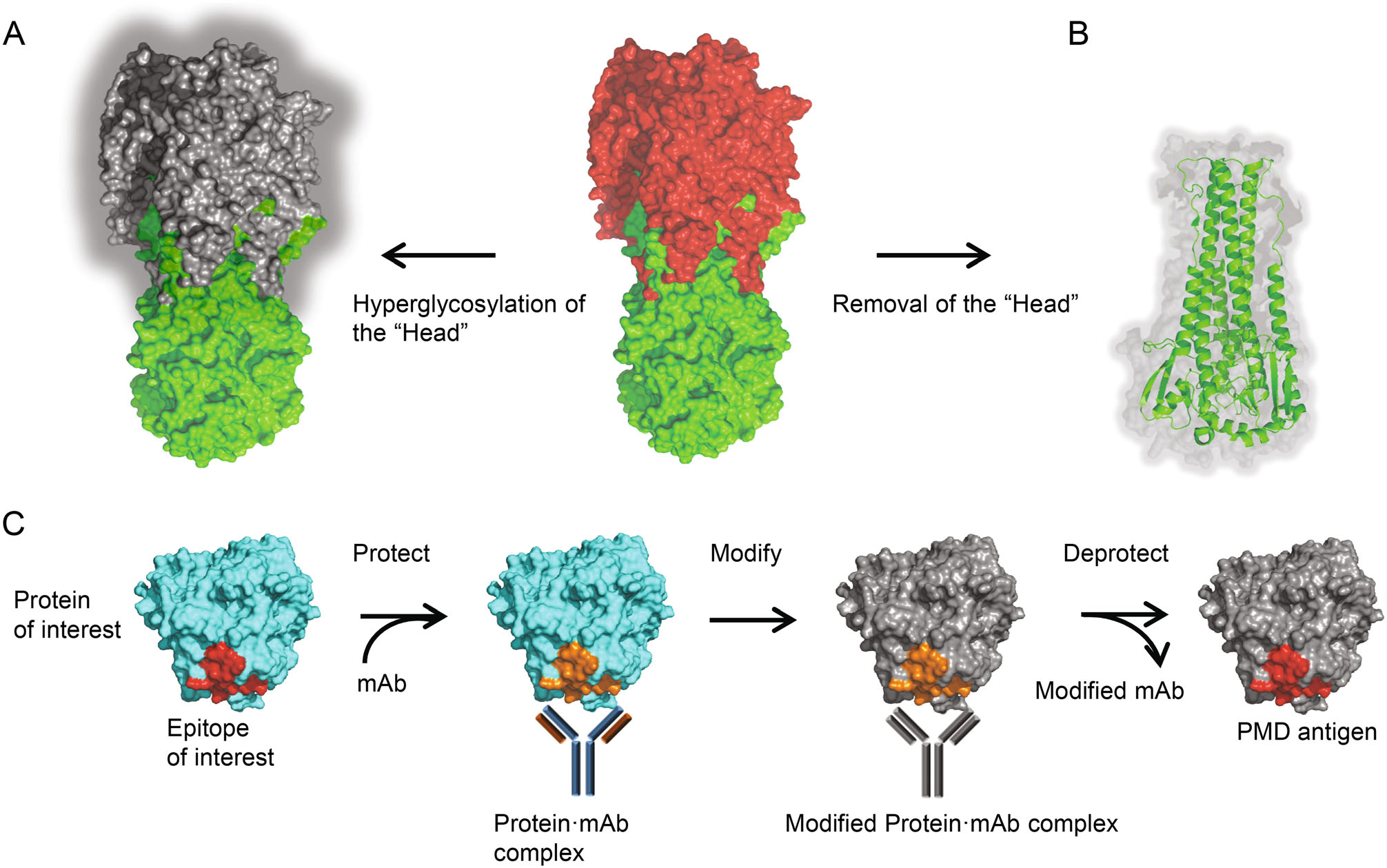
Figure 3. Immunofocusing approaches for a universal flu vaccine. A Head masking. The "Head" region of trimeric HA was hyperglycosylated, leaving only the HA stalk immunogenic. B Mini-HA. The "Head" region of trimeric HA was removed completely, while the conformation of the stalk domain maintains well. C Schematic of the PMD strategy. First, the epitope of interest is protected by binding of specific mAb. Then the surface of the protein·mAb complex is modified and rendered nonimmunogenic (shown as grey). By removal of the mAb, the epitope is deprotected and exposed as PMD antigen.
More recently, Weidenbacher and Kim developed a novel method, called protect, modify, deprotect (PMD), for creating immunogens aimed to elicit antibodies targeting a specific epitope (Weidenbacher and Kim 2019). A monoclonal antibody that recognizes the specific epitope is used to protect the target epitope on the protein, followed by modifications of the remaining exposed surfaces of the protein to render them nonimmunogenic. After removal of the monoclonal antibody, the epitope is deprotected and the resultant protein is modified at the surface other than the target epitope (Fig. 3C). The increasing numbers of broadly-neutralizing mAbs and corresponding conserved epitopes may provide attractive targets for the application of PMD in universal influenza vaccine design.
-
The success of mini-HA in inducing robust stalk-specific antibodies strongly implies that the HA stalk is not intrinsically poorly immunogenic, but is the victim of immunodominance, where the immune system tends to respond to complex antigens in a reproducibly hierarchical manner, that is, higher-ranking antigens sometimes suppress responses to lower ranking antigens (Angeletti and Yewdell 2018). In the case of IAV HA, there is clear head immunodomination over stalk.
However, immunodominance can be subverted by various strategies. For example, an "antigen imprinting" strategy was inspired by the phenomenon that natural infection and vaccination with pandemic H1N1 in 2009 preferentially boosted broadly binding antibodies in humans including antibodies targeting the stalk domain (Wrammert et al. 2011; Li et al. 2012; Pica et al. 2012). Generally, immune response against newly emerging strains of influenza virus would be affected by immune memory acquired by past influenza exposure; memory B cells that are cross-reactive with newly emerging strains is activated, while induction of strain-specific antibodies is prevented (Henry et al. 2018). In this strategy, chimeric HAs that share the same stalk domain but express different head domains were constructed and used for vaccination sequentially, allowing repeated exposure of the same stalk in the context of an irrelevant globular head domain (Fig. 4A) (Hai et al. 2012; Krammer et al. 2013; Margine et al. 2013; Ermler et al. 2017). The first immunization with a chimeric HA construct leads to a primary response against the globular head domain and, though much less efficiently, against the stalk domain. Upon subsequent booster vaccinations, a recall response against the stalk is anticipated, but immune response against the immunodominant head domain is restricted.
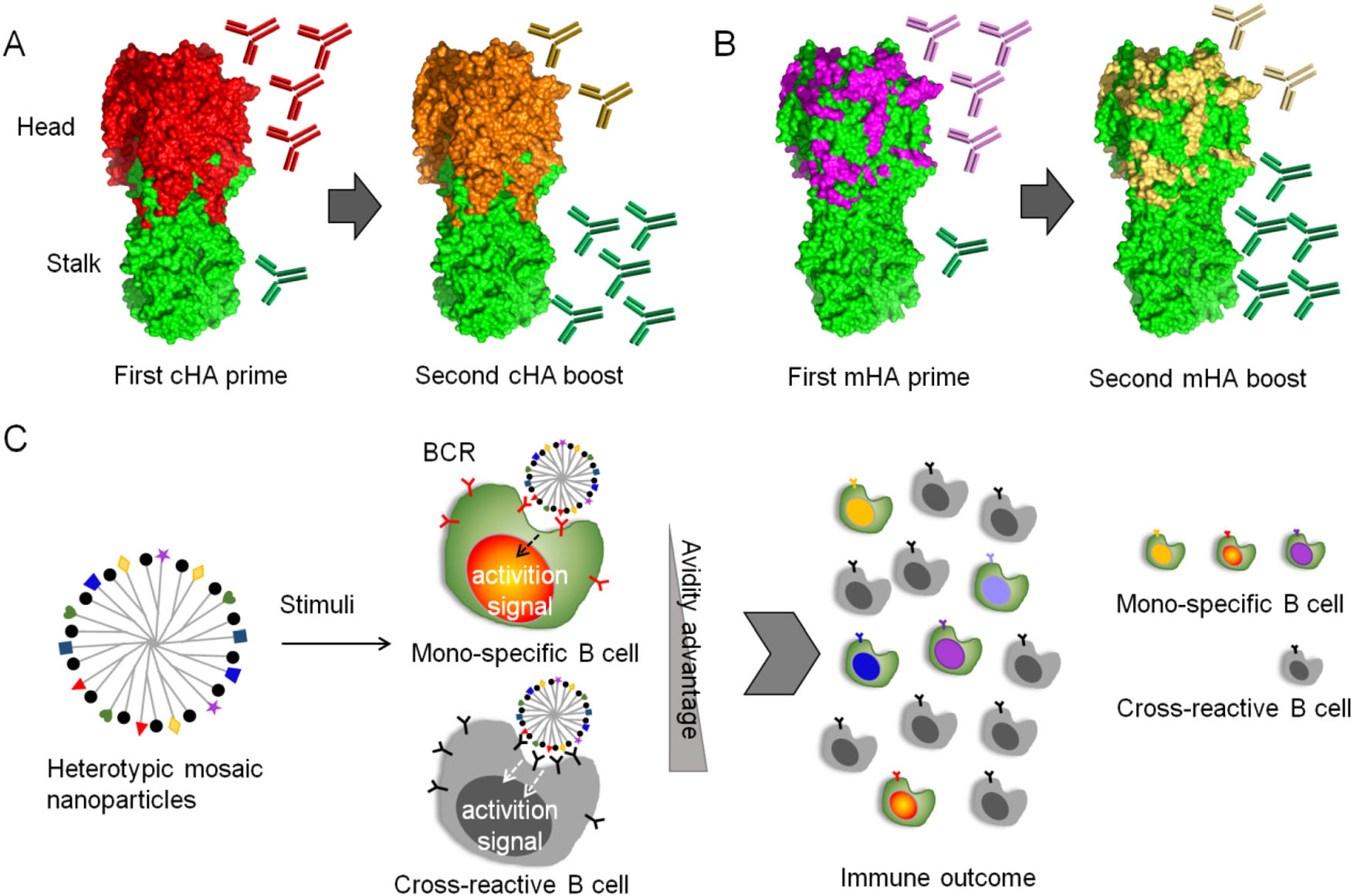
Figure 4. immunosubversion strategies for a universal flu vaccine. A Chimeric hemagglutinin (cHA) constructs and vaccination strategy. The cHAs consist of a conserved stalk domain in combination with "heads" from different avian influenza virus HA subtypes. Vaccination with the first cHA leads to primary antibody responses mainly targeting the immunodominant head domain and low-level priming against the stalk domain. Upon boosting with a second cHA that has the same stalk domain but a completely different head domain, the immune system induces a primary response against the novel head domain and a strong memory response against the stalk domain. B Mosaic hemagglutinin (mHA) constructs and vaccination strategy. Only the variable immunodominant antigenic sites in mHAs are replaced with antigenic site equivalents from different influenza virus HA subtypes, and the resulting mHA displays conserved epitopes in both the stalk and head domains. Vaccination with the first mHA likely induces a strong primary response against the grafted antigenic sites and low-level priming for conserved epitopes in the stalk and head domains. Revaccination with a second mHA with antigenic sites that have been grafted from a different subtype might result in a primary response against the antigenic sites and a strong recall response against conserved epitopes in both the head and stalk domains. C Model of immunosubversion approach with mosaic antigen array. Colored and black simbols indicates strain specific and conserved antigens respectivley. B cells possessing BCRs specific to multiple antigenic variants are colored accordingly. The avidity advantage of the mosaic antigen to cross-reactive B cells over mono-specific B cells is anticipated, promoting proliferation of cross-reactive B cells.
To further include head-specific cross-reactive epitopes into consideration for universal vaccine design, Krammer and Palese proposed a novel concept of mosaic HAs, which were constructed by replacing the immunodominant antigenic sites in the head with sequences from exotic, avian HA subtypes, instead of changing the whole head domain (Fig. 4B) (Broecker et al. 2018; Liu et al. 2018; Krammer and Palese 2019). Such a vaccination regimen can induce antibodies targeting both conserved HA stalk and conserved epitopes in the head domain, possessing improved potential as universal vaccines (Sun et al. 2019).
Recently, Kanekiyo et al. developed a novel immunosubversion strategy by enhancing the avidity of immunogens to cross-reactive B cells (Fig. 4C) (Kanekiyo et al. 2019). Heterotypic influenza hemagglutinin antigens were co-localized on a single nanoparticle, generating a mosaic array. After vaccination, the avidity advantage of conserved epitopes to B cells can be achieved over strain-specific B cells, i.e., the B cell responses can adaptively target conserved antigenic surfaces, promoting cross-reactive antibody secretion. This heterotypic mosaic nanoparticle immunogen provides a new tool to combat viral genetic plasticity and antigenic variation.
Of note, a latest study using an African green monkey IAV model revealed that although mismatched prime-boost generated a pool of stem-specific memory B cells, head-specific B cells and serum Abs still substantially dominated the immune response (Jegaskanda et al. 2019). Moreover, it's theoretically proven that, at least in prime-boost model of mice, memory B cell (MBC) clones seldom reenter secondary germinal centers upon boosting (Mesin et al. 2020). The rearrangement of MBC clones were therefore blocked, and the diversity and specificity were limited, restricting the breadth and effectiveness of the ensuing antibody response (Mesin et al. 2020). If this clonality bottleneck is also the case of humans, our ability to elicit antibodies to non-immunodominant epitopes may be further improved once the bottleneck is solved in future.
-
"Original Antigenic Sin (OAS)" refers to the phenomenon that the antibody response to the first influenza virus variant one encounters dominates the anti-influenza virus antibody response lifelong (reviewed in (Henry et al. 2018)). To date, the knowledge of how past influenza exposure shapes the response to subsequent antigenically distinct influenza strains remains obscure. Although "antigen imprinting" strategy has shown positive impact in the context of the development of a "stalk-based" universal influenza virus vaccine, the complex immune response underlying OAS more or less stymies the development of universal influenza vaccines (Cohen 2018).
Additive approach by either serial vaccination or combined vaccination with multiple antigenic types has been successfully employed to produce pneumococcal conjugate vaccines (13 different antigens) and human papillomavirus vaccines (up to nine different antigens), which can elicit accumulated responses with non-overlapping specificities. For influenza vaccines, admixture of multivalent influenza vaccines may result in antigenic competition and eventually lead to loss of efficacy for one or more components (Kanekiyo et al. 2019). Moreover, broad immunity has not been achieved by serial immunization with conventional flu vaccines (Belongia et al. 2017).
It is argued by Worobey et al. that the propensity of initial influenza virus exposure to establish a lifelong immunological imprint also presents a remarkable opportunity: immunization of infants prior to their initial, natural virus exposure with multiple versions of common human subtypes simultaneously may promise extended immunological imprinting across all currently circulating strains and against potential pandemic strains of IAV. This approach might be a possible step toward a universal vaccine (Cohen 2018; Worobey et al. 2020).
Low Immunogenicity and Immunofocusing
Immunodominance and Immunosubversion
Original Antigenic Sin Versus Additive Approach
-
Decades have been spent to concoct universal influenza vaccines, and some solid progress has been achieved. Currently, several vaccine candidates are in early human clinical trials, with each vaccine candidate exploiting conserved regions of the virus to maximize breadth [reviewed in (Nachbagauer and Krammer, 2017)]. Although some vaccine candidates such as those featured HA stalk have underwhelmed vaccine developers due to their poor effectiveness (Cohen 2018), increasing numbers of new effective broad-protective immunogens (epitopes) have been discovered, providing attractive targets that should be explored in the development of novel universal influenza vaccines (Bajic et al. 2019; Bangaru et al. 2019; Stadlbauer et al. 2019; Watanabe et al. 2019).
Of particularly note, some of the novel conserved epitopes are discontinuous (Stadlbauer et al. 2019) or even occluded in their native state (Bajic et al. 2019; Bangaru et al. 2019; Watanabe et al. 2019). How one can obtain and immunize against these epitopes remains a big issue. The advantage of synthetic epitope mimetics and de novo protein design may facilitate immunogens design for universal influenza vaccines in the future (Zerbe et al. 2017; Grayson and Anderson 2018).
-
The huge growth in high-resolution 3D structures of proteins promotes the emergence of a novel paradigm in the study of biomolecular recognition, the design and synthesis of protein epitope mimetics. Based on the natural structure of interested epitope, the mimetics (synthetic peptides) can be designed and synthesized, with diverse chemistries and cross-links (e.g., helix-stabilizing staples) harnessed to achieve desired folding (Zerbe et al. 2017).
Recent technological advances have suggested the promising potential of synthetic peptides as antigens to generate focused immune responses. By coupling a mimetic of the V3 loop from HIV-1 GP120 to synthetic virus-like nanoparticles, Riedel et al. successfully designed a candidate HIV-1 vaccine which can elicit antibodies in rabbits that recognized recombinant gp120 (Riedel et al. 2011). Moreover, both folded B- and linear T- epitopes can be displayed on the engineered nanoparticles, inducing antibody response and T-cell response respectively (Zerbe et al. 2017).
-
The computational design of new proteins sparks hopes in the field of rational vaccinology, particularly to elicit targeted neutralizing antibody responses (Correia et al. 2014; Sesterhenn et al. 2019). While most de novo proteins designed so far are either functionless or present functions that are encoded by regular, continuous secondary structures. Interestingly, a great progress has been made recently by Sesterhenn et al., who developed a novel computational design strategy to build de novo proteins presenting complex structural motifs (Sesterhenn et al. 2020). By using this strategy, the authors further engineered epitope-focused immunogens mimicking irregular and discontinuous RSV neutralization epitopes. And excitingly, the de novo-designed immunogens can induce robust neutralizing responses against the respiratory syncytial virus in both mice and nonhuman primates (Sesterhenn et al. 2020).
Synthetic Epitope Mimetics
De Novo Immunogen Design
-
The ever-changing influenza surface proteins, HA and NA, arouse the desire of a universal flu vaccine. By offering life-long protection against all influenza strains, an universal vaccine can greatly reduce the need for yearly vaccine reformulations and vaccination. So far, many universal influenza vaccine candidates that are in clinical trials focus on targeting conserved epitopes of influenza, including either surface proteins or highly conserved internal proteins. Moreover, the new identified highly conserved epitopes and advanced techniques will lead to the fast development of next-generation universal flu vaccines.
-
This work was supported by The Drug Innovation Major Project (Grant No. 2018ZX09711001), the Key Research and Development Projects of Science and Technology Department of Shandong Province (Grant No. 2017CXGC1309) and Shandong Provincial Natural Science Foundation of China (Grant No. ZR2019MH078; ZR2017MH086).
-
The authors declare that they have no conflict of interest.
-
This article does not contain any studies with human or animal subjects performed by any of the authors.













 DownLoad:
DownLoad: