HTML
-
EV71 belongs to family Picornairidae, and has a single-stranded positive-sense RNA genome. First identified in California in 1969, the virus is a newly emerging health threatening pathogen in the Asia-Pacific region, including Japan, Singapore, China, and Korea (McMinn et al. 2001; Ryu et al. 2010; Shimizu et al. 1999; Singh et al. 2002). It is reported to be the main cause of hand, foot, and mouth disease (HFMD) in young children (Ooi et al. 2010). There have been millions of cases in China, with hundreds of deaths every year since 2008 (Qiu 2008). Currently, two inactivated EV71 vaccines were shown safe and efficacious in phase Ⅲ trials in China and have recently been approved for vaccination in China (Li et al. 2014; Zhu et al. 2013, 2014). However, there are presently no effective drugs for clinical treatment of EV71 infection. In order to develop effective treatment for EV71 infection, the molecular mechanism underlying its replication must first be understood.
The EV71 genome encodes four structural proteins (VP1, VP2, VP3, and VP4) and seven nonstructural proteins (2A, 2B, 2C, 3A, 3B, 3C, and 3D), and the 5′UTR of EV71's genomic RNA contains a cloverleaf-like structure and an IRES region (Shingler et al. 2013; Thompson and Sarnow 2003b). It has been reported that most IRES-dependent translation of these proteins requires the binding of canonical initiation factors, as well as IRES trans factors (ITAFs) (Holcik and Sonenberg 2005). ITAFs with multiple RNA-binding domains (RBDs) are known to bind to the IRES region (Faye and Holcik 2015), and many ITAFs have been found to interact with EV71 5′UTR, including heterogeneous nuclear ribonucleoprotein A1 (hnRNP A1), far-upstream element-binding protein 2 (FBP2), far-upstream element-binding protein 1 (FBP1), heterogeneous nuclear ribonucleoprotein K (hnRNP K), and poly(C)-binding protein 1 (PCBP1). HnRNP A1, FBP2, and FBP1 have been shown to regulate the EV71 IRES-related translation initiation activity, while hnRNP K and PCBP1 are required for the efficient synthesis of viral RNA (Huang et al. 2011; Lin et al. 2008, 2009a, b; Luo et al. 2014).
PTB has four RBDs, which are also known as RRMs (RNA recognition motif), is predominantly expressed in the nucleus, and participates in the alternative splicing of many pre-mRNAs (Auweter and Allain 2008; Spellman et al. 2005). PTB has been reported to function as ITAFs in several viruses, including poliovirus (PV), encephalomyocarditis virus (EMCV), hepatitis A virus (HAV), and hepatitis C virus (HCV) (Gosert et al. 2000; Hellen et al. 1994; Hellen and Wimmer 1995; Schmid and Wimmer 1994; Venkatramana et al. 2003). PTB can bind to the 5′UTR of the EV71 genome and compete with the negative regulator protein FBP2 for IRES binding (Lin et al. 2009a). Preventing the interaction of PTB and the EV71 IRES region through the use of quinacrine inhibits EV71 replication (Wang et al. 2013). However, the details of PTB's interaction with the EV71 IRES and how it affects the EV71 life cycle remain largely unknown.
In this study, biotinylated EV71 IRES RNA probes were used to pull-down the associated cellular proteins in T98G cells. The domains in EV71 IRES which were responsible for PTB binding to the EV71 IRES were studied. And the impact of PTB on EV71 translation initiation and viral production was also assayed. This study aimed to identify the cellular protein interaction with EV71 IRES and reveal the molecular mechanism of this interaction to improve our understanding of the EV71 life cycle, and to identify potential therapeutic targets of EV71 virus infection.
-
Human glioblastoma cell line T98G, human bone marrow neuroblastoma cell line SH-Sy5Y, and human embryonic kidney cell line 293ET were purchased from the ATCC and cultured according to the recommended guidelines (http://www.atcc.org). Human embryonal rhabdomyosarcoma cells (RD) from the Cell Center of Peking Union Medical College were cultured in Dulbecco's modified Eagle's medium (DMEM) with 10% fetal bovine serum (FBS), 100 U/mL penicillin, and 100 μg/mL streptomycin sulfate (Invitrogen, USA). Human astrocyte cell line HA was purchased from ScienCell Research Laboratories and cultured in astrocyte medium (catalog 1801). All cells were maintained at 37 ℃ in a 5% CO2 incubator.
EV71 virus strain (Human enterovirus 71 strain 87-2008 Xi'an Shaanxi, GenBank ID: HM003207.1) was kindly supplied by Professor Wenbo Xu in Chinese Center Disease Control and Prevention. Cells were infected by EV71 and incubated at 37 ℃ for 1 h. Unbound viruses were washed away by PBS. Virus titration was determined by a modified plaque assay using RD cells (Deng et al. 2012). Generally, RD cells (2 × 105 cells) were seeded onto a 12-well plate, incubated overnight, and then infected with serially diluted virus suspension. After incubation for 1 h, the virus suspension was replaced with DMEM containing 2% FBS and 0.8% methylcellulose. After 96 h post-infection, the medium was removed. The cells were fixed with 10% formaldehyde and stained with 1% crystal violet. The virus titer was expressed as PFU (plaque forming unit) per mL.
-
To make full-length and truncated PTB expression constructs, cDNA from T98G cells was used as template for amplification by specific primers (Supplementary Table S1). The resultant PCR products were cloned into pcDNA4.0 vector (Invitrogen, USA). The secondary structure of EV71 5′UTR was predicted by the mfold web server and used to design full-length and truncated IRES probes (Bellaousov et al. 2013), which were then amplified from EV71 cDNA using specific primers (Supplementary Table S1). The EV71 IRES element contains five major stem-loops (SL Ⅱ-Ⅵ). The PCR products were cloned into pGEM-3zf vector (Promega, USA) to generate pGEM-3zf-IRES, pGEM-3zf-SL Ⅱ (121-180), SL Ⅲ (190-230), SL Ⅳ (241-450), SL Ⅴ (451-563), SLⅥ (564-742, stem-loop Ⅵ, and linker region), respectively. All the constructs were confirmed by DNA sequencing.
The siRNAs against PTB and negative control siRNA (siNC) were purchased from Invitrogen (Cat. AM16708). Antibodies used in this study were as follows: rabbit anti-PTB polyclonal (generated in our lab) antibody, mouse anti-β-actin monoclonal antibody (1:3000, Sigma, USA), mouse anti-CREB monoclonal antibody (1:1000, CST, USA), mouse anti-his monoclonal antibody (1:1000, Qiagen, Germany), rabbit anti-EV71 VP1 polyclonal antibody (1:1000, Abnova, Taiwan, China), and mouse anti-EV713C monoclonal antibody (1:200, Millipore, USA).
-
The plasmids of pGEM-3zf-IRES and six IRES truncation variants were linearized with Hind Ⅲ restriction enzyme. EV71 IRES RNA was synthesized using a T7 transcription kit and with biotin NTP (Takara, Japan, GE Healthcare, USA) following the manufacture's protocol. The RNAs were purified using mini quick spin RNA columns (Roche, USA) and analyzed on 1% agarose gels.
-
T98G, HA, RD, and SH-SY5Y cells were pelleted, and then prepared using nuclear protein extraction reagents (Thermo, USA). A Bio-Rad protein assay was used to determine the protein concentrations of cell extracts. The RNA pull-down assay protocol was modified from method previously reported (Lin et al. 2008). Biotinylated EV71 IRES was used to capture interacting proteins. Bands of interest were excised, digested in gel with trypsin, and identified by Bruker Ultraflex matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS).
-
Mass data were exported to the Biotool 2.0 software package, and the Mascot (http://www.matrixscience.com) algorithm was used to identify the proteins.
-
T98G cell extract was prepared and used for co-immunoprecipitation after 6 h of EV71 infection at a multiplicity of infection (MOI) of 20. Lysates were pre-cleaned by incubation with protein A-agarose beads (50% in lysis buffer) on ice for 1 h. After centrifugation at 10, 000 ×g at 4 ℃ for 10 min, the supernatants were used for the immunoprecipitation assay. 100 μL of lysate, 400 μL of lysis buffer, and 10 μL of PTB antibody were mixed together and incubated on ice for 2 h. Pre-washed protein A-agarose beads (v/v, 50% in PBS) were added into the mixtures and incubated on ice for another 1 h. Mixtures were pelleted by centrifugation at 1000 ×g at 4 ℃ for 5 min and washed five times with lysis buffer. Each reaction mixture was resuspended in 400 μL of proteinase K buffer (100 mmol/L Tris-HCl, pH 7.5, 12.5 mmol/L EDTA, 150 mmol/L NaCl, 1% SDS, and 100 μg of proteinase K) and incubated for 30 min at 37 ℃. RNA was extracted with phenol/chloroform. RT-PCR was performed, with the fragments amplified using primers specific to either the EV71 5′UTR or to the ribosomal protein S16 (Supplementary Table S1).
-
T98G cells were seeded on 20-mm cover slips to 90% confluence in 24 wells and infected with EV71 at MOI of 5. Culture medium was removed, and the cells were washed three times with PBS 6 h post-infection, then fixed with 4% PFA for 20 min at room temperature. Cells were blocked in PBS containing 5% BSA for 1 h at 37 ℃ and incubated with antibodies at 4 ℃ overnight. Next, the cells were washed three times with PBS, and incubated with secondary antibody: either donkey anti-mouse antibody labeled with Alexa Fluor 488 (CA-21202, Invitrogen), or donkey anti-rabbit antibody labeled with Alexa Fluor 594 (A-21207, Invitrogen). Images were captured using an Olympus microscope and digital camera system (BX51 and DP70, or BX53 and DP72, Olympus).
-
T98G cells were transfected with either PTB siRNA or pcDNA4.0-PTB. After two days, dicistronic construct pRF or pRF-EV71 IRES was co-transfected with siRNA duplexes into T98G cells. Forty-eight hours later, Renilla luciferase (Rluc) and firefly luciferase (Fluc) activities were determined in a LB9507 bio-luminometer using a dual-luciferase reporter assay (Promega, USA) according to the manufacturer's instructions. PTB expression was determined by Western blotting of cell lysates.
-
Cells were harvested and washed three times with PBS, and then incubated with TNTE lysis buffer (50 mmol/L Tris-HCl, pH 7.4, 150 mmol/L NaCl, 1 mmol/L EDTA, 10 mmol/L sodium pyrophosphate, 0.5% Triton X-100, 1 mmol/L sodium vanadate, and 25 mmol/L sodium fluoride) supplemented with protease inhibitors (2 mg/mL aprotinin, 2 mg/mL pepstain, 2 mg/mL leupeptin and 2 mg/mL PMSF) on ice for 30 min to extract cells proteins. Lysates were centrifugated at 10, 000 ×g at 4 ℃ for 30 min, after which the supernatants were boiled with SDS-PAGE sample buffer for 10 min. The cell lysates were electrophoresed on a 12% SDS-PAGE, and transferred to nitrocellulose membranes (Amersham, USA). The membranes were blocked with Tris-buffered saline containing 0.2% (v/v) Tween-20 and 5% non-fat dry milk, and incubated with the indicated antibodies. Blots were visualized by an x-ray film (Kodak, USA).
-
All experiments were reproducible, and each assay was repeated at least three times. The data were analyzed using a two-tailed paired Student's t test and values with P < 0.05 were considered statistically significant.
Cell Lines and Virus
Plasmids, Small Interfering RNAs, and Antibodies
In vitro Transcription
Biotinylated RNA Pull-Down Assays
Database-Searching Algorithm
Co-immunoprecipitation and RT-PCR
Immunofluorescence Microscopy
Dicistronic Expression Assay
Western Blot Assays
Statistical Analysis
-
To identify proteins that interact with the IRES of EV71, biotinylated EV71 IRES probes were used to capture the interacting proteins. As shown in Fig. 1A, compared with nonbiotinylated RNA (lane 2), the biotinylated EV71 IRES RNA enriched 6 differential bands (lane 1). Using MALDI-TOF MS analysis, 12 proteins were identified, of which PTB, poly(rC)-binding protein 2 (PCBP2), and proliferation-associated protein 2G4 (ITAF45) have been found to interact with the IRES of picornavirus (Table 1). PTB has been found to be an important protein during the life cycle of enterovirus. However, the details of PTB's interaction with the EV71 IRES remain to be determined.
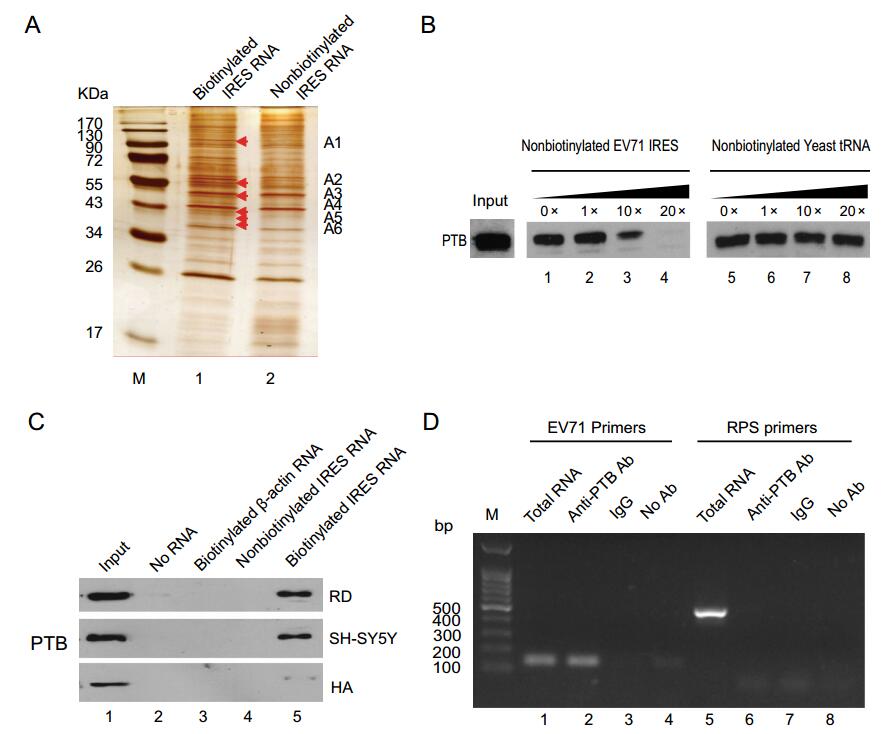
Figure 1. Interaction of PTB with the EV71 IRES region. A Pull down of proteins with the EV71 IRES in T98G cell lysate. The specific bands interacting with EV71 IRES are shown by silver staining. Lane 1: biotinylated IRES RNA, Lane 2: nonbiotinylated EV71 IRES RNA. B A specific association between PTB and the EV71 IRES region was confirmed by Western blotting and a competition assay. Increasing amounts of unlabelled RNA were added to compete with the biotin-labeled EV71 IRES RNA interacting with PTB. The eluted proteins were separated by 12% SDS-PAGE. Lanes are as follows: lanes 1-4, unlabeled EV71 IRES RNA; lanes 5-8, unlabeled yeast tRNA. C Extracts of RD cells, SH-SY5Y cells, and HA cells were prepared and then incubated without RNA (lane 2), with biotinylated actin RNA (lane 3), non-biotinylated EV71 IRES RNA (lane 4), or biotinylated EV71 IRES RNA (lane 5). After pull-down assay, the bound proteins were eluted, boiled, and subjected to 12% SDS-PAGE. PTB protein was detected by western blot with a rabbit anti-PTB antibody. The inputs were cell extracts of RD, SH-SY5Y and HA (lane 1). D EV71 IRES RNA was pulled down with PTB from EV71-infected T98G cell lysate. T98G cells were infected with EV71 at an MOI of 20 for 6 h and then cell extracts were incubated with rabbit anti-PTB antibody (lanes 2 and 6), normal rabbit IgG (lanes 3 and 7), or without antibody (- Ab) (lanes 4 and 8). Following washing and dissociation, the RNA extract was prepared and subjected to RT-PCR analysis with primers specific for the ribosomal protein S16 (PRS16) RNA (lanes1-4) or for EV71 IRES region RNA (lanes 5-8). Lane 1, cell lysate without immunoprecipitation as a positive RT-PCR control; Lane 2, anti-PTB antibody incubated with 200 mg infected-cell lysate; Lane 3, negative control with rabbit IgG; Lane 4, negative control with no antibody.

Table 1. MALDI-TOF MS results of cellular proteins associated with the EV71 IRES.
To elucidate the specific interaction of PTB and the EV71 IRES region, a competition assay and Western blotting were carried out. The specific interaction was outcompeted by non-biotinylated EV71 IRES (Fig. 1B, lanes 1-4), but not by non-biotinylated yeast tRNA (Fig. 1B, lanes 5-8). To confirm this interaction, additional cell lines were assayed. RNA pull-down and Western blot results showed that PTB interacted with EV71 IRES in all of the cell lines tested, including the RD, SH-SY5Y, and HA cell lines (Fig. 1C). Non-biotinylated IRES RNA or biotinylated β-actin probe were used as negative control.
To further confirm the results in infected cells, PTB antibody was used to pull-down the EV71 IRES. EV71 IRES was detected in total RNA extract and in the assay with PTB antibody, but not with rabbit IgG or in the absence of antibody (Fig. 1D). Primers specific to ribosomal protein S16 (RPF16) were used as negative control. These results illustrated that PTB interacts with EV71 IRES region.
-
To identify the binding region of EV71 IRES with PTB, the secondary structure of EV71 5′UTR was predicted, and found to be conserved in picornavirus (Fig. 2A). Three functional regions including cloverleaf-like (CL), IRES, and linker regions, were identified based on a previous report (Lin et al. 2008). Stem-loops Ⅱ-Ⅵ, which are important for initiation of virus translation, were contained in the EV71 5′UTR. To map the regions of interaction between PTB and EV71 IRES, six different stem-loop RNA probes were generated, including SL Ⅱ (nt 121-180), SL Ⅲ (nt 190-230), SL Ⅳ (nt 241-450), SL Ⅴ (nt 451-563), SL Ⅵ (nt 564-742), and the entire IRES (nt 90-742) (Fig. 2B). After RNA pull-down assay, Western blot results showed that PTB specially binds to the SL Ⅵ (stem-loop Ⅵ and linker region) and whole IRES, but not other SLs (Fig. 2C).
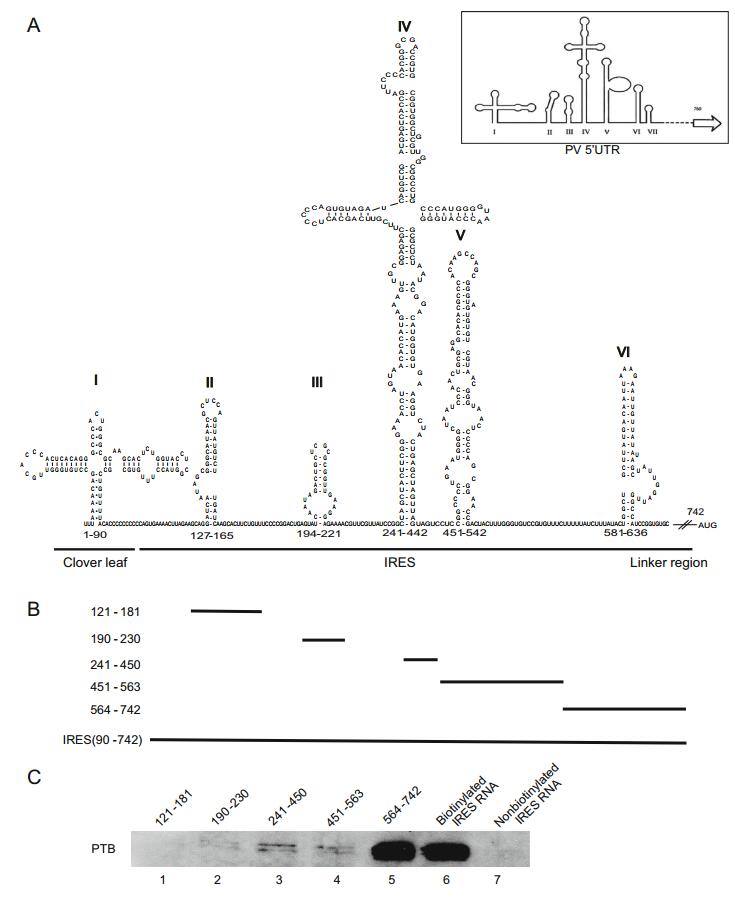
Figure 2. Determination of the EV71 IRES sequences required for the binding of PTB. A Prediction of the RNA secondary structure of the EV71 5′UTR by the mfold web serve. The modified schematic representation of the secondary structure of poliovirus (PV) 5′UTR (modified from Hellen et al. 1994) is shown in the top right corner. B Plasmids carrying different deletions in the five stem-loops of EV71 IRES: pGEM-3zf-(SL Ⅱ, 121-181), (SL Ⅲ, 190-230), (SL Ⅳ, 241-450), (SL Ⅴ, 451-463), (SL Ⅵ, 564-742). C Analysis of the regions responsible for the interaction in the EV71 IRES region using various truncated RNA forms, transcribed in vitro and biotinylated. T98G cell lysate were incubated with these biotin-labeled RNAs and the non-biotinylated IRES RNA probes were used as controls. After being pulled down by streptavidin, the protein complex was separated by SDS-PAGE and Western blot was carried out to detect PTB in the pulled-down complex (lanes 1-7).
-
PTB protein contains four RRMs (RRM1, RRM2, RRM3, and RRM4), which are important for RNA binding (Clery et al. 2008). The amino acids constituting these four RRM domains are underlined in Fig. 3A. To identify the domains that account for interaction with the EV71 IRES region, three plasmids expressing the one full-length and two truncated PTB proteins (RRM1-2, RRM3-4) were constructed (Fig. 3B). These plasmids were individually transfected into 293ET cells, and the resulting cell lysates were used in pull-down assays with biotinylated EV71 IRES. The results showed that all the three PTB proteins were expressed in transfected cells, and that they were detected in the input lanes. PTB RRM1-2 protein, but not RRM3-4 protein, was detected in the presence of biotinylated EV71 IRES (Fig. 3C). These results indicated that RRM1-2 of PTB is responsible for the binding of PTB to the EV71 IRES.
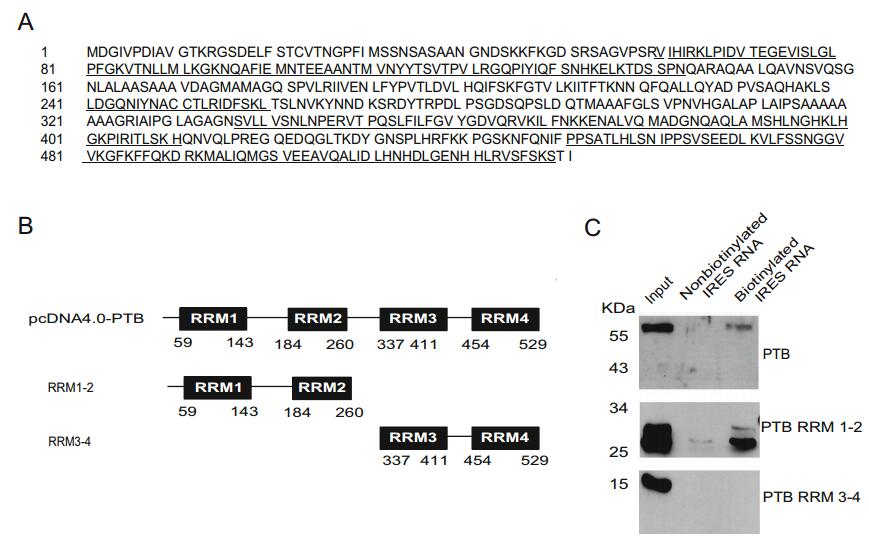
Figure 3. Analysis of the expression of PTB in neuro-related cells and RRMs required for its binding to EV71 IRES. A The RRM domains of full-length PTB protein as described. The amino acids of the four RRM domains, RRM1, RRM2, RRM3, and RRM4, are underlined. B Three constructed plasmids expressing the full-length PTB, or truncated PTB proteins in which RRM3-4 was deleted (RRM1-2), or RRM1-2 was deleted (RRM3-4), respectively. C RNA-protein binding assay showing PTB protein as determined by Western blot analysis using anti-His antibody, from 293ET cells transfected with plasmids coding the His-Tagged proteins, pcDNA4.0-PTB, RRM1-2, RRM3-4, respectively.
-
PTB is localized mainly in the nucleus, but can shuttle between nucleus and cytoplasm to complete different functions (Xie et al. 2003). To further illustrate whether EV71 infection induces changes in PTB localization, T98G cells were infected with EV71, and the localization of PTB was monitored. Confocal microscopy demonstrated that PTB was mainly expressed in the nucleus in mock cells; however, after EV71 infection, a portion of the PTB proteins were shuttled from the nucleus to the cytoplasm (Fig. 4A). To further confirm this changed expression pattern, the PTB expression pattern during EV71 infection was examined. The results showed that partial PTB protein was translocated into the cytoplasm in EV71-infected cells, while PTB was expressed exclusively in the nuclei in non-infected cells (Fig. 4B). Consistent with the confocal results, EV71 VP1 protein was exclusively located in the cytoplasm. The positive controls, β-actin and CREB, were exclusively expressed in the cytoplasm and nucleus, respectively. These results illustrated that, upon EV71 infection, PTB translocated from the nucleus to the cytoplasm, implying that PTB might be recruited by EV71 to participate the viral life cycle.
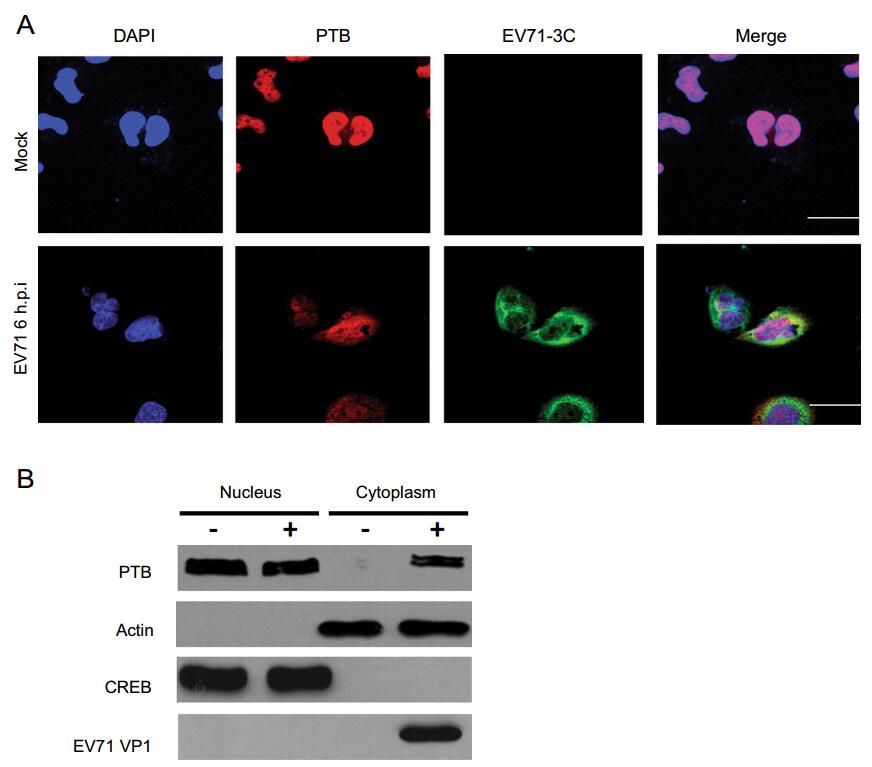
Figure 4. Subcellular distribution of PTB protein during EV71 infection. T98G cells mock-infected (upper panels) or infected with EV71 at an MOI of 20 (lower panels). At 6 h post infection, cells were fixed with formaldehyde, washed, and detected with antibody against PTB or EV71 3C protein. FITC-conjugated goat anti-rabbit IgG or TRITC-conjugated goat anti-mouse IgG was used as secondary antibody and stained with DAPI. Images were captured by confocal laser scanning microscopy. PTB (red), EV71 3C (green), DAPI (blue). Bar = 10 µm. B T98G cells infected with or without EV71 at an MOI of 20 for 6 h. Whole cell lysate, cytoplasm, and nucleus extractions were prepared from mock infected and EV71-infected cells, respectively. Endogenous proteins as detected by Western blot analysis using antibodies to CREB, EV71 VP1, PTB, or β-actin. The blot is representative of three independent experiments with similar results.
-
Cap-independent initiation of translation is a major pathway for picornavirus translation, and IRES provides an alternative means of viral polyprotein translation initiation. To evaluate the effects of PTB on EV71 IRES activity, a dicistronic reporter plasmid was used (Fig. 5A). The first cistronic (Renilla luciferase, Rluc) is cap-dependent, while the translation of the second cistronic (Firefly luciferase, Fluc) is dependent on EV71 IRES activity. One stem loop was added upstream of Rluc to prevent a high Rluc background signal arising from false initiation of translation. The Ratio of Fluc to Rluc expression could be considered as the relative IRES activity. Compared with the siNC negative control, knockdown of PTB decreased EV71 IRES activity to 70% (P < 0.05) (Fig. 5B), while overexpression of PTB increased EV71 IRES activity to 160% (P < 0.05). These results suggest that PTB can positively regulate EV71 IRES activities related to initiation of translation.
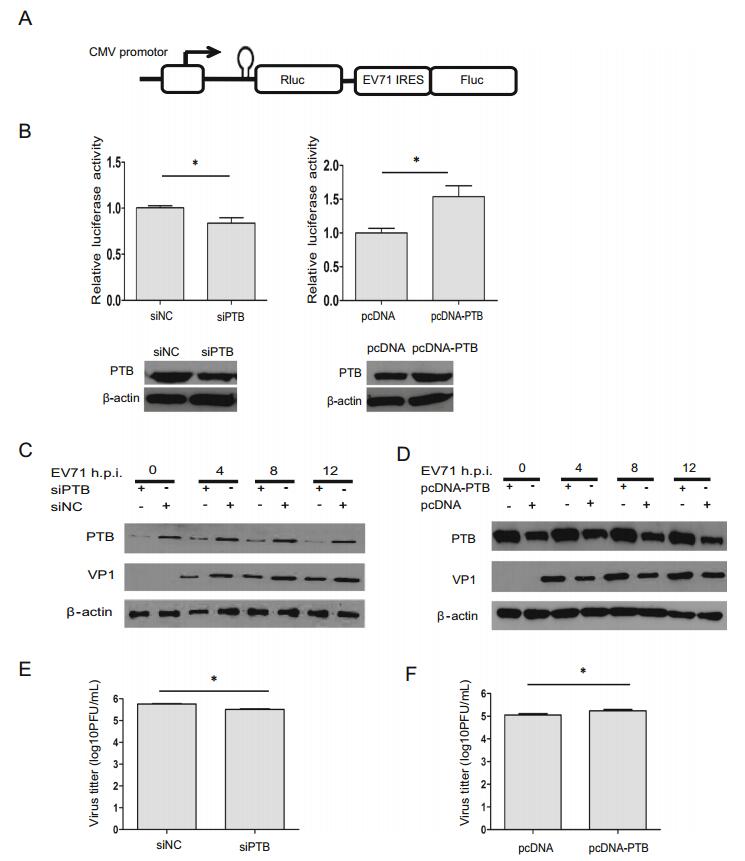
Figure 5. Analysis of PTB in the regulation of EV71 viral translation and virus production. A Schematic diagram of dicistronic reporter plasmids pRF-EV71 IRES. B T98G cells were transfected with either PTB siRNA or pcDNA4.0-PTB. 48 h later, dicistronic construct pRF or pRF-EV71 IRES and siRNA duplexes were co-transfected into T98G cells. After 48 h, the relative ISRE activity were analyzed by monitoring the luciferase activity. Western blot were used to analyze the expression level of PTB and β-actin. C-F T98G cells were transfected with siRNA targeting PTB (siPTB) and control siRNA (siNC), respectively. 48 h later, T98G cells were transfected with plasmids containing either full-length PTB cloned into pcDNA4.0 vector (pcDNA-PTB), or vector without PTB (pcDNA). After 24 h, cells were infected with EV71 at an MOI of 20 for 4, 8 and 12 h. Cell culture supernatants from EV71-infected cells were prepared and subjected to plaque assays. Western blotting was used to determine the expression level of EV71 VP1 protein, PTB or β-actin protein. E, F The effects of PTB on EV71 growth. Cell culture supernatants from C and D were determined by plaque assay. Each bar represents the average of triplicate data points with the standard deviation represented as the error bar. *P < 0.05 and **P < 0.01 versus negative control.
-
To study the roles of PTB in virus translation, the expression level of EV71 VP1 was examined. The expression level of EV71 VP1 was decreased in PTB knockdown cells at 4 h, 8 h, and 12 h post-EV71 infection, respectively (Fig. 5C), while the expression level of EV71 VP1 was increased in PTB-overexpression cells during infection (Fig. 5D), indicating that PTB promotes the expression of viral protein.
The effects of PTB on EV71 growth were also evaluated. As illustrated in Fig. 5E, 5F, virus production and viral titer in EV71-infected cells were enhanced by overexpression of PTB, but reduced by knockdown of PTB, suggesting that PTB positively regulates viral replication.
PTB Interacts with EV71 Viral RNA in IRES Region
PTB Binds to the Stem-loop Ⅵ within the EV71 IRES Region
RRM1-2 of PTB Interacts with EV71 IRES
PTB Transfers to the Cytoplasm during EV71 Infection
PTB Promotes IRES-Dependent Translation
PTB Promotes EV71 Viral Protein Expression and Virus Production
-
Translation initiation of many RNA virus genomes is cap-independent, but IRES-dependent in viral genomic RNA 5′UTR. Viral IRESs contain four major structure groups, which depend on different initiation of translation patterns, with EV71 containing a type Ⅰ IRES element (Jackson et al. 2010; Thompson and Sarnow 2003a). The interactions between viral IRES and ITAFs are crucial for understanding initiation of viral translation. ITAFs play an important role in regulating IRES-related translation initiation (Hung et al. 2016; Lin et al. 2008; Lu et al. 2004; Tolbert et al. 2017). The molecular mechanisms underlying interactions between ITAFs and EV71 IRES remain poorly understood. In this study, using a biotinylated RNA-protein pull-down assay combined with MALDI-TOF analysis, a total of 12 different proteins associated with the EV71 IRES were shown to bind to the EV71 IRES (Table 1). PTB, PCBP2, and ITAF45 have been implicated in the initiation on Type Ⅰ picornavirus IRESs (Sweeney et al. 2014). The interaction of PTB with the EV71 IRES region was demonstrated to be common to different cell lines, but relatively weak in HA cells. It has been reported that PTB expression was also strongly increased in malignant glioblastoma multiforme tumors relative to adjacent normal tissue (Wei et al. 2000). We speculate that PTB expression may be lower in HA cells compared to other cell lines. Other proteins such as SFPQ, cationic trypsinogen, and CGI-55 protein were recently found to bind to the EV71 IRES region. Further investigation into the roles of the interactions between these identified proteins and EV71 IRES in EV71 life cycle is needed.
PTB belongs to the hnRNP family, and preferentially binds to pyrimidine-rich sequences of RNA (Ghetti et al. 1992). PTB has multiple functions, which include alternative splicing, polyadenylation, stabilization, localization, and IRES-mediated translation of cellular and viral genes (Black 2003; Knoch et al. 2004; Sawicka et al. 2008; Tischendorf et al. 2004). PTB has also been reported to participate in translation initiation in many viruses such as poliovirus, EMCV, HRV14 and FMDV (Hellen et al. 1993; Niepmann 1996; Rojas-Eisenring et al. 1995; Witherell et al. 1993). In the present study, the expression of EV71 VP1 was decreased in cells with PTB knockdown, and increased in cells in which PTB was overexpressed during infection, suggesting a regulation cascade through the binding of PTB to the EV71 IRES. The predicted secondary structure of the EV71 5′UTR contained a cloverleaf-like region, IRES, and linker region. Picornavirus IRES elements are known to contain conserved secondary structures.
In poliovirus, three noncontiguous PTB-binding sites have been identified using UV cross-linking assays, and reported to be located within the 5′UTR, between nt 70 and 288, and 443 and 539 (domain V), and 630 and 730 (linker region) (Hellen et al. 1994). PTB binds to the basal half of domain V and stimulates the initiation of translation by modulating eIF4G binding (Kafasla et al. 2010). In our study, a panel of RNA transcripts corresponding to the stem-loop segments of EV71 IRES was generated. Using an RNA pull-down assay, we found that PTB acted as an ITAF and interacted with the 564-742 nt of the EV71 IRES region, which might form stem-loop Ⅵ and a linker region within the 5′UTR, and was important for the initiation of viral translation. Stem-loop Ⅵ of EV71 was reported to be the main region bound by ribosomal preinitiation complexes (Sweeney et al. 2014). Whether the linker region within EV71 5′UTR alone can interact with PTB, as in poliovirus, needs to be studied further.
In addition to loop Ⅵ of IRES region, PTB also interacts with 1-90 nt of viral RNA 5′UTR, implying that PTB may take part in viral RNA replication. Quinacrine negatively regulates EV71 infection by inhibiting the interactions between PTB and the EV71 5′UTR region (Wang et al. 2013), indicating a potential target against EV71 infection. Furthermore, the regions on which quinacrine acts to impair EV71 replication should be studied further. 3′UTR and the cloverleaf-like structure were reported to be crucial for virus replication, but whether PTB affects EV71 transcription by binding to these parts requires further investigation.
PTB has four RRMs, with flexible linkers between RRM 1 and 2, and between RRM 2 and 3. RRM 3 and 4 appear to act as a di-domain separated by a short linker (Oberstrass et al. 2005). In poliovirus, IRES activation was abrogated by the mutation of RRM1, 2, and 4, while RRM 3 appears to have few functions (Kafasla et al. 2011). Here, we found that the interaction domain of PTB with the EV71 5′UTR was mainly located in the RRM1-2 region (Fig. 3), which was revealed to be important for the interaction between PTB and the EV71 IRES, either directly or together with other cellular proteins. However, identification of the domain that plays the critical role in stimulating EV71 IRES initiation requires further research.
PTB protein is mostly accumulated in the nuclear compartment, while the EV71 life cycle mainly plays out in the cell cytoplasm. The mechanism by which PTB and the virus interact is worthy of further exploration. Viruses rely on their hosts for replication and for the maintenance of viability, and the two become engaged in a dynamic interaction during infection. Previous research has revealed that viruses can reorganize the host's secretory pathway and induce the formation of specialized organelles for viral RNA replication (Hsu et al. 2010). In DENV infection, PTB translocated noticeably from the nucleus to the cytoplasm (Agis-Juarez et al. 2009). In our immunofluorescence and Western blotting experiments, a significant portion of total PTB location was found in cytoplasm following EV71 infection. To our knowledge, it is the first evidence that PTB proteins translocate to the cytoplasm during EV71 infection, and suggests an important role for PTB in the EV71 viral life cycle. With knockdown of PTB, both the expression of EV71 VP1 and the viral titer were decreased. In contrast, when PTB was overexpressed, EV71 VP1 expression and the viral titer were both increased (Fig. 5), indicating that PTB positively regulated viral production.
In conclusion, our results reveal that PTB functions as an ITAF and promotes EV71 IRES-related translation as well as virus production. Our findings allow for improved understanding of the function of PTB in EV71 IRES-related translation and suggest potential antiviral targets in EV71 infection.
-
This study was supported by the National Basic Research Program of China (973 Program) (Grant No. 2011CB504902); the Chinese Science and Technology Key Projects (Grant No. 20132X10004601-002); the CAMS Innovation Fund for Medical Sciences (CIFMS) (Grant No. 2016-I2 M-1-004); the National Key Research and Development Program of China (Grant No. 2016YFA0100702); the Key Project of Institute of Medical Biology, the Chinese Academy of Medical Sciences (Grant No. 2018IMBCAMS001).
-
XP, BQ, and JY designed the experiments. FY, JX, GW, and BY carried out the experiments. JX, FY, WH, and ZW analyzed the data. JX and FY wrote the paper. JX, FY, ZW checked and finalized the manuscript. All authors read and approved the final manuscript.
Acknowledgements
Author Contributions
-
The authors declare that they have no conflict of interest.
-
The authors declare that they have no conflict of interest. This article does not contain any studies with human or animal subjects performed by any of the authors.
-
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Conflict of interest
Animal and Human Rights Statement
Open Access
-

Table S1. Primers used in the study.
















 DownLoad:
DownLoad: