HTML
-
Porcine reproductive and respiratory syndrome (PRRS) is one of the most devastating global viral swine diseases caused by porcine reproductive and respiratory syndrome virus (PRRSV) (Benfield et al. 1992; Neumann et al. 2005; An et al. 2010). It is mainly characterized by severe reproductive failure in adult sows and respiratory disease in piglets, resulting in tremendous economic losses to the pork industry worldwide (Corzo et al. 2010; Lunney et al. 2010, 2016). As an enveloped single-stranded positivesense RNA virus, PRRSV belongs to the family of Arteriviridae in the order of Nidovirales (Snijder and Meulenberg 1998; Kuhn et al. 2016). Its genome is approximately 15 kb in size, including at least ten open reading frames (ORFs), of which ORF1a and ORF1b, three-quarters of the viral genome in the 5′-terminal region, encode the replication-related polymerase proteins, and the remaining quarter of the viral genome in the 30-terminal region, encodes eight structural proteins (Murtaugh et al. 2010; Kappes and Faaberg 2015).
Among the RNA viruses in the order of Nidovirales, the members of Arteriviridae including PRRSV are prone to mutate and possess the highest mutation rate due to the lack of 3′–5′ exonuclease proofreading activity (Forsberg 2005; Ogando et al. 2019). According to the differences in geographical origin and genetic variation, PRRSV is divided into two major genotypes, namely European genotype (PRRSV-1) and North American genotype (PRRSV-2), sharing about 55%–70% nucleotide homology and about 50%–80% amino acid homology (Benfield et al. 1992; Wensvoort et al. 1992; Suarez et al. 1996). PRRSV-2, the predominant PRRSV existing in China, is further divided into nine lineages based on phylogenetic analysis of ORF5 sequence (Shi et al. 2010). Classical PRRSV strains represented by CH-1a was the first PRRSV strain isolated in 1996 and had been predominant in China (Valicek et al. 1997). In 2006, the highly pathogenic PRRSV (HPPRRSV) characterized by a unique discontinuous 1 + 29 amino acid deletion in the nsp2 coding region emerged and then became the major circulating strain in pig farms in China. It threatened the commercial pork production seriously (Tong et al. 2007; An et al. 2011; Guo et al. 2012; Xie et al. 2013). Subsequently, the NADC30 strain with a characteristic deletion of 131 amino acids in the nsp2 coding region was first reported in the United States in 2008, and then the NADC30-like PRRSV isolates spread rapidly throughout China in 2013 (Zhao et al. 2015; Zhou et al. 2015, 2018a; Yu et al. 2017), that increased the genetic diversity of PRRSV.
In addition to amino acid deletions, insertions and substitutions in the PRRSV genome, recombination also played a vital and indispensable role in the evolution of PRRSV (Bian et al. 2017; Jiang et al. 2020). Currently, multiple PRRSV lineages including lineage 1, 3, 5 and 8 co-exist in swine herds in China, promoting the PRRSV recombination between different lineages (Shi et al. 2010; Gao et al. 2017; Sun et al. 2020). Recombination plays an important role in the epidemiology of PRRSV and results in the increasing number of PRRSV variants (Zhou et al. 2017, 2018a; Zhang et al. 2019). However, the virulence of these recombinant viruses between different PRRSV lineages varies greatly, especially NADC30-like PRRSVs in lineage 1, which share low sequence homology with HPPRRSV and are extremely prone to recombine with the other PRRSVs, including the wild and vaccine strains. It has been posed new challenges to effective prevention and control of PRRSV using the existing vaccines (Liu et al. 2017; Chen et al. 2018; Wang et al. 2018). Therefore, a timely understanding of the genetic evolution and pathogenic characteristics of PRRSV will be essential to provide scientific basis for the prevention of novel PRRSV variants widespread and the formulation of effective control strategies (Jiang et al. 2020; Sun et al. 2020; Xie et al. 2020). In this study, we successfully isolated a novel PRRSV variant named HBap4-2018, which was confirmed to be a natural recombinant PRRSV derived from HPPRRSV and NADC30-like PRRSV. Further analysis showed that HBap4-2018 retained the genetic and biological characteristics related to the virulence of HP-PRRSV, and was pathogenic to piglets.
-
In 2018, lung samples of dying pigs were collected from a pig farm in Hebei Province of China, where the fattened pigs suffered from severe clinical symptoms of high fever, dyspnoea and high mortality. Lung tissues were cut into pieces and homogenized in phosphate-buffered saline (PBS), and the total RNA was extracted and reverse transcribed into cDNA as described previously (Chen et al. 2019, 2021). Then the cDNA was used for PCR with specific primers designed based on the conserved ORF7 gene of PRRSV. Meanwhile, differential primers were designed based on the nsp2 gene of CH-1a (GenBank: AY032626), HuN4 (GenBank: EF635006) and NADC30 (GenBank: JN654459), so as to be identified as the classical strains (1021 bp), HP-PRRSV-like strains (931 bp) or NADC30-like strains (628 bp) based on the length of PCR products (Supplemental Table S1).
-
The supernatant of homogenate was harvested after filtration through a 0.22 μm filter, and was then inoculated onto MARC-145 for virus isolation. When cytopathic effects (CPE) on MARC-145 cells were apparent, the plates were treated with freeze-thaw cycle twice to collect the cell culture supernatant (named HBap4-2018). Subsequently, the first generation of the HBap4-2018 was inoculated onto the monolayers of MARC-145 cells grown in T25 cell culture flask for serial passage and plaque assay. The purified 5th isolation was identified based on differential primers of the nsp2 as well as the specific primers of porcine circovirus type 2 (PCV2), pseudorabies virus (PRV) and classical swine fever virus (CSFV) (Zhou et al. 2008). HP-PRRSV HuN4 strain-specific nsp2, GP5 and N protein monoclonal antibodies were used simultaneously for indirect immune fluorescent assay (IFA) to confirm the existence of isolated virus (Zhou et al. 2006, 2009). Then the growth curve of the HBap4-2018 was determined by measuring the virus titers at different time points postinfection.
-
The whole genome of HBap4-2018 was divided into eight overlapping fragments, and eight primers (Supplemental Table S1) were designed to amplify these fragments by PCR using Q5 High-Fidelity DNA Polymerase Premix (New England Biolabs, M0492L). PCR products were purified using a Gel Extraction Kit (OMEGA, D2500-01, Norcross, GA, USA) and cloned into the pMD18-T vector (TaKaRa, 6011, Dalian, China) for sequencing. Meanwhile, SMART 5′RACE and 30RACE kits (Clontech, 634858, San Francisco, USA) were used to amplify the 5′ and 3′ ends of the genome. At least three identified positive clones of each fragment were sequenced by a commercial service provider (Shanghai Sunny Biotechnology, China). Finally, the complete genome sequence of the HBap4-2018 isolate was obtained by assembling the overlapping fragments using the SeqMan program of DNASTAR7.0 software (DNASTAR Inc., Madison, WI, USA).
-
To determine the genetic evolutionary relationships between the HBap4-2018 and other representative PRRSV strains, a total of 44 reference strains with the complete genome sequences available in the GenBank database were obtained in this study (Supplemental Table S2). Phylogenetic trees based on the complete genome, ORF5 and nsp2 were constructed respectively by the neighbor-joining method (NJ) with 1000 bootstrap replicates using MEGA 6.0 (Tokyo Metropolitan University, Tokyo, Japan). And the Clustal W method of MegAlign (DNASTAR Inc., Madison, WI, USA) was used for amino acid alignment and homology analysis of nsp2 genes between HBap4-2018 and some representative PRRSV strains.
-
To detect the potential recombination events in the genome of the isolated HBap4-2018, RDP4 (University of Cape Town, Cape Town, South Africa) software including seven different algorithms (RDP, GENECONV, BootScan, MaxChi, Chimaera, SiScan and 3Seq) was used, and the recombination sites of HBap4-2018 were identified. At the same time, recombinant events and breakpoints in the genome of HBap4-2018 were verified and visualized by similarity plot analysis using SimPlot version 3.5.1 (Lole et al. 1999). Phylogenetic trees based on different regions divided by breakpoints in the genome of HBap4-2018 were constructed to support the putative recombinant events.
-
Fifteen 4-week-old piglets that were free of PRRSV, CSFV, PRV, and PCV2 were purchased from a commercial pig farm with no previous history of PRRS outbreak or PRRSV vaccination. They were randomly assigned into three groups, including a HuN4-inoculated group (n = 5), a HBap4-2018-inoculated group (n = 5), and a Dulbecco's modified eagle medium (DMEM)-inoculated group (n = 5). Pigs in each group were inoculated intramuscularly (1 mL) and intranasally (2 mL) with 3 × 105 median tissue culture infective dose (TCID50)/mL of PRRSV strain HuN4, HBap4-2018 or DMEM. Clinical symptoms and rectal temperature of each pig were recorded daily after inoculation. The weight of each pig was measured every five days, and the serum samples were collected weekly to measure the viral load by TaqMan real-time RT-PCR, and the antibodies by HerdCheck*PRRS × 3 ELISA Kit (IDEXX, 99-18070, Westbrook, USA). In addition, nasal swabs were collected every another day for virus shedding detection by TaqMan real-time RT-PCR. All the surviving pigs were euthanized at 28 days post-inoculation (dpi) for pathological examination. Lung tissues from each pig were fixed in 4% paraformaldehyde for hematoxylin-eosin (HE) staining and immunohistochemistry (IHC) examination as described previously (Chen et al. 2021), and the monoclonal antibody against PRRSV N protein was used for IHC staining diluted at 1:100 (Zhou et al. 2006).
-
All data shown in this report are expressed as the mean ± standard deviation (SD). GraphPad Prism 6 (GraphPad, La Jolla, CA, USA) was used to perform the statistical analysis, and statistical significance was assessed using Student's t-test. Differences were considered statistically significant when P-value was lower than 0.05.
Sample Collection and Detection
Virus Isolation and Identification
Complete Genome Sequencing of the HBap4-2018
Phylogenetic Analysis and Sequence Alignments
Recombination Analysis of HBap4-2018
Pathogenicity Analysis of the Isolated HBap4-2018
Statistical Analysis
-
A total of 11 lung tissue samples collected from a pig farm in Hebei Province were detected by RT-PCR, and the results showed that all the samples were PRRSV ORF7 gene positive. Subsequently, a PRRSV strain named HBap4-2018 was isolated in MARC-145 cells that inoculated with the filtered lung homogenate. The typical PRRSV-induced CPE was observed at 48 h post-infection (hpi) (Fig. 1A). The purified 5th HBap4-2018 was identified by RT-PCR, and the results showed that the length of nsp2 from HBap4-2018 was almost the same with that from NADC30-like strains, while other common pathogens including PCV2, PRV and CSFV were negative (Fig. 1B). The presence of PRRSV was confirmed by IFA staining, and the results showed that HBap4-2018 could be identified by HP-PRRSV HuN4 strain-specific GP5 and N protein monoclonal antibodies, but not the nsp2 monoclonal antibody (Fig. 1C), indicating that mutations occurred in nsp2 gene between HBap4-2018 and HP-PRRSV. Multi-step growth curve showed that the viral titer of HBap4-2018 reached the peak at 48–60 hpi, and its growth kinetics was similar with that of HP-PRRSV HuN4 in the early stages of infection, but significantly lower (P < 0.05) than that of the attenuated HuN4-F112 (Fig. 1D). In addition, the size of the plaques formed by HBap4-2018 was a little smaller than that formed by HuN4 (Fig. 1E).
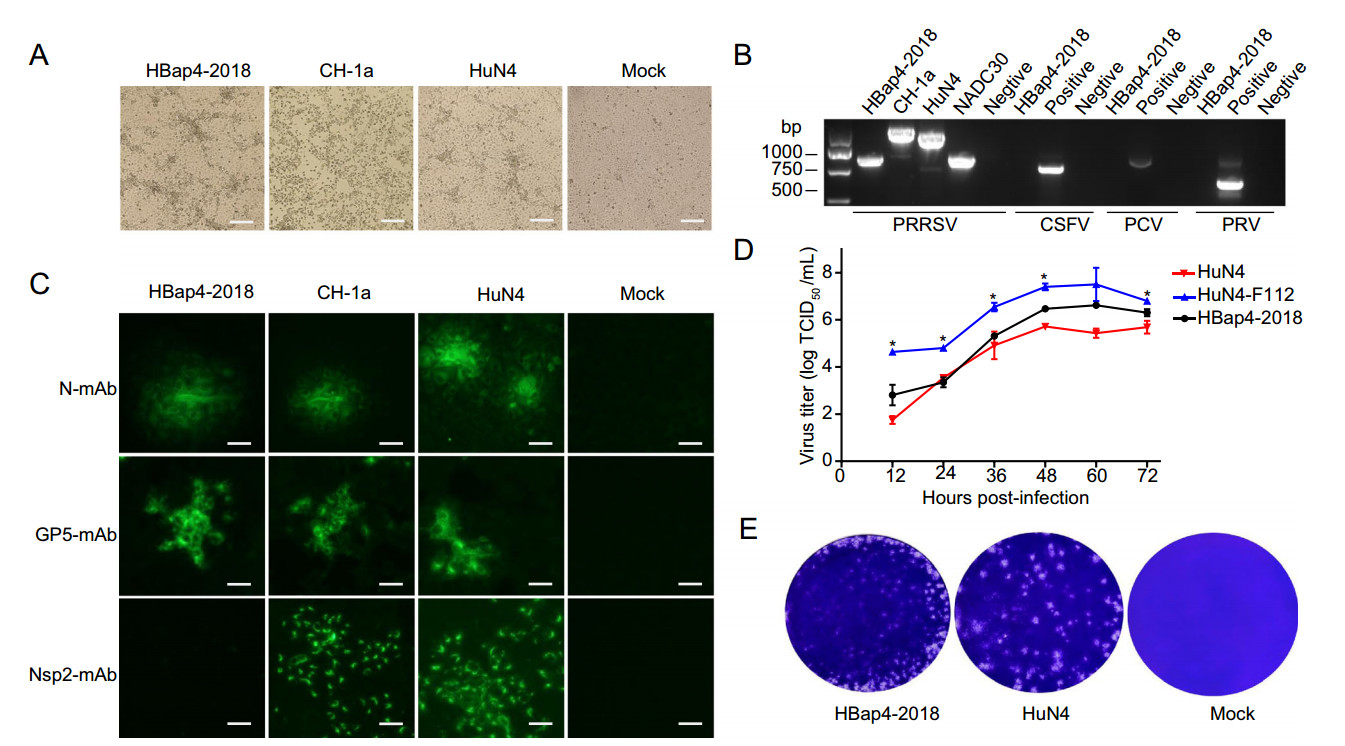
Figure 1. Isolation of PRRSV strain HBap4-2018 and identification for its biological characterization in MARC-145 cells. A Cytopathic effects (CPE) in MARC-145 cells at 48 h post-infection with HBap4-2018, CH-1a, HuN4 or mock, respectively. Scale bar = 200 μm. B RT-PCR was performed to identify the purified 5th HBap4-2018. C The presence of HBap4-2018, CH-1a or HuN4 was identified by IFA staining using HuN4 strain-specific N, GP5 and nsp2 protein monoclonal antibodies, respectively. Scale bar = 200 μm. D Multistep growth kinetics of HBap4-2018, HuN4 and HuN4-F112 in MARC-145 cells. E Crystal-violet-stained plaques formed on the monolayers of MARC-145 cells inoculated with HBap4-2018, HuN4 or mock. IFA, immune fluorescent assay; mAb, monoclonal antibody; TCID50, median tissue culture infective dose; PRRSV, porcine reproductive and respiratory syndrome virus; CSFV, classical swine fever virus; PCV, porcine circovirus; PRV, pseudorabies virus.
-
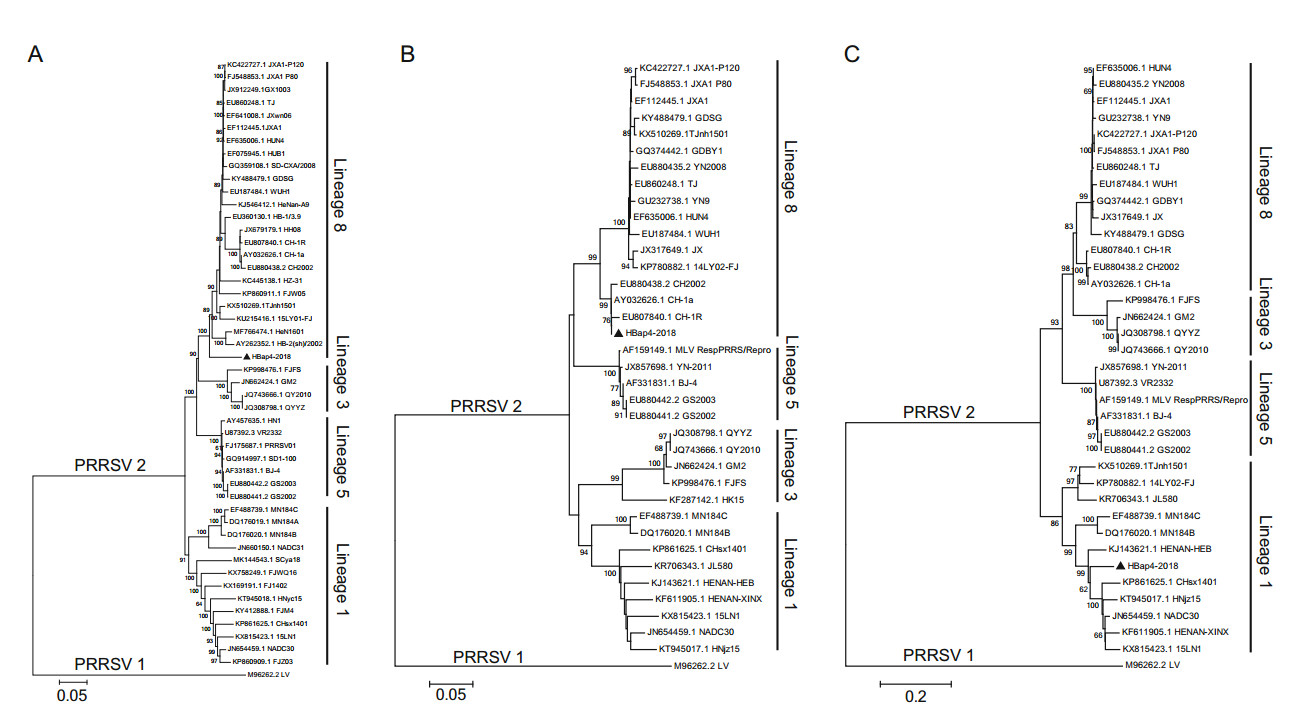
The complete genome of HBap4-2018 was 15, 003 bp in length except for the poly (A) tails, and it has been submitted to GenBank (MZ579701). HBap4-2018 shared 88.9%–89.0%, 84.9%–87.1%, and 87.8%–89.0% homology in nucleotide sequence with HP-PRRSV-like strains, classical PRRSV strains and NADC30-like strains, respectively (Table 1). Notably, nsp2 of HBap4-2018 displayed the highest amino acid homology (85.3%–85.5%) with NADC30-like strains, significantly higher than that of HP-PRRSV strains (66.8%–67.1%) and classical PRRSV strains (63.9%–64.2%) (Table 1). Amino acid sequence alignment results showed that nsp2 of NADC30-like strains had a discontinuous deletion of 131 amino acids (111 + 1 + 19 aa) as compared with VR-2332 in the region of 323–433 aa, 484 aa and 505–523 aa (Fig. 2A). Interestingly, nsp2 of HBap4-2018 had additional continuous deletion of five amino acids in the region of 463–467 aa on the basis of NADC30-like strains related region, which displayed a novel mutation pattern (111 + 5 + 1 + 19 aa) (Fig. 2B). Phylogenetic trees were constructed based on the complete genome, the ORF5 and nsp2 genes of HBap4-2018 strain as well as other representative PRRSV strains, and the results showed that HBap4-2018 belonged to the lineage 8 together with HPPRRSV strain HuN4 based on the complete genome and the ORF5 gene (Fig. 3A, 3B). However, in the analysis based on the nsp2, HBap4-2018 was located in lineage 1, which was represented by NADC30-like strains (Fig. 3C). The above results suggested that HBap4-2018 might be a recombinant virus between HP-PRRSV strains and NADC30-like strains.
Regions HP-PRRSV-like Classical PRRSV-like NADC30-like JXA1 HuN4 VR-2332 CH-1a NADC30 CHsx1401 nt aa nt aa nt aa nt aa nt aa nt aa Complete 88.9 89.0 84.9 87.1 89.0 87.8 ORF1a 80.8 80.0 80.9 80.0 78.3 78.0 79.2 77.4 89.3 89.8 87.6 87.9 ORF1b 96.9 98.2 96.9 98.4 89.9 95.5 94.2 97.1 87.8 95.9 85.9 94.9 Nsp1 95.1 94.0 95.5 94.2 88.0 93.7 92.4 90.0 84.0 83.7 82.0 82.4 Nsp2 80.8 67.1 80.9 66.8 78.3 64.2 79.2 63.9 89.3 85.5 87.6 85.3 Nsp3 83.3 90.0 83.5 90.5 84.7 91.8 83.7 90.0 92.4 97.0 90.9 95.2 Nsp4 96.7 98.0 96.7 98.0 89.0 94.1 93.6 95.6 84.0 93.1 83.3 93.1 Nsp5 94.8 94.1 95.5 94.1 85.9 88.8 91.6 90.6 87.8 91.8 85.1 88.2 Nsp6 79.9 93.8 97.9 93.8 97.9 100.0 95.8 93.8 95.8 100.0 85.4 93.8 Nsp7α 80.6 83.4 81.0 83.4 84.4 88.4 81.9 84.9 93.8 95.4 92.0 93.4 Nsp8 89.1 95.7 89.1 95.7 91.3 95.7 89.9 95.7 98.6 100.0 97.1 97.8 Nsp9 96.6 98.4 96.5 98.8 90.8 96.9 94.5 97.8 94.4 96.9 87.6 95.6 Nsp10 96.7 97.5 96.7 98.2 89.0 95.0 93.7 96.1 85.4 94.8 84.7 94.8 Nsp11 97.8 98.2 97.6 96.0 88.9 93.3 93.7 96.9 90.3 95.5 89.4 93.7 Nsp12 97.4 99.4 97.6 99.4 95.0 94.8 90.0 96.8 88.9 95.5 88.1 94.2 ORF2 99.2 96.1 99.2 96.1 93.3 92.2 96.4 93.4 86.3 87.2 86.6 87.5 ORF3 96.2 96.5 96.2 96.1 86.9 89.9 93.5 92.2 83.3 82.0 83.7 82.0 ORF4 95.0 95.5 95.9 97.8 88.9 89.9 94.3 97.2 87.5 87.7 86.0 87.7 ORF5 95.5 95.0 95.7 95.5 86.8 87.0 92.3 92.0 84.2 85.5 83.7 84.5 ORF6 94.5 97.1 94.5 97.1 92.0 94.9 93.1 95.4 90.3 94.9 89.1 94.9 ORF7 88.7 87.9 88.7 87.1 91.4 91.1 89.2 89.5 94.9 94.4 93.3 91.91 Data were shown as percentage (%).
nt, nucleotide; aa, aimino acid.Table 1. Nucleotide and amino acid homology analysis of the HBap4-2018 strain.
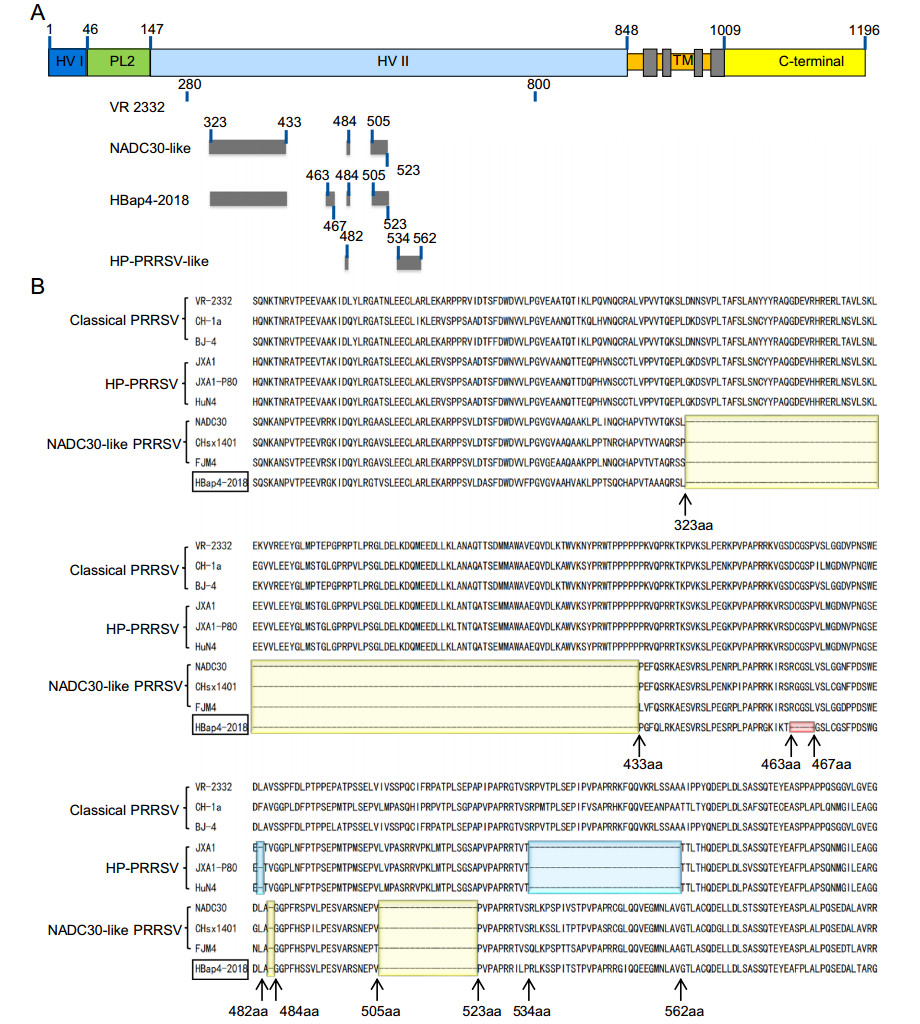
Figure 2. Amino acid sequence alignment of nsp2 from HBap4-2018 and other representative PRRSV strains. A Schematic of different deletion patterns of nsp2 from different PRRSV lineages. B Amino acid sequence of nsp2 from HBap4-2018 (marked with black box) and other representative PRRSV strains were aligned using Clustal W. Regions of nsp2 amino acid deletion in HP-PRRSV, NADC30-like PRRSV and HBap4-2018 compared with VR-2332 are highlighted in yellow, blue and red, respectively. PRRSV, porcine reproductive and respiratory syndrome virus; HP-PRRSV, highly pathogenic PRRSV.
-
Potential recombinant events in the complete genome of HBap4-2018 were analyzed using RDP4 and SimPlot software. Five recombination break points were identified in the genome of HBap4-2018 at 2000, 5105, 6292, 7412 and 14, 249 nt, which located in nsp2, nsp3, nsp5, nsp9 and ORF6, respectively. These recombination break points separated the whole genome of HBap4-2018 into six regions, namely 1–2000 nt (Region A), 2001–5105 nt (Region B), 5106–6292 nt (Region C), 6293–7412 nt (Region D), 7413–14, 249 nt (Region E) and 14, 250–15, 003 nt (Region F) (Fig. 4A). Phylogenetic trees were constructed based on these six regions, and the results showed that HBap4-2018 was located in a branch of HP-PRRSV (lineage 8) in the phylogenetic tree constructed based on Region A, C, and E. However, HBap4-2018 belonged to lineage 1. Lineage 1 was represented by NADC30-like strains in the phylogenetic tree, which was constructed based on Region B, D, and F (Fig. 4B). It was indicated that recombination events occurred in the genome of HBap4-2018, in which HP-PRRSV served as the major parental strains, while NADC30-like PRRSV served as the minor parental strains.

Figure 4. Recombination analysis of HBap4-2018. A Detection of recombination in the query genome (HBap4-2018). The x-axis showed the genomic position of HBap4-2018, while the y-axis indicated the similarity between HBap4-2018 and NADC30, JXA1, HuN4, and TJ. The genome of HBap4-2018 was divided by five break points into six regions, and the recombinant regions B, D and F were shaded orange. Number 1–12 below the ORF1a and ORF1b represents nsp1–nsp12. B Phylogenic trees were constructed based on the region A, B, C, D, E and F, respectively. Blue represents HPPRRSV, red represents NADC30-like PRRSV.
-
The pathogenicity of the recombinant PRRSV strain HBap4-2018 was evaluated in 4-week-old piglets, and all the piglets infected with HuN4 and HBap4-2018 showed typical clinical signs including depression, anorexia, dyspnea and cough, which were scored according to the clinical scoring criteria (Li et al. 2014). As shown in Fig. 5A, piglets infected with HBap4-2018 showed obvious clinical signs at the early stage of infection, even though lighter than those of the HP-PRRSV HuN4 infected piglets (Fig. 5A). Subsequently, clinical signs of piglets infected with HBap4-2018 reduced gradually and recovered to normal state at the late stage. Piglets infected with HuN4 showed high fever at 1 dpi, as the rectal temperature exceeded 40.5 ℃ for more than 3 consecutive days, and some piglets even exceeded 41 ℃ until they died (Fig. 5B). While the rectal temperature of the HBap4-2018-inoculated group began to rise to exceed 40.0 ℃ at 1 dpi, three piglets in this group even exceeded 40.5 ℃, and the other two piglets recovered gradually after 5 dpi. During the whole experimental period, the rectal temperature of the DMEM-inoculated group was maintained at the normal level. The weight of each pig was measured every five days, and the results showed that the average daily weight gains in the HuN4 and HBap4-2018-inoculated groups were significantly lower (P < 0.05) than those in the DMEM-inoculated group at the early stage of infection (Fig. 5C). While in the late stage of infection, the average daily weight gains of the two recovered piglets in the HBap4-2018-inoculated group increased gradually. Three piglets in the HBap4-2018-inoculated group died at 6 dpi, 16 dpi and 28 dpi, respectively, and the other two piglets survived with a mortality rate of 60% (Fig. 5D). By contrast, all the piglets in the HuN4-inoculated group died, yielding a 100% mortality rate, while all the piglets in the DMEM-inoculated group survived throughout the experiment.
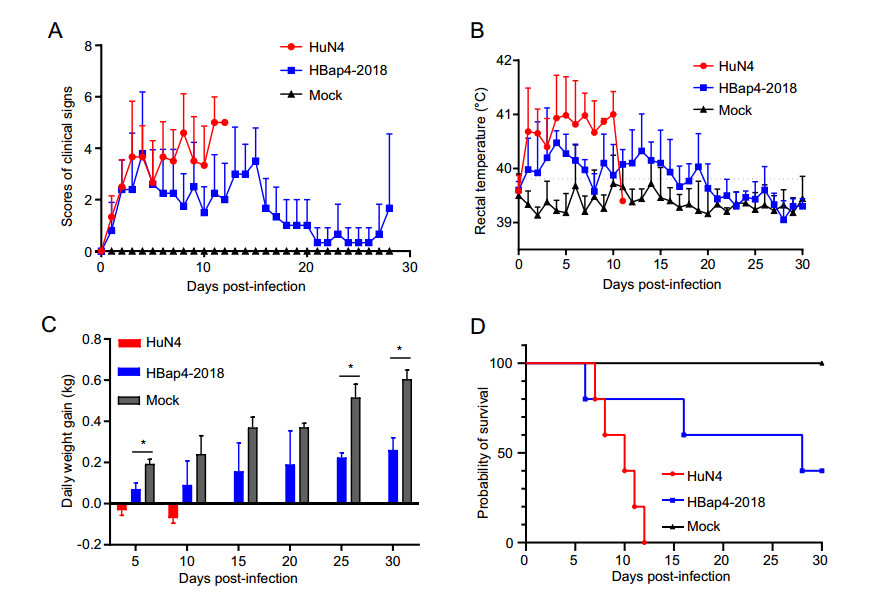
Figure 5. Clinical scores, rectal temperature, average daily weight gain and mortality rates in the piglets. A Clinical signs of piglets inoculated with HuN4, HBap4-2018 and DMEM were scored according to the clinical scoring criteria. B Rectal temperatures of piglets inoculated with HuN4, HBap4-2018 and DMEM. The fever cut-off value was set at 40.0 ℃. C Daily weight gain of inoculated piglets were shown every 5 days over 30 days. D Mortality rates in each group were shown during the whole experiment. All data were shown as the mean ± SD (error bars). SD, standard deviation.
-
The virus shedding of nasal swabs was detected by TaqMan real-time RT-PCR. We found that piglets infected with HuN4 shed virus at the high level of shedding titers ranging from 6.58 × 103 copies/mL to 2.38 × 104 copies/mL within 3–7 dpi, and peaked at 5 dpi. While in the HBap4-2018-inoculated group, high level of shedding titers could be detected within 5–7 dpi, later than that of HuN4-inoculated group. Shedding titers of piglets infected with HBap4-2018 peaked 6.54 × 104 copies/mL at 5 dpi and reduced continually, while no virus shedding was detected in DMEM-inoculated group during the whole experiment (Fig. 6A). Serum samples were collected weekly to detect the viral loads and the antibodies. As shown in Fig. 6B, piglets infected with HuN4 had the high level of viral load (1.42 × 107 copies/mL) at 7 dpi, significantly higher (P = 0.0036) than that of piglets infected with HBap4-2018. In addition, viral load in the HBap4-2018-inoculated group reached the highest level of 4.04 × 107 copies/mL at 14 dpi, a week later than that of piglets infected with HuN4. No viral load was detected in the DMEM-inoculated group throughout the experiment. PRRSV N protein antibody in serum samples was measured using a commercially available ELISA kit, and the results showed that both the HuN4 and the HBap4-2018-inoculated groups showed serologically positive (S/P [ 0.4) at 7 dpi, then antibody levels increased continuously (Fig. 6C). Piglets infected with HBap4-2018 had high levels of antibody at 21 dpi, lasting for more than a week. DMEM-inoculated group showed serologically negative (S/P < 0.4) throughout the experiment.
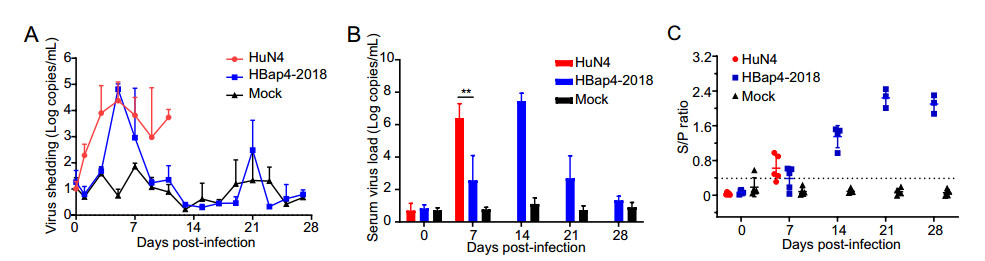
Figure 6. Virus shedding, viremia and PRRSV-specific antibody levels in the piglets. A The virus shedding of piglets inoculated with HuN4, HBap4-2018 and DMEM were detected weekly by TaqMan real-time RT-PCR. B Detection of PRRSV RNA copies in serum of each group over 28 days by RT-qPCR. C PRRSV N protein antibody in serum samples collected from all the piglets weekly were measured using a commercial IDEXX ELISA kit. All data were shown as the mean ± SD (error bars). SD, standard deviation. Serum was confirmed to be positive when S/P value > 0.4.
-
Lung tissues of piglets infected with HuN4 and HBap4-2018 showed severe macroscopic lesions including edema, hemorrhage and pulmonary consolidation on the surface, whereas no obvious macroscopic lesions were observed in the DMEM-inoculated group (Fig. 7A). Histopathological observation revealed typical interstitial pneumonia characterized by severe hemorrhage, alveolar interstitial thickening, pulmonary consolidation and infiltration of numerous inflammatory cells in alveolar spaces, while the lung tissue of the DMEM-inoculated piglets did not exhibit any histopathological lesions (Fig. 7B). In addition, IHC staining was performed to examine PRRSV antigen in the lungs of inoculated piglets, and large amounts of PRRSV antigen-positive epithelial cells were observed in the lungs of piglets inoculated with HuN4 and HBap4-2018 (Fig. 7C). However, no PRRSV antigen-positive epithelial cells were found in the lungs of piglets inoculated with DMEM, indicating that the macroscopic and histopathological lesions in the lungs resulted from PRRSV inoculation.
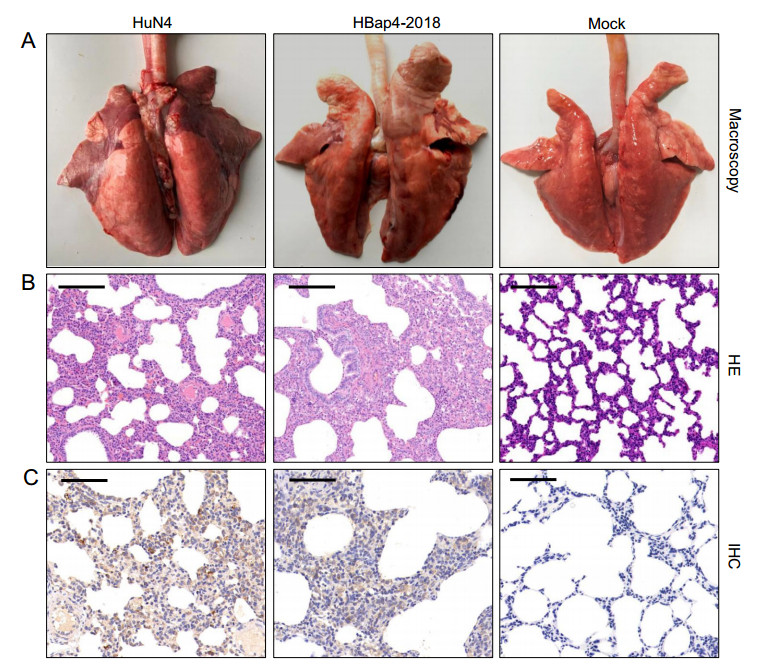
Figure 7. Macroscopic and histopathological lesions in the lungs. A Macroscopic lung lesions of piglets inoculated with HuN4, HBap4-2018 and DMEM. B Lungs of piglets inoculated with HuN4, HBap4-2018 and DMEM were stained with HE for pathological examination. C Detection of PRRSV in lung tissues of piglets inoculated with HuN4, HBap4-2018 and DMEM by IHC. Scale bar = 100 μm. PRRSV, porcine reproductive and respiratory syndrome virus; HE, hematoxylin-eosin; IHC, immunohistochemistry.
-
PRRS is one of the most pandemic and destructive diseases of the global swine industry in recent years (Lunney et al. 2010). The accelerated genetic diversity of PRRSV resulted from continuously mutating, contributed to the complicated and confusing epidemic status, which posed new challenges to the effective control of PRRSV infection using existing vaccines (Zhang et al. 2020). Analysis of the genomic characteristics of PRRSV prevalent in the field is beneficial to fully understand the prevalence status and to track the evolutionary dynamics of PRRSV, which is driven by genetic variations including amino acid deletion, insertion, substitution and particularly the genome recombination (Park et al. 2020; Yu et al. 2020; Li et al. 2021). CH-1a-like PRRSV strains have been predominant in China since the first PRRSV strain CH-1a was isolated in 1996 (Valicek et al. 1997). Ten years later, the HP-PRRSV characterized by a unique 1 + 29 amino acid deletion in the nsp2 coding region emerged and then became the major circulating strain in Chinese mainland (Tong et al. 2007; Zhou et al. 2008; An et al. 2011; Guo et al. 2012). Subsequently, the NADC30 strain with a characteristic deletion of 131 amino acids in nsp2 coding region was reported in 2013 and spread across the country (Zhao et al. 2015; Zhou et al. 2015), which threatened the pork production due to a lack of ideal cross-protection provided by current commercial vaccines. Therefore, deletions played an important role in the evolution of PRRSV. However, recombination occurred extensively and played an increasingly important role in the evolution of PRRSV after 2006. Both the HPPRRSV and the NADC30-like PRRSV had strong recombination capacities (Jiang et al. 2020). Particularly, an increasing number of NADC30-like isolates and various recombination events were reported in recent years, indicating that the NADC30-like PRRSV contributed to PRRS outbreaks by recombining with other strains (Zhou et al. 2017; Sun et al. 2018; Huang et al. 2019).
To further analyze the genomic characteristics of the variant PRRSV, HBap4-2018 was isolated from the PRRSV-positive samples in this study. The isolate could be propagated in MARC-145 cells, and the biological characteristics of HBap4-2018 in vitro were similar with that of HuN4, a HP-PRRSV that isolated and identified in our laboratory previously (Tong et al. 2007; Zhou et al. 2008). Further identification showed that HBap4-2018 could be recognized by HP-PRRSV HuN4 strain-specific GP5 and N protein monoclonal antibodies, but couldn't be recognized by the nsp2 monoclonal antibody, indicating that the mutations occurred in nsp2 gene between HBap4-2018 and HP-PRRSV. As the largest non-structural protein encoded by PRRSV, nsp2 is extremely easy to mutate, especially in the hypervariable region, the size of which is quite variable, reminding the existence of a nonessential region for replication in nsp2 (Xu et al. 2012; Yu et al. 2018). A recent study reported that 323–521 aa in the nsp2 hypervariable region of PRRSV JXwn06 was associated with viral cellular tropism to primary porcine alveolar macrophage cell (Song et al. 2019). Notably, according to the PRRSV nsp2 polymorphic classification system proposed by Yu et al. (Yu et al. 2020), nsp2 of HBap4-2018 displays additional specific continuous deletion of five amino acids in the region of 463–467 aa compared with the related region of NADC30-like strains. It displayed a specific "111 + 5 + 1 + 19" amino-acid-deletion pattern. Besides, many other amino acid substitutions were found in nsp2, but these deletions and substitutions did not affect the adaptation of HBap4-2018 to MARC-145 cells. It could be speculated that these mutations might be a self-protecting evolutionary strategy of PRRSV to regulate its adaptability to piglets. In this study, no unique characteristics such as amino acid insertions and deletions were found in the functional domains of GP5 in HBap4-2018 as compared with some representative PRRSV strains. Nsp9 and nsp10 contributed to the fatal virulence of HP-PRRSV emerging in China (Li et al. 2014; Zhao et al. 2018). But in this study we found that both nsp9 and nsp10 of HBap4-2018 showed high homology with those of HP-PRRSV, indicating that the pathogenicity of HBap4-2018 might be similar with HP-PRRSV, which had been confirmed by the viral challenge test.
Currently, the co-existence of multiple PRRSV lineages in swine herds not only increases the genetic diversity of PRRSV, but also provides suitable condition for recombination of PRRSV in the field. Recombination is an important evolutionary strategy for PRRSV. Both interlineage and intra-lineage recombination occurred, contributing to the emergence of new epidemic isolates (Guo et al. 2018, 2019; Zhou et al. 2018b). A recent study reported that during 2014–2018, intro-lineage recombination hot spots were scattered across the genome of both Chinese and US PRRSV strains, while the high-frequency inter-lineage recombination regions were both located in nsp9 and GP2 to GP3 (Yu et al. 2020). In this study, HBap4-2018 was confirmed to be an inter-lineage recombinant PRRSV, and five recombination break points were identified to locate in nsp2, nsp3, nsp5, nsp9 and ORF6, respectively. The whole genome of HBap4-2018 was separated into six regions of different length by these five recombination break points. It was indicated that HBap4-2018 was a variant PRRSV that recombined by HP-PRRSV (lineage 8) and NADC30-like PRRSV (lineage 1). The three short regions of nsp2–nsp3 (2000–5105 nt), nsp5–nsp9 (6292–7412 nt) and ORF6–30UTR (14, 249–15, 003 nt) were provided by NADC30-like PRRSV, while most of the HBap4-2018 genome was provided by the major parental isolates HP-PRRSV, which reserved the most of the regions that associated with virulence in the HP-PRRSV genome. This novel recombinant pattern was different from other patterns reported in the past (Yu et al. 2020). It could enrich the understanding of such recombination events. Nsp9 region encodes RNAdependent RNA polymerase, which is involved in the replication of PRRSV (Zhao et al. 2018). To explore whether the recombination break point located in nsp9 might affect its replication, multistep growth curve was constructed. Considering that HP-PRRSV instead of classical PRRSV served as the major parental strains in the genome of HBap4-2018, we compared the growth kinetics among HBap4-2018, HP-PRRSV HuN4, and HuN4-F112, an attenuated vaccine strain of HuN4 via serial passage in vitro. The results showed that the growth kinetics of HBap4-2018 was similar to that of HP-PRRSV HuN4 in the early stages of infection, but significantly lower than that of HuN4-F112, further indicating the recombinant HBap4-2018 might be derived from HP-PRRSV strains instead of their attenuated vaccine strains.
Vaccine immunization plays an important role in the prevention and control of PRRS. At present, various PRRS modified live virus vaccines are available in China, which can provide appreciable protection against genetically homologous PRRSV (Tian et al. 2009; Yu et al. 2015). However, ideal cross-protection cannot be provided by current commercial vaccines against heterologous PRRSV (Bai et al. 2016; Wang et al. 2016; Zhou et al. 2017). Moreover, NADC30-like PRRSV has been reported to recombine with field isolates and live-vaccine strains, which increases the difficulty of PRRS control (Lu et al. 2015; Dong et al. 2018). Evidences showed that recombination among multiple lineages were capable of making a difference on the virulence of PRRSV, which could not be predicted in the wild (Liu et al. 2018; Wang et al. 2018). Piglets infected with the highly pathogenic recombinant PRRSV 14LY01-FJ and SD17-38 showed sustained high fever, dyspnea, typical interstitial pneumonia and pulmonary lesions (Liu et al. 2017; Chen et al. 2018), while recombinant PRRSV TJnh1501 and CHsx1401 were considered to be the moderately virulent PRRSV because of the moderate clinical signs (Bian et al. 2017; Zhou et al. 2017). By contrast, FJ1402 and SCN17 infected piglets just showed transient fever and slight dyspnea (Zhang et al. 2016; Zhou et al. 2018b). To explore whether the existence of recombination events in HBap4-2018 affected its virulence, its pathogenicity was evaluated in 4-week-old piglets, and HuN4 was regarded as the HP-PRRSV. Piglets inoculated with HBap4-2018 displayed typical clinical signs such as sustained high fever, dyspnea and slow growth. The virus shedding of piglets infected with HBap4-2018 peaked at 5 dpi, and the viral load reached the highest level at 14 dpi, both of which delayed slightly than those of piglets infected with HuN4. Although two piglets in the HBap4-2018-inoculated group survived at the end of experiment, three piglets died at 6 dpi, 16 dpi and 28 dpi, respectively, with the mortality rate of 60%. It was indicated that HBap4-2018 was highly pathogenic to piglets. The above results suggested that nsp2–nsp3 (2000–5105 nt), nsp5–nsp9 (6292–7412 nt) and ORF6–30UTR (14, 249–15, 003 nt) regions in the PRRSV genome couldn't significantly affect the virulence of HP-PRRSV, and these regions were not critical for determining the virulence of HP-PRRSV.
In conclusion, HBap4-2018 was identified to be a novel natural recombinant PRRSV with three recombinant fragments in the genome, in which HP-PRRSV served as the major parental strains, while NADC30-like PRRSV served as the minor parental strains. HBap4-2018 reserved the most of the regions that associated with virulence in the HP-PRRSV genome, and was confirmed to be highly pathogenic to piglets. Although HP-PRRSV strains are still considered to be predominant in swine herds in China, they suffer extremely frequent recombination events with diverse recombinant patterns, which cause considerable difficulty in PRRS prevention. This study highlights the importance of continuous monitoring the prevalence status of PRRSV.
Virus Isolation and Identification
Genomic Characteristics and Phylogenetic Analysis of HBap4-2018
Recombination Analysis of HBap4-2018
Clinical Signs Observation
Virus Shedding, Viral Loads and Antibody Detection
Macroscopic and Histopathological Lesions in the Lungs
Discussion
-
The study was supported by the Shanghai Municipal Agriculture Science and Technology Project (2020-02-08-00-08-F01465), the National Natural Science Foundation of China (32072861), the Natural Science Foundation of Shanghai (20ZR1469600), and the earmarked fund for Modern Agro-industry Technology Research System of China (CARS-35).
-
YJZ and GZT conceived and designed the experiments. XMT, PFC, MQL, XW, STZ, and XWZ performed the experiments. PFC, XMT, JRY and JRZ analyzed the data. LXY, WT, and FG contributed reagents/materials/analysis tools. HY, CLL and YFJ participated in analysis and discussion. PFC and XMT wrote the paper. YJZ checked and finalized the manuscript. All authors read and approved the final manuscript.
-
The authors declare that they have no conflict of interest.
-
All animal experiments were approved by the Ethical Committee of the Shanghai Veterinary Research Institute, Chinese Academy of Agricultural Sciences (SHVRI-SZ-20191112-01).
















 DownLoad:
DownLoad: