-
Dear Editor,
Omsk hemorrhagic fever virus (OHFV) belongs to the tickborne virus group within Flavivirus, in Flaviviridae family (Holbrook et al. 2005) which consists of several important human pathogens including tick-borne encephalitis virus (TBEV), Powassan virus (POWV), Louping ill virus (LIV), Langat virus (LGTV) and Kyasnaur Forest disease virus (KFDV) (Yoshii and Holbrook 2009). OHFV is an enveloped virus containing a single-stranded positive-sense RNA genome of approximately 11, 000 nucleotides. The genome has a single open reading frame, which encodes three structural proteins [capsid (C), pre-membrane (prM) or membrane (M), and envelop (E)] and seven non-structural proteins (NS1, NS2a, NS2b, NS3, NS4a, NS4b and NS5) between the 5' untranslated region (UTR) and 3' UTR (Chambers et al. 1990).
Omsk hemorrhagic fever (OHF) disease was characterized as acute fever, headache, cough companied with ocular suffusion, cervical lymphadenopathy, capillary toxicosis, papulovesicular rash on the soft palate, and neurological diseases occasionally, as well as a distinct hemorrhagic syndrome including visceral hemorrhages in the nose, gums, uterus, and lungs (Ruzek et al. 2010). The fatality rate ranged from 0.5% to 2.5%. Because OHF may be mild or nearly asymptomatic in about 80% of cases, many OHF cases may be missed or misdiagnosed. Thus, the real morbidity might be higher than reported, and the case-fatality rate might be lower (Ruzek et al. 2010).
Since its first isolation in 1947 from a human presenting with hemorrhagic fever (Dobler 2010; Lani et al. 2014), OHFV was mainly prevalent in the Omsk, Tjumen, Novosibirsk, and Kurgan regions, and other regions of western Siberia has been suggested in recent years (Dobler 2010). The virus is transmitted mainly by Dermacentor reticulatus ticks. People can be infected through tick bites or contact with the blood, feces, and urine of the infected muskrats that are highly susceptible to OHFV. In addition, OHFV had also been reported to be transmitted through the milk of infected goats or sheep to humans (Holbrook et al. 2005), and laboratory infections through the contaminated aerosol had been reported (Jelinkova-Skalova et al. 1974), implying a variety of transmission routes of OHFV. Overall, the widespread presence of ticks, as well as multiple transmission routes makes OHFV be a potential threat and prevalence worldwide. Currently, no effective antivirals or approved vaccines against OHFV are available, therefore, reliable diagnostic methods for rapid discovery and surveillance of OHFV are of great importance for virus control.
Currently, the diagnosis of OHFV infection was carried out mainly by virus isolation (Dobler 2010; Lani et al. 2014), RNA detection by reverse transcriptase PCR (RT-PCR), and virus-specific serum detection including ELISA, haemagglutination-inhibition assay (Porterfield, 1957), and complement-fixation (Casals 1961) as well as neutralization tests. OHFV is classified as a biosafety level 4 (BSL-4) agent, the high requirement of the facilities to manipulate the virus limits the virus isolation and several serological methods based on the live virus. The RT-PCR assays including conventional RT-PCR and real-time RT-PCR are widely used techniques in molecular biology and diagnostics. As obvious advantages exhibited in terms of rapidity, the higher sensitivity, specificity and quantitative measurement compared to the conventional RT-PCR, the real-time RT-PCR assay has been used for detection of a variety of pathogens (Kong et al. 2009; Ho et al. 2010; Zhu et al. 2013; Hanaki et al. 2014).
Several real-time reverse-transcription PCR assays for the universal detection and identification of flaviviruses including OHFV have been published (Maher-Sturgess et al. 2008), these methods, however, have not yet been assessed specifically in the diagnosis of OHFV. Thus, we aim to establish a specific real-time RT-PCR assay using SYBR green chemistry for the quantification and detection of OHFV.
To design the specific primers for the real-time RT-PCR assay, we chose different sequences within OHFV virus strains and other TBEV group members for sequence alignment, and the region between NS4A and NS4B of the viral genome was targeted for primer design, for which sequences were highly conserved among different OHFV virus strains whereas variable between viruses of TBEV group (Fig. 1A). Four sets of primers were designed taking into account of mismatches among different strains and to avoid primer-dimer formations. The primer name, detailed sequences, and positions within the genome as well as the length of products for all oligonucleotide primers were listed in Supplementary Table S1.
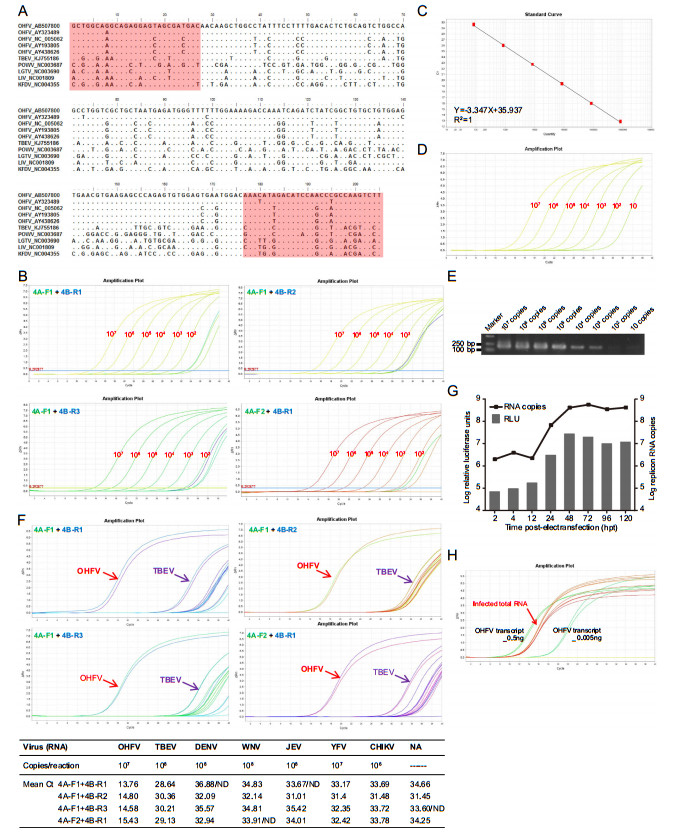
Figure 1. Development of a specific real-time RT-PCR assay using SYBR green chemistry for quantification and detection of OHFV. A Designing of primers. Diagram of sequence alignment of OHFV related TBEV group virus in flavivirus for partial NS4A and NS4B genes. GenBank accession numbers of representative sequences are specified on the left. Dot (·) indicates conserved nucleotides. Primer sequences are highlighted in light pink fields. B Amplification analysis of four sets of primers by the real-time RT-PCR assay using tenfold serially diluted in-vitro transcribed OHFV RNA. Amplification plots were exhibited for primer pairs "NS4A-F1+NS4B-R1", "NS4A-F1+NS4B-R2", "NS4A-F1+NS4B-R3", and "NS4A-F2+NS4B-R1". C Standard curve based on 4A-F1+4B-R1 primers. Standard curves for OHFV specific Green-Ⅰ-based real-time RT-PCR were generated from the CT values obtained against the tenfold serially diluted OHFV RNA ranging from 102 to 107 copies and done in triplicate. The sensitivity of real-time RT-PCR based on 4A-F1+4B-R1 primers using the real-time RT-PCR assay (D) and conventional RT-PCR assay (E). F Specificity of the OHFV real-time RT-PCR assay. Amplification plots of the real-time RT-PCR assays based on different primer pairs using OHFV and other RNA templates including other flaviviruses TBEV, DENV2, WNV, YFV and JEV, as well as an alphavirus CHIKV as templates. The CT values of each real-time RT-PCR reaction are shown in table. G Application of OHFV real-time RT-PCR assay for replicative measurement of replicon in cell culture. OHFV-Rluc replicon RNAs were transfected into BHK-21 cells, at the indicated time points after transfection, cells were lysed and Rluc activity was measured using a Multimode Microplate Reader (Varioskan Flash, Thermo Fisher, Finland) by mixing 20 μL cell lysates with 50 μL substrate (Promega). At the same time, total cellular RNAs were extracted using Trizol reagent (TaKaRa, China) according to the manufacturer's protocols and subjected to the real-time RT-PCR using 4A-F1+4B-R1 primer pair. H Detection of OHFV RNAs from virus infected cells. The BHK-21NS1 cell line were infected with replication-defective OHFV-DNS1 virus at MOI = 1. At 24 h p.i.. Total RNAs of the infected cells were extracted and subjected to real-time RT-PCR assay using all four sets of primer pairs. Different amounts of the transcribed OHFV RNAs were used as positive controls and detected by the primer pair "NS4A-F1+NS4B-R1". Three independent experiments were performed in triplicate, and the representative data were presented.
To validate the amplification ability of the four primer pairs for OHFV RNA detection based on SYBR Green-Ⅰ-based real-time RT-PCR, we employed the in vitro transcribed genomic RNA ranging from 102 to 107 copies per reaction as the template for initial experiments. As shown in Fig. 1B, all four sets of primers performed amplification with OHFV RNA efficiently, except a pair primers of 4A-F1+4B-R2 with 103 copies limit detection, others conducted amplification with 102 copies template. Besides, the 4A-F1+4B-R1 primer set performed better during the real-time RT-PCR process as they need few cycles by which the fluorescence of the sample increased to a level higher than the background fluorescence compared with other primer sets (Fig. 1B). The melting curves of the reactions using the four primer pairs were displayed in Supplementary Fig. S1 to show the specificity of the assays.
Then, the 4A-F1+4B-R1 primer pair and genomic RNAs transcribed from full-length infectious cDNA clones were used to establish the standard curves for OHFV. The standard curves were generated by plotting the threshold cycles (CT) values (Y-axis) (Supplementary Fig. S2) against the logarithm of copy numbers of serially diluted (tenfold) RNA transcripts (X-axis) with a range of 102 to 107 copies per reaction. The amplification plots and the amplicon about 190 bp of the assay were exhibited in Supplementary Fig. S2A and S2C. The linear regression equation analysis revealed that the correlation coefficient (R2) was 1 with a slope value of 3.347 (Fig. 1C).
To compare the detection limits of the SYBR Green Ⅰ-based real-time RT-PCR and conventional RT-PCR, tenfold serially diluted OHFV RNA transcripts were tested using the above two methods with specific primer pairs 4AF1+4B-R1. The detection limit of the real-time RT-PCR assay was found to be 10 copies (Fig. 1D), meanwhile, conventional RT-PCR was found to be 1000 copies which was 100-times less sensitive than real-time RT-PCR assay (Fig. 1E).
In order to assess the specificity toward the target viruses of OHFV, other RNA templates including other flaviviruses: TBEV, DENV2, WNV, YFV and JEV, as well as an alphavirus CHIKV were tested. Real-time RT-PCR assay was carried out using designed all four primer pairs with the same quantity of RNAs precisely 1.0 ng per reaction. As shown in Fig. 1F, the 4A-F1+4B-R1 primers were highly specific, the CT values of those viral RNAs were more than 32 comparable to a background signal detected in the negative control without RNA template except for the CT values of TBEV was about 29, while the CT value of OHFV was approximately 13.76, suggesting the high specificity of the designed primers targeting OHFV when comparing to other arboviruses.
To evaluate the ability of quantification for replication of OHFV genomic RNA, OHFV-Rluc replicon RNAs replicating in cell culture were parallelly measured by realtime RT-PCR and luciferase assay. Briefly, OHFV-Rluc replicon RNAs were transfected into BHK-21 cells, at the indicated time points after transfection, cells were lysed and Rluc activity was measured. At the same time, total cellular RNAs were extracted and followed by OHFV realtime RT-PCR assay and RNA copies were calculated through CT value employing standard curve established in Fig. 1C. As shown in Fig. 1G, the luciferase activities were positively correlated with the RNA copies of OHFVreplicon, suggesting that the real-time RT-PCR assay we established is a reliable method for detection of the quantification of the OHFV genome.
Further, we investigated the sensitivity of the developed qRT-PCR assay in the real condition of virus infection. As OHFV is classified as a BSL-4 agent, we used the replication-defective OHFV-DNS1 virus to infect the BHK-21NS1 cell line in which OHFV-DNS1 could replicated efficiently in BSL-2 laboratory for biosafety reasons (Zhang et al. 2019). At 24 h p.i., total RNAs of the infected cells were extracted and subjected to real-time RT-PCR assay using all four sets of primer pairs, and different amounts of the in vitro transcribed OHFV genome were used as control and detected by the primer pair "NS4A-F1+NS4B-R1". As shown in Fig. 1H, the OHFV RNAs from the virus infected cells could be detected sensitively, indicating that our developed real-time RT-PCR assay was highly sensitive to detect OHFV RNAs from cell samples.
In summary, the SYBR Green Ⅰ-based one-step real-time RT-PCR assay described here had been proven to be a simple, sensitive, specific, rapid and economic approach for the detection and quantitation of OHFV. These features will make it an excellent tool for epidemiological surveillance and clinical diagnosis of OHFV.
HTML
-
This work was supported by the National Major Science and Technology Project (2018ZX10711001-006), and the Wuhan Science and Technology Project (2018201261638501).
-
The authors declare that they have no conflict of interest.
-
This article does not contain any studies with human or animal subjects performed by any of the authors.


















 DownLoad:
DownLoad: