-
Dear Editor,
Severe fever with thrombocytopenia syndrome (SFTS), caused by SFTS virus (SFTSV) in the genus Banyangvirus of the family Phenuiviridae, is an emerging tick-borne viral zoonosis (Liu et al. 2014). Typical clinical symptoms of SFTS include fever, headache, thrombocytopenia, and leukocytopenia (Yu et al. 2011). Since SFTSV was identified in 2009, it has been found in more than 20 provinces in China and is closely related to strains from Japan and South Korea (Kim et al. 2013; Takahashi et al. 2014). In Northeastern China, the disease was discovered in Liaoning Province in 2010 (Wang et al. 2016), and the virus was detected in Haemaphysalis longicornis in Jilin Province in 2013 (Liu et al. 2016). Here we describe the first fatal case of SFTS in Jilin Province and conduct phylodynamic analysis of SFTSV.
The patient was a 51-years-old woman living in Ji'an city of Jilin Province, northeastern China. She collected Chinese medicinal herbs on the mountain during June 24-25, 2018, and presented fever (with a maximum temperature of 39.4 ℃) on June 28, followed by nausea, vomiting, diarrhea, and chills. Laboratory tests showed a low white-cell count of 1650 cells per cubic millimeter, and a low platelet count of 70, 000 cells per cubic millimeter. Chest X-ray examination revealed bilateral lung texture enhancement. On July 1, the platelet count was decreased to 40, 000 cells per cubic millimeter. On July 2 (day 4), the number of platelets, white blood cells, and neutrophils decreased continuously, and proteinuria, hematuria and occult blood appeared in the urine. On July 3 (day 5), the patient showed bilateral inguinal lymph node enlargement, multiple organ dysfunction and hypoproteinemia (Supplementary Figure S1).
Leukopenia continued until day 7 and turned to a high level on day 8-9, and the same trend was observed in neutrophils, monocytes, basophils, and lymphocytes, which may be resulted from the application of glucocorticoid and granulocyte colony stimulating factor (Supplementary Figure S2 and Table S1). Erythrocyte count, hemoglobin, and hematocrit were within normal range until day 9(Supplementary Figure S2 and Table S1). The fibrinogen concentration was decreased from day 7 to day 9, and the serum D-dimer levels were elevated to more than 10, 000 μg/L (normal range: 0-232) on day 9(Supplementary Table S1). Collectively, these data indicated that disseminated intravascular coagulation occurred in the patient. After treatment with recombinant human interleukin-11, thrombopoietin, and granulocyte colony stimulating factor, the number of platelets was increased to 60, 000 cells per cubic millimeter on day 8-9 (Supplementary Figure S2 and Table S1).
Hypersensitive C-reactive protein, lactated hydrogenase, creatine kinase, creatine kinase isoenzymes, and α-hydroxybutyric dehydrogenase were significantly elevated on day 6-9. Aspartate aminotransferase and alanine aminotransferase were increased significantly from day 4 and reached an extremely high value on day 9, with a level of 351 and 3102 U per liter, respectively. The creatinine was increased slightly from day 4-7, and elevated moderately on day 8-9. Blood urea nitrogen was in normal range until day 9, with an increased concentration of 13.8 μmol/L (normal: 2.6-7.5 μmol/L). These results indicated heart, liver, and kidney dysfunction gradually aggravated during the disease course. The patient died on July 7 (day 9) (Supplementary Figure S1 and Table S2).
Based on the clinical symptoms of the patient, SFTSV was suspected to be the causative pathogen. Serum, urine, nasal and throat swabs were collected at day 9after illness onset, the pathogen was confirmed by quantitative reverse transcription PCR (RT-qPCR) with the MiniBEST Viral RNA/DNA Extraction Kit (Takara Biomedical Technology, Beijing, China). Virus-induced cytopathic effect (CPE) was observed on Vero cells inoculated with patient's serum or urine. Negative staining revealed typical viral particles of bunyaviruses (Fig. 1A). The isolated virus was designated as strain JA1-2018 (GenBank access Numbers MH822136- MH822138).
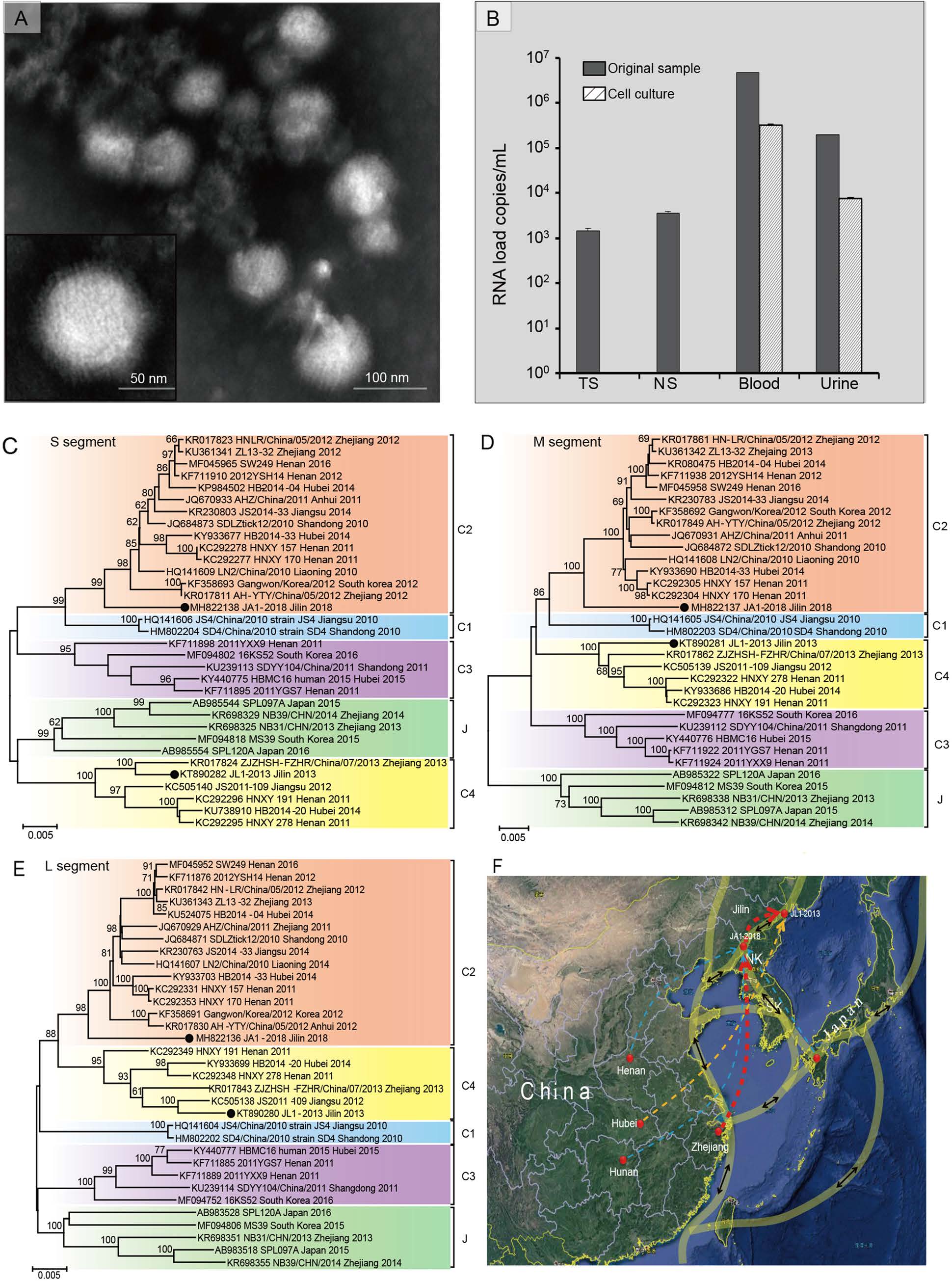
Figure 1. Identification and phylodynamic analysis of SFTSV in the first fatal patient in Jilin Province. A Negatively stained virions that purified from SFTSV-infected Vero cells show typical structural characteristics of bunyaviruses. B RNA loads of SFTSV in the nasal swabs, throat swabs, serum, and urine samples from the patient at day 9 after illness onset, and their third passages cell culture supernatants detected by realtime RT-PCR. NS, nasal swabs; TS, throat swabs. Phylogenetic analysis of S segment (C), M segment (D) and L segment (E) of SFTSV. The nucleotide sequences of S, M, and L segments were analyzed by the Neighbor-Joining method. Bootstrap testing (1000 replicates) was performed, and the bootstrap values higher than 70 are indicated. Sequences are identified by their GenBank accession number, strain name, origin, and isolated date. The two Jilin strains are marked with black dots, and different genotypes C1, C2, C3, C4, and J are represented the indicated colors. The scale bars in each panel indicate 0.005 substitutions per site. F Predicted migration routes of SFTSV strain JL1- 2013 and JA1-2018 from Jilin Province. Significant migration pathways were summarized based on the BF>3 and PP>0.5. Locations related to migration of Jilin strains are labeled with red solid circles. Red dotted arrows, decisive rates with BF>1000; orange dotted arrow, very strongly supported rates with 100 ≤ BF ≤1000; and light blue dotted arrows, weekly supported rates with 3 ≤ BF ≤10. Thickness of dotted lines represents posterior probability between 0.4 and 1.0. Light transparent yellow lines with double sided black arrows represent the possible migrating routes of migratory birds using the East Asia/ Australasia flyway. SK, South Korea; NK, North Korea.
RT-qPCR showed that RNA loads in the specimens of sera (4.71×106 copies/mL) and urine (1.98×105 copies/mL) were significantly higher than those in throat swabs (1.46×103 copies/mL) and nasal swabs (3.56×103 copies/mL). Moreover, high viral RNA loads were detected in serumcultured cell supernatant (3.17×105 copies/mL) and urinecultured cell supernatant (7.49×103 copies/mL) in the first passage, but viral RNA was not found in both throat swab and nasal swab-cultured cell supernatants in the first three passages (Fig. 1B).
Immunofluorescence assay (IFA) revealed a titer of 1:80 for IgM antibodies to SFTSV, and a titer of less than 1:20 of IgG (Supplementary Figure S3, see Supplementary Materials and Methods for details). Moreover, antibodies that against cytomegalovirus, hepatitis B and C viruses, Epstein-Barr virus, human immunodeficiency virus, and Crimen-Congo hemorrhagic fever virus were all tested negative. Taken together, the patient was diagnosed as SFSTV infection by molecular detection, virus isolation and serological testing.
We measured the levels of inflammatory cytokines and chemokines in the serum of the patients (see Supplementary Materials and Methods for details). The results showed that interleukin (IL)-6, IL-8, IL-10, IL-15, monocyte chemotactic protein (MCP)-1, granulocyte-colony stimulating factor (G-CSF), interferon-gamma-induced protein (IP)-10, and macrophage inflammatory protein (MIP)-1β levels in the patient were all significantly higher than healthy individuals, while IL-12, granulocyte/macrophage (GM)-CSF, tumor necrosis factor (TNF)-α, eotaxin, especially platelet-derived growth factor (PDGF)-BB, and RANTES were significantly reduced (Supplementary Figure S4). These results are consistent with previous studies and suggest that the inflammatory response mediated by cytokine and chemokine production imbalance is associated with the disease progression of SFTSV infection (Deng et al. 2012).
The complete sequences of the small (S), medium (M), and large (L) segments of the SFTSV strain JA1-2018 were obtained by overlapping RT-PCR (primers shown in Supplementary Table S3) and contained 1744, 3378, and 6368 nucleotides, respectively (GenBank accession numbers MH822136-MH822138). The phylogenetic trees constructed based on the S, M, and L segments presented similar topological structures as reported (reference strains information was shown in Supplementary Table S4), showing that all the strains of SFTSV were clearly clustered into five genotypes (C1-C4 and clade J) (Shi et al. 2017). SFTSV strain JA1-2018 formed a monophyletic cluster within the clade C2 strains of SFTSV, and was genetically related to isolates from Zhejiang and Henan provinces. However, the strain JL1-2013, a tick-derived SFTSV strain in Jilin, was clustered within the clade C4 (Fig. 1C-1E).
The topology of bayesian tree for SFTSV S segment showed that the included viruses were clustered into five genotypes, in which genotypes C1-C4 included the Chinese isolates while genotype J mainly contained Japanese and South Korean viruses (Supplementary Figure S5). The estimated time to most recent common ancestor (TMRCA) for all 5 genotypes was found to be 1937. The TMRCA for C2, C3, C4, and J genotypes were estimated to be 1981, 1968, 1967, and 1964, respectively. The C2 and C1 genotypes diverged from each other in 1964, and the genotypes C1 and C2 separated from genotype J in 1946, and genotype C4 diverged from genotypes C1, C2 and J in 1944. Strain JA1-2018 diverged from other strains in genotype C2 in 1981. Strain JL-1 2013 shared a TMRCA in 1995 with other five strains in the clade C4. Thus, the human strain virus in Ji'an may appear earlier than the tick strain virus in Hunchun in Jilin Province.
The migration route of SFTSV was predicted based on the time-scaled bayesian analysis. The location.rates.log file generated by BEAST software packages was loaded by SpreaD3 software (https://rega.kuleuven.be/cev/ecv/software/SpreaD3) to calculate the bayes factor (BF) and posterior probability (PP) for the diffusion between discrete locations. As shown in Fig. 1F, there were two significant migration links with decisive support, with the Bayes factor (BF) of more than 1000 and the posterior probability (PP) of 1, including one route that originated from Zhejiang Province to Ji'an, Jilin Province, where the strain JA1-2018 was isolated; and another that originated from Ji'an to Dunhua, where the strain JL1-2013 was isolated (red arrows). Strongly supported diffusion rate was also indicated from Hubei to Dunhua, Jilin Province (100 < BF < 1000, PP = 0.99) (orange arrows). Weakly support of migration links appeared originating from Henan, Hunan Province of China, or Japan to Jilin Province (3 < BF < 10) (light blue arrows).
SFTSV was detected in H. longicornis five years ago, showing that the region may be natural epidemic locus of SFTSV. The Yalu River that connects China and North Korea flowing through the Ji'an is an important habitat of migratory bird. Additionally, SFTSV has been reported in Dandong, Liaoning Province in the downstream of the Yalu River, while Ji'an is in the upstream with a minimum distance of 200 km. Migratory birds have been reported to act as long-distance carriers of tick-borne pathogens (Cerutti et al. 2018; Geller et al. 2013). SFTSVs in South Korea are genetically related to those in Japan and China, whose transmission may be mediated by migratory birds (Yun et al. 2015). Our spatial DIFFUSION analysis showed a potential migration route of SFTSV originating from Zhejiang Province to Ji'an, Jilin Province, and then from Ji'an to Hunchun, Jilin Province. Interestingly, both Ji'an and Hunchun are the stopover sites for migratory birds according to the East Asia/Australasia flyway of migratory birds (Takekawa et al. 2010). All these analyses support that the Jilin strain may originate from Zhejiang Province and is transmitted by migratory birds and H. longicornis.
In conclusion, the first fatal case of SFTSV infection was found in Jilin Province, showing that the geographical distribution of SFTSV is further expanding to northeastern China, necessitating surveillance of SFTSV infection in northeastern China, especially along the migration routes of migratory birds.
HTML
-
This study was supported by the National Key Research and Development Program of China (2017YFD0501702, and 2017YFD0500104), and the National Natural Science Foundation of China (31672542 and 31372430).
-
The authors declare that they have no conflict of interest.
-
The study was approved by the human bioethics committee of the First Hospital of Jilin University.
















 DownLoad:
DownLoad: