HTML
-
Severe fever with thrombocytopenia syndrome (SFTS) is an emerging tick-mediated human infectious disease. The virus responsible for this syndrome, severe fever with thrombocytopenia syndrome phlebovirus (SFTSV), belongs to the genus Phlebovirus of the family Phenuiviridae (the order Bunyavirales) and consists of three single-stranded RNA segments, L, M and S (Jin et al. 2012; Liu et al. 2014). Following the first report of SFTS in China in 2009 (Xu et al. 2011), SFTSV was also identified in South Korea (2013) and Japan (2013) (Kim et al. 2013; Takahashi et al. 2013). Although the name of this virus is derived from the clinical symptoms of the disease, severe fever and thrombocytopenia, clinical manifestation of SFTS is nonspecific, depends on the infected host, and can be difficult to distinguish from human granulocytic anaplasmosis, leptospirosis, Orientia tsutsugamushi infection, and hemorrhagic fever with renal syndrome (Zhang and Xu 2016). Moreover, SFTSV has also been isolated from wild and domesticated farm animals in areas with high seroprevalence of SFTSV in humans (Niu et al. 2013; Zhao et al. 2012).
The immunofluorescence assay (IFA) is commonly used as a reference test to detect the antibodies of conserved viral protein in infected hosts due to its high sensitivity. However, this assay sometimes allows nonspecific interactions with similar antigens of different genus viruses resulting in poor specificity (Scott et al. 1986; Rudd et al. 2013). Further, the IFA requires use of live virus, which must be handled in special facilities with high-level biosafety equipment making processing of large numbers of samples prohibitively laborious (Xu et al. 2011). Due to these limitations, various enzyme-linked immunosorbent assays (ELISAs) and double-antigen sandwich ELISAs have been developed to detect immunoglobulins specific for the nucleocapsid protein (N) and glycoprotein N proteins (Fukuma et al. 2016; Jiao et al. 2011; Lee et al. 2016; Yu et al. 2015). Further, recent studies revealed that there is large amount of sequence diversity in SFTSV strains in East Asia (genotypes A to F) (Fu et al. 2016; Liu et al. 2016). Therefore, sensitive and specific serological diagnosis methods are needed to determine the sero-prevalence of multiple SFTSV genotypes. In our previous study, we developed antibody capture ELISAs using recombinant N and Gn proteins (rN-based ELISA and rGn-based ELISA) (Yu et al. 2018b). The results showed that while the rGnbased method worked well against homologous SFTSV strains, the rN-based ELISA test showed the ability to detect different genotypes of SFTSV.
Breeding of domesticated deer began in Korea in 1956 and by 2014 there were rapid increased up to 37, 314 head of deer on farms (MAFRA 2014). In addition, the number of domesticated deer farmers grew to 2505, underscoring the large amount of manpower required to rear deer. Numerous studies of the use of deer byproducts in traditional medicine for treatment of various diseases have been conducted by Chinese and Korean scientists (Chen et al. 2014; Guo et al. 1997). Further, there is a long tradition in East Asian countries regarding the medicinal use of deer blood and antlers.
Given the increasing incidence of SFTSV human infections and the resulting concern for public health due to the more than 20% fatality rate (Kato et al. 2016; Wang et al. 2017), we investigated SFTSV infection and its seroprevalence on breeding deer farms in Korea. To this end, serum samples were collected from farms throughout the country from 2015 to 2017, and RT-PCR and ELISA assays were performed for viral RNA detection and seroprevalence studies, respectively.
-
A total of 215 blood samples were collected in Ethylenediaminetetraacetic acid (EDTA) tubes from deer between March of 2015 and October of 2017. Blood samples were placed on ice prior to being sent to the laboratory for serum isolation. For this, samples were centrifuged at 10,000 × g for 15 min and the sera transferred to fresh tubes and stored at -80 ℃ until the time of analysis.
-
RNA was extracted from deer serum samples using the TRIzol® Reagent (Thermo Fisher, Massachusetts, USA) according to the manufacturer's instructions. Single strand cDNA was synthesized using viral RNA and primers specific for SFTSV M segments (5′-ACACAAA GACCGGCCAACACTTCA-3′) and random hexamers (Omniscript Reverse Transcriptase, QIAGEN, Hilden, Germany). Using the cDNA, PCR amplification of the M gene of SFTSV (a surface protein) was performed using a specific manufactured primer set (M gene forward: 5′CAATGCACTGGCTGCCATCT-3′ and M gene reverse: 5′-TATCCAAGGAGGATGACAATAAT-3′), also used in our previous study (Yun et al. 2017).
-
Samples in which SFTSV was detected by RT-PCR were diluted with PBS to a concentration of 1 × 101 to 1 × 103 and then used to infect Vero E6 cells in 6-well plates. Culture medium was harvested every 2 days after infection, and virus isolation was confirmed by RT-PCR using the method described above.
-
To analyze the nucleotide sequences of the M gene segment, PCR products were purified using a gel extraction kit (GeneAll, Seoul, Korea) and analyzed by sequencing (Bionics, Seoul, Korea). The sequences were aligned using MegAlign version 5 (DNASTAR Inc., USA) and compared for similarity to sequences deposited in GenBank (NCBI, USA). For phylogenetic analysis, sequence alignment and construction of the phylogenetic tree was performed using the maximum likelihood (ML) method and MEGA Version 7.0 software.
-
The rN and rGn protein of CB1 strain were purified as described elsewhere for use as ELISA coating antigens (Yu et al. 2018b). Both rN-and Gn-based ELISAs were conducted with collected deer sera as previously described (Yu et al. 2018a, b). Then, 0.2 μg per well of each purified protein in 100 μL of 0.1 mol/L carbonate-bicarbonate buffer (Sigma, St. Louis, USA) was coated onto Polysorp ELISA plates (Nunc, Rochester, NY, USA) for 12 h at 4 ℃. Plates were then blocked using 5% skim milk in 0.05% PBS-Tween 20 (0.05% PBST) for 12 h at 4 ℃ followed by five washes with 200 μL/well of 0.05% PBST to prevent non-specific binding. After the tenfold diluted deer serum was incubated on the coated plates for 3 h at 37 ℃, the plates were washed five times. Horseradish peroxidase (HRP)–conjugated anti-Deer IgG(H + L) (KPL, USA) (1:1000) was then added and plates were incubated for 2 h at 37 ℃. Following five washes with 0.05% PBST, 100 μL of the substrate solution, orthophenylenediamine peroxidase substrate (Sigma, St. Louis, USA), and 30% H2O2 in 1 mol/L citrate buffer (Sigma, St. Louis, USA) was added. Color development was terminated with 100 μL of 0.5 mol/L H2SO4. The optical density (O.D) was measured spectrophotometrically at 450 nm. All deer serum samples were considered positive or negative when compared to a cut-off value, which was calculated using the formula O.D = mean + 2 SDs (standard deviations) (Lardeux et al. 2016; Zhao et al. 2005).
-
Vero E6 cells in 6-well plates were infected with 1 mL of 1 × 103 TCID50/mL of CB1 SFTSV for 2 h at 37 ℃ and incubated for 3 days. The infected cells were fixed with 80% acetone solution prior to permeabilization with 10% Triton X-100 (Sigma, St. Louis, USA) in PBS and blocking with 3% BSA in PBS. After washing five times, diluted deer serum (1:20) samples were incubated with fixed cells for 3 h at 37 ℃, and the fluorescence was detected using fluorescein-labeled antibody against Deer IgG (H + L) (Seracare, MA, USA). The fluorescence was observed using an Olympus Ⅸ 71 (Olympus, Tokyo, Japan) microscope and DP controller software to capture images.
Sera Collection from Domesticated Deer
RNA Extraction from Sera and Reverse Transcription (RT)-Polymerase Chain Reaction (PCR) Assay
SFTSV Isolation
Sequencing and Phylogenetic Analysis
Indirect ELISA
Indirect Immunofluorescence Analysis (IFA)
-
We collected sera from 215 deer (5 to 10 samples per farm) on farms throughout South Korea from March of 2015 to October of 2017 (Fig. 1). To detect the M gene segment of SFTSV, we conducted RT-PCR with total RNA extracted from each sample. Two sera samples were positive for viral RNA, suggesting active infections of Korean deer during this time. To isolate the virus, we attempted to infect Vero E6 cells with the positive sera; however, no virus was isolated from the cell cultures following three blind passages. The M gene sequencing results revealed that the two viruses belonged to the B genotype and they showed the highest homology with KADGH/Human/Korea strain (97.0% nucleotide identity), which was isolated from a human serum sample in South Korea in 2013 (Fig. 2).
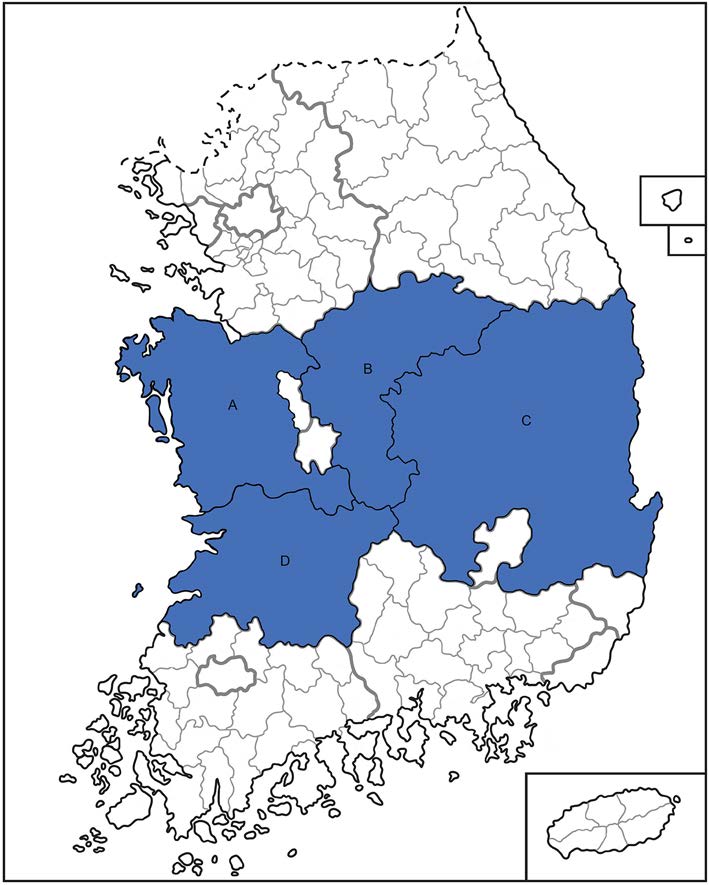
Figure 1. Sampling locations of domesticated deer serum in South Korea. Deer sera were collected from farms in Chungnam (A), Chungbuk (B), Gyeongbuk (C) and Jeonbuk (D) provinces marked by blue in Korea respectively.
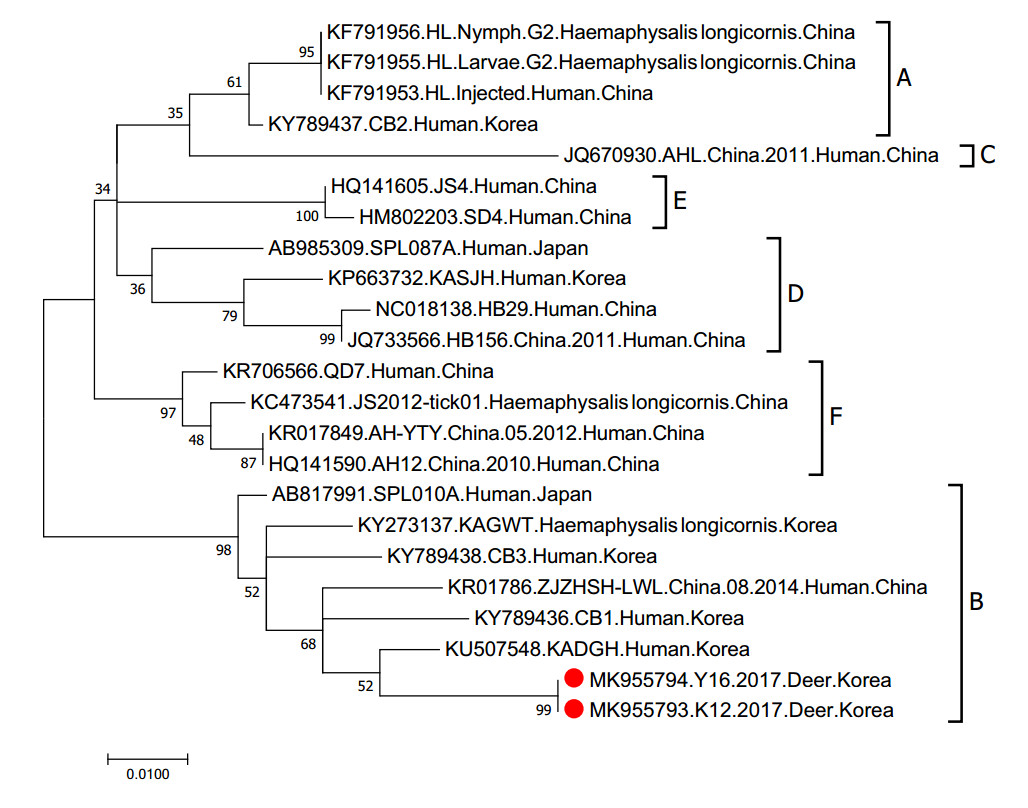
Figure 2. Phylogenetic analysis based on the nucleotide sequences of M segment of SFTSV strains. This figure shows the phylogenetic relationship of the M gene segment within SFTSV. The numbers on the branches indicate bootstrap percentages based on 1000 replications. The scale bar indicates the nucleotide substitutions per site and the tree was generated with the Maximum Likelihood (ML) method based on the Kimura 2-parameter model using MEGA Version 7.0 software. The branches were designated as SFTSV strains belonging to the genotypes A, B, C, D, E, and F, respectively. The deer-isolated SFTSV sequences analyzed in this study are marked with red closed circles. Sequenced data have been deposited in GenBank under the following accession numbers: Y16 (MK955794) and K12 (MK955793). These sequence will be released when the print version release in public.
-
To evaluate the sero-prevalence of SFTSV in domesticated deer in South Korea, deer sera were evaluated for the presence of the SFTSV antibody using two different ELISAs, rN-and Gn-based. With the rN-based ELISA, 30 (14.0%) of the 215 samples were found to be positive (O.D value greater than 0.7). The rGn-based ELISA showed 7.9% (17 of 215 specimens) of samples were positive (O.D greater than 0.7) for this surface protein. It should be noted that while only one serum sample was positive for SFTSV Gn antibody in 2015, the SFTSV sero-positive rates increased to 12.1% in 2016 and 28.2% in 2017 (Table 1). Moreover, the two samples found to be positive by RTPCR only exhibited a positive result in the rGn-based ELISA, suggesting increased genotype-specific sensitivity of the rGn-based ELISA. Overall, 40 samples were found to be positive by ELISA (Table 1). These data suggest that about 18.6% of domesticated deer were exposed to SFTSV infection over the three years surveyed and thus, exhibit potential as a reservoir for SFTSV during zoonotic infection.
Years Positive Negative (both ELISAs) Total rN* positive rGn+ positive rN and rGn Positive rN or rGn Positive 2015 0 (0.0%) 1 (2.6%) 0 (0.0%) 1 (2.6%) 38 (97.4%) 39 (100%) 2016 6 (9.1%) 3 (4.5%) 1 (1.5%) 8 (12.1%) 58 (87.9%)# 66 (100%) 2017 24 (21.8%)# 13 (11.8%) 6 (5.5%) 31 (28.2%) 79 (71.8%)## 110 (100%) Total 30 (14.0%) 17 (7.9%) 7 (3.3%) 40 (18.6%) 175 (81.4%) 215 (100%) *rN, recombinant N protein-based ELISA; +rGn, recombinant Gn protein-based ELISA.
#Indicates statistically significant differences in positive or negative rates compared with 2015 isolates as determined by Gehan–Breslow– Wilcoxon test. Statistical analyses were performed using GraphPad Prism version 8.00 for Windows (GraphPad Software, La Jolla, CA) (#indicates P < 0.05, ##indicates P < 0.0001).Table 1. The result of rN and Gn protein-based ELISAs of domestic deer serum samples.
-
To confirm the results of the ELISAs, IFA was performed with the 40 SFTSV-positive deer sera. Vero E6 cells in 6-well plates were infected with 1 × 103 TCID50/mL CB1 SFTSV strains (genotype B) and primary deer sera (1:20 dilutions) were incubated with cells for 3 h. Following the incubation, fluorescence was detected using fluoresceinconjugated secondary deer antibodies and fluorescence microscopy. For positive and negative controls, an in-house generated ferret antisera was used (described elsewhere) (Yu et al. 2018b). The IFA results show that all 30 rNand 17 rGn-based ELISA positive sera were clearly IFA positive (Fig. 3).
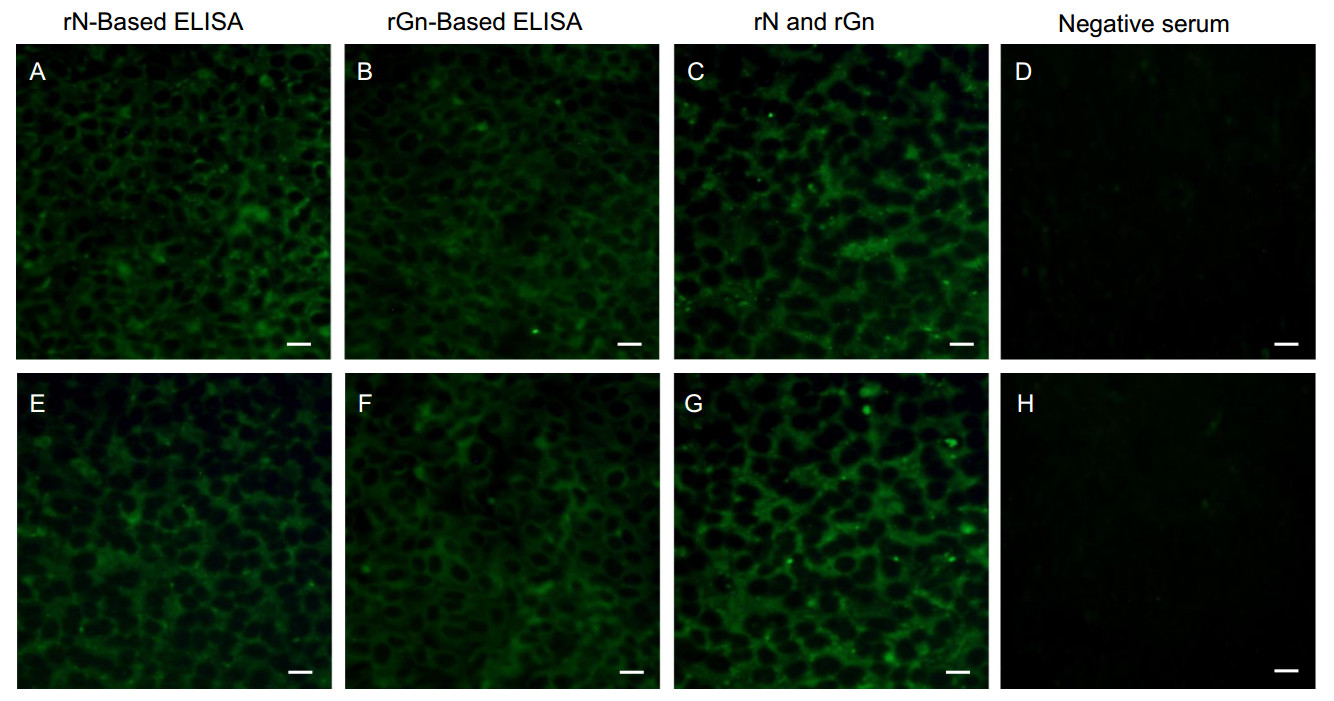
Figure 3. Immunofluorescence assay (IFA) of deer serum. IFA was performed using the rNor rGn-based ELISA positive or negative deer serum. Vero E6 cells were infected with 1 × 103 TCID50/mL CB1 SFTSV strain (genotype B) and incubated with (A, E) rN-based ELISA positive or (B, F) rGn based ELISA positive or (C, G) both rN-based ELISA and rGn based ELISA positive, or (D, H) both rNELISA and rGn ELISA negative deer sera (1:20 dilutions) and fluorescein-labeled deer IgG was used as the secondary antibody. The bar on the bottom right of each image represents the scale of 20 μm.
SFTSV Detection and Phylogenetic Tree Analysis
Evaluation of SFTSV Sero-prevalence in Breeding Deer in South Korea
IFA Assay
-
Domesticated deer have been bred in Korea since 1956 with numbers increasing to more than 30, 000 head by 2014 (Ministry of Agriculture Food and Rural Affairs, Korea). While the deer farms are relatively distributed across the country, approximately 20% of the whole breeding population is located in Chungcheong Province. In addition, the number of farm staff on domesticated deer farms is 2505, indicating a significant workforce is required for rearing deer in Korea. Further, the mean mortality rate of SFTSV cases in humans is relatively high in Japan (27%), South Korea (23.3%), and China (5.3–16.2%). (KCDC 2018; Li et al. 2018; NIID 2018; Zhan et al. 2017). Therefore, because of public health concerns, the seroprevalence of SFTSV in domesticated deer must be monitored due to the possibility of zoonotic infection by this pathogen. RT-PCR amplification revealed that two sera were positive for viral RNA, suggesting active infection of SFTSV in breeding deer in South Korea. Although we failed to isolate the virus, genetic characterization of the M gene segment revealed that the SFTSV from infected deer belonged to the B genotype (the most prevalent genotype in Korea and Japan) (Robles et al. 2018) and showed the highest nucleotide homology with the 2013 KADGH-Human-Korea isolate. Thus, this suggests that the Korean deer were infected with the same genotype as seen in human SFTSV infections.
Further, ELISA results showed approximately 18.6% of deer sera were positive for SFTSV antibodies over the course of this three-year survey. It is noteworthy that the sero-positive rate was quite low in 2015 (2.6%) and rapidly increased to 12.1% in 2016 and 28.2% in 2017, suggesting marked increases are occurring in the SFTSV infection rates of breeding deer.
In the current study, more than 18% of the domesticated deer tested were found to have been exposed to SFTSV, confirming that active infections are common on Korean farms. Further, since deer farms are generally located in thick, mountainous areas, the possibility of contact with ticks harboring SFTSV is relatively high. Therefore, special care must be taken by farmers and farm laborers to prevent SFTSV infection during the process of managing diseases and collecting deer antlers, such as wearing proper personal protective equipment to protect those directly touching animal blood and to guard against possible tick bites.
-
We thank all involved staffs in the diagnosis of SFTS during this study. We thank veterinarians at Sejeong Farms for collecting the samples. This work was supported by the Ministry of Health & Welfare (government—wide R & D fund project for infectious disease research (HG18C0029).
-
M-AY and YKC conceptualized the study. M-AY, KMY, S-JP, Y-IK, NJR, Y-JS, E-HK, and H-IK, conducted the investigation. HWJ, M-SS, S-YK and YKC wrote the paper.
-
The authors do not have any competing interests to declare.
-
This work was supported by the Ministry of Health & Welfare (government – wide R & D fund project for infectious disease research (HG18C0029). Deer serum samples for the study were collected with the deer owner's approval, and the experiments were conducted under the guidelines of the Chungbuk National University.














 DownLoad:
DownLoad: