-
Ascoviridae is an insect-specific family of viruses with double-stranded circular DNA genomes of 110–200 kb (Bigot et al., 2011; Wei et al., 2014). Ascoviruses can cause a chronic but ultimately fatal disease in the larvae of members of the Noctuidae, Crambidae, and Plutellidae families (Bigot et al., 2011). Based on virion morphology, DNA sequence information, host range, and tissue tropism, two genera, Ascovirus and Toursvirus, have been recognized by the International Committee on Taxonomy of Viruses (ICTV) (Asgari et al., 2017). The genus Ascovirus contains three species–Spodoptera frugiperda ascovirus 1a (SfAV-1a), Trichoplusia ni ascovirus 2a (TnAV-2a), and Heliothis virescens ascovirus 3a (HvAV-3a) (Bigot et al., 2011)–while the newly assigned genus Toursvirus consists of only one species, Diadromus pulchellus ascovirus 4a (DpAV-4a), that was recently removed from the genus Ascovirus (Asgari et al., 2017).
A total of 18 ascoviral isolates or strains have been reported globally so far (Bigot et al., 2011; Huang et al., 2012a). Among these, the genomes of only 6 isolates have been sequenced: SfAV-1a (Bideshi et al., 2006), TnAV-6a (previously named as TnAV-2c, Wei et al., 2014) (Wang et al., 2006), HvAV-3e (Asgari et al., 2007), HvAV-3f (Wei et al., 2014), HvAV-3g (Huang et al., 2012b), and DpAV-4a (Bigot et al., 2009). Among the sequenced isolates, HvAV-3g contains the largest genome (199, 721 bp), while DpAV-4a has the smallest one (119, 343 bp) (Bigot et al., 2009; Huang et al., 2012b).
Based on the analysis of biological characteristics, the species HvAV-3a, which consists of 8 isolates, has been found to be the most diverse and widely distributed species from America to Asia and Australia (Hamm et al., 1998; Huang et al., 2012a). Among HvAV-3a species, the genome sequences of 3 isolates–HvAV-3e, HvAV-3g, and HvAV-3f–have been reported from Australia, Indonesia, and USA, respectively (Asgari et al., 2007; Huang et al., 2012b; Wei et al., 2014). However, no ascovirus isolated from China has been sequenced so far. To understand the diversity and phylogeny of HvAV-3a isolates, we aimed to sequence the complete genome of a HvAV-3h, which was isolated from China and transmitted by Microplitis similis (Li et al., 2016). We then went on to compare the sequence with the other 6 previously published ascovirus genome sequences. The results of our study will be helpful for providing insights into the origin and evolution of ascoviruses.
-
For this study, HvAV-3h was propagated in Spodoptera exigua larvae as described previously (Huang et al., 2012a). The virions were then purified and viral DNA was extracted as previously described (Federici et al., 1990).
-
The genome was sequenced with the Roche 454 GS FLX system by using shotgun strategy. The reads were assembled with Roche GS De Novo assembler software. A few regions that were not assembled into contigs were further amplified using PCR and sequenced.
Hypothetical ORFs were predicted using ORF finder (NCBI) with characteristics of containing a standard ATG start codon and a stop codon and potentially encoding at least 50 amino acids (aa). Gene annotation and comparisons were performed using the NCBI protein-protein BLAST algorithm (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Homologies among ascovirus genomes were investigated using Basic Local Alignment Search Tool (BLAST) (blastp) in NCBI (Altschul et al., 1997) and two sequences were aligned using blastp. MIROPEATS program (Parsons, 1995) was used to find repetitive regions. Restriction sites were predicted and the genome map framework was drawn using the program Geneious vesion 8.04. The annotated genome sequence data were uploaded to GenBank under the accession number KU170628.
-
Phylogenetic analysis of the ascoviruses was performed using the amino acid sequences of DNA polymerase and Major capsid protein (MCP) from HvAV-3h and the 6 other sequenced Ascovirus and Toursvirus isolates, with homologs from invertebrate iridescent virus 22 and Wiseana iridescent virus as the outgroups (Piégu et al., 2015). The sequences were then aligned using ClustalW (Larkin et al., 2007) with default parameters of MEGA 6.0 (Tamura et al., 2013). The maximum likelihood (ML) method was used with the program raxmlGUL 1.5 (https://sourceforge.net/projects/raxmlgui/) (Silvestro and Michalak, 2012), in which the analysis setting was chosen “ML+rapid bootstrap” and the “number of bootstrap replicates” was 1, 000. The program MrBayes 3.1.2 (http://morphbank.Ebc.uu.SE/mrbayes/) was used to perform Bayesian inference (BI) analysis (Ronquist et al., 2003), with the MCMC analysis run for 300,000 generations and a burn-in series of 1,000.
-
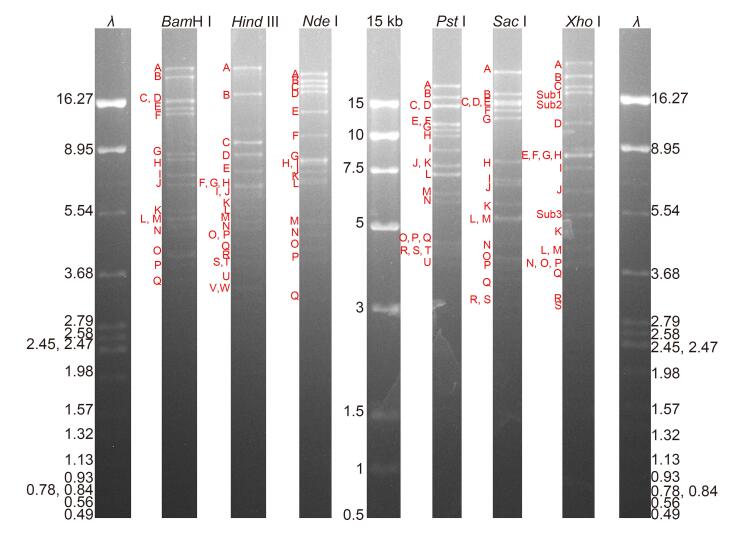
The HvAV-3h genome was sequenced using the Roche 454 GS FLX system with the shotgun strategy. A total of 43, 218 reads were obtained with the average length of 423 bp, and the genome was covered about 53 times. The HvAV-3h genome was assembled using Roche GS De Novo assembler software and Geneious version 8.04. A few regions that were not assembled into contigs were further amplified using PCR with the primers listed in Supplementary Table S1, cloned, and sequenced. The final assembled contig representing the entire HvAV-3h genome sequence was confirmed using restriction digestion with 6 enzymes (BamH I, Hind III, Nde I, Pst I, Sac I, and Xho I). The restriction profile (Supplementary Figure S1) matched well with the predicted fragments of the assembled genome (Supplementary Table S2). However, there were still some submolar bands (Supplementary Figure S1), indicating the presence of different genotypes in the isolate.
The HvAV-3h genome was assembled into a circular contiguous sequence of 190, 519 bp, which was significantly larger than the genome size (165 kbp) estimated previously (Huang et al., 2012a). This was possibly largely due to the missing small size fragments in the previously reported restriction enzyme digestion profiles (Huang et al., 2012a). A genomic map showing the organization of the ORFs with the predicted protein of over 50 aa in the HvAV-3h genome is presented in Figure 1. The genome has a G+C content of 45.5% and encodes 185 predicted ORFs, in which 104 are in the forward orientation and 81 are in the reverse orientation (Figure 1). The coding region accounts for 88.8% of the total sequences.
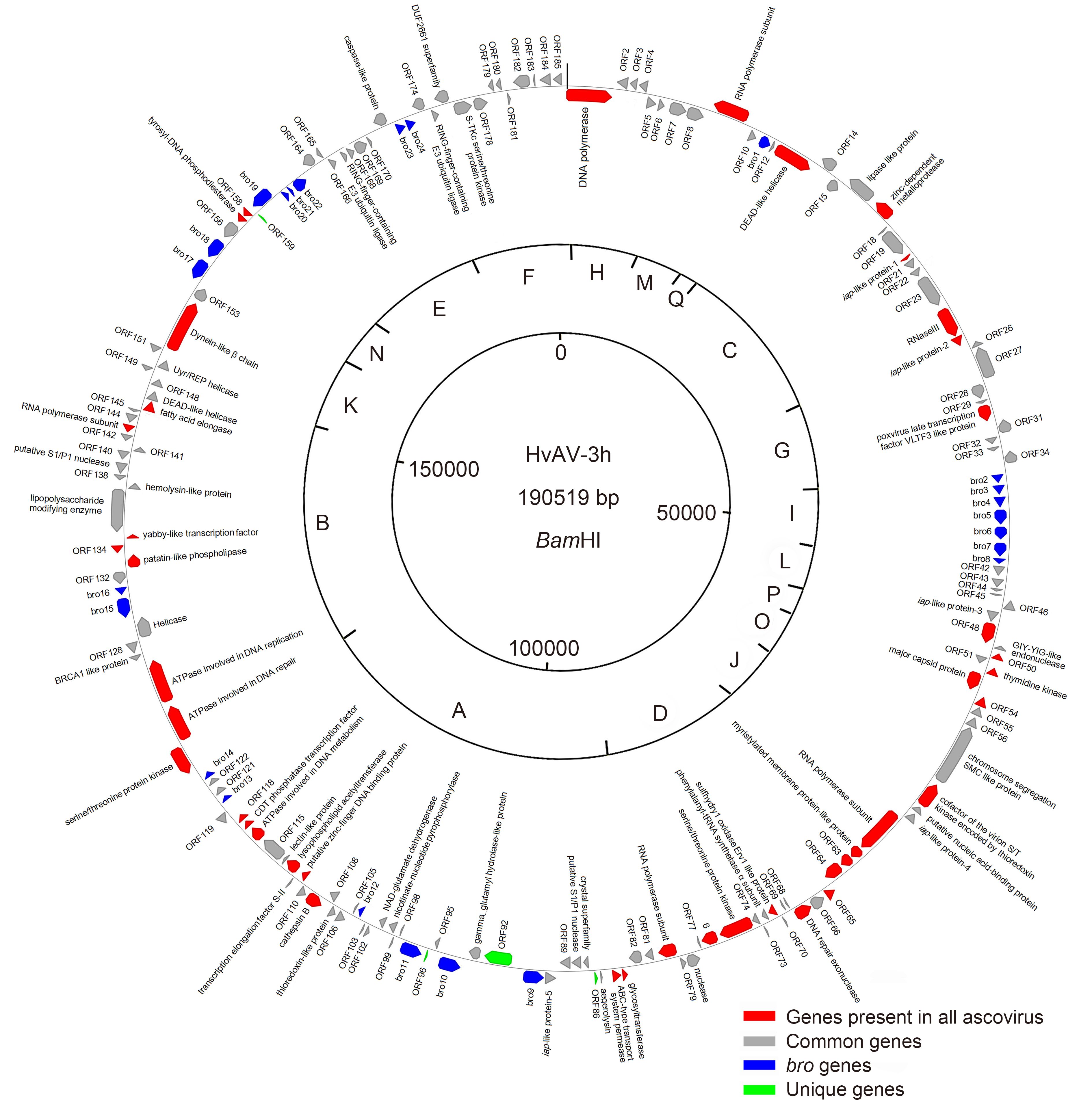
Figure 1. Circular map and gene organization of the HvAV-3h genome. The sites for the restriction enzyme BamH I are presented; the fragments are indicated as A to Q according to the size from the largest to the smallest restriction fragment in Supplementary Figure S1. (Red color indicates core genes present in all ascovirus isolates, grey color indicates common genes, blue color indicates bro genes, and green color indicates unique genes).
-
The size of the HvAV-3h genome was found to be smaller than that of the HvAV-3g (199, 721 bp) and HvAV-3f genomes (198, 157 bp), but larger than that of the HvAV-3e genome (186, 262 bp). It is thus the third largest ascovirus genome described so far. Gene-parity plots of HvAV-3h against the 6 ascovirus isolates revealed high co-linearity of the gene order between HvAV-3h and HvAV-3g, HvAV-3f, or HvAV-3e, with only a few inversions and drifts, which may partially account for the difference in the genome size. Much lower co-linearity was found between HvAV-3h and SfAV-1a or TnAV-6a. No obvious collinear region could be found between HvAV-3h and DpAV-4a (Figure 2). The collinear regions between HvAV-3h and other ascovirus isolates, with the exception of DpAV-4a, indicated the presence of conserved regions among ascoviral genomes and their derivation from an ancestor ascovirus.
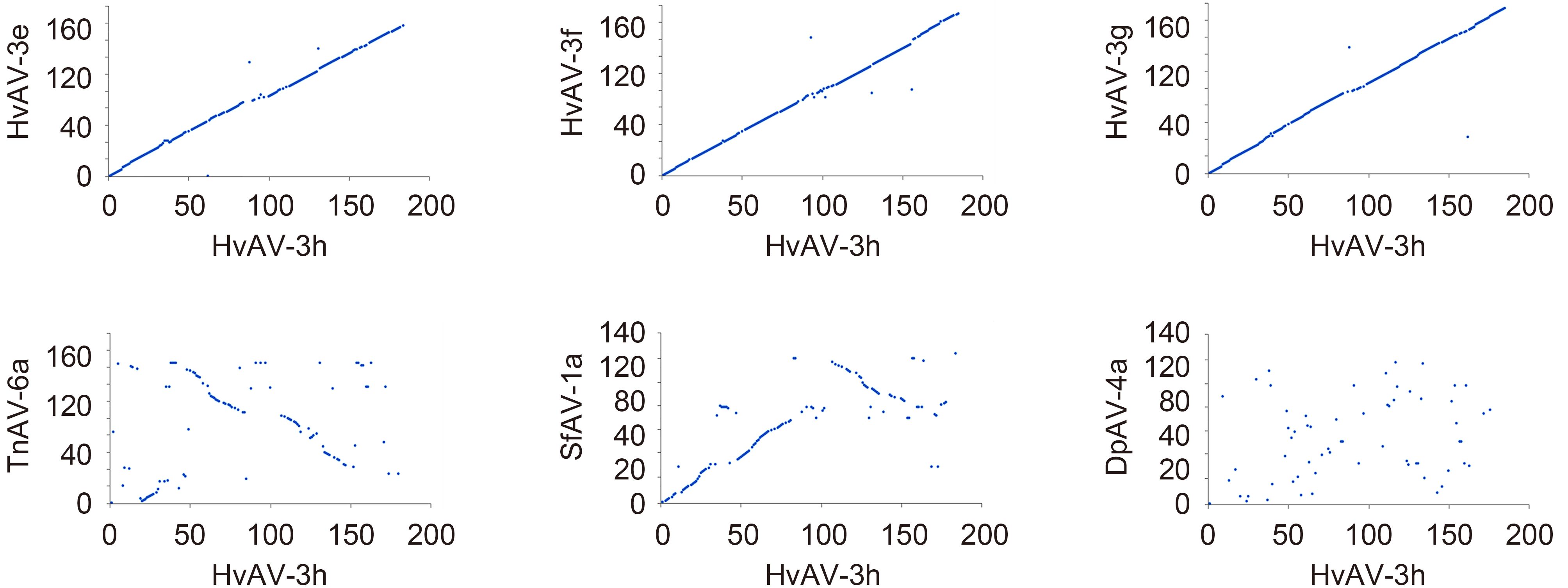
Figure 2. Gene Parity Plot analyses of HvAV-3h with the 6 other sequenced Ascovirus and Toursvirus isolates. Homologous genes were plotted based on their relative locations in the genomes. The x-axis represents ORFs from HvAV-3h and y-axis represents ORFs from each isolate indicated in the title of each panel.
Phylogenetic analyses of the conserved genes of DNA polymerase and MCP from HvAV-3h and the 6 other Ascovirus and Toursvirus isolates also revealed a close relationship between HvAV-3h, HvAV-3g, HvAV-3f, and HvAV-3e (Figure 3). In fact, analysis using ML and Bayesian methods with a bootstrap value of 100% and a posterior probability of 1.00, respectively, revealed that these four HvAV-3a isolates form a monophyletic clade at high levels. The HvAV-3a isolates also appeared to have a distant relationship with SfAV-1a and TnAV-6a and an even more distant relationship with DpAV-4a (Figure 3).
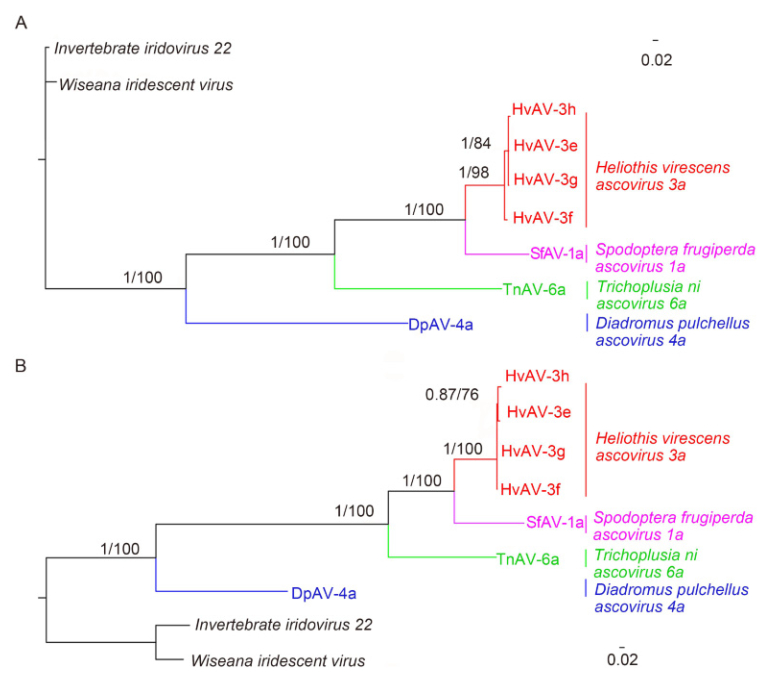
Figure 3. Phylogenetic tree based on amino acid sequences of MCP (A) and DNA polymerase (B) from the genomes of HvAV-3h and the 6 other ascovirus isolates, which were determined using Bayesian and Maximum likelihood (ML) analysis. The scale bar indicates the number of substitutions per spot. Numbers represent the ncxJe supports; the number on top indicate posterior probabilities for the Bayesian Inference and those below are based on 1000 bootstrap replicates for ML. The colors represent the haplogroups specific to each species.
-
BLAST analysis revealed that, among the identified ORFs, 181 ORFs were related to genes reported from the other 6 ascovirus isolates, including HvAV-3g (174 ORFs, with average amino acid identity of 94.0%), HvAV-3f (172 ORFs, 92.9%), HvAV-3e (162 ORFs, 87.5%), SfAV-1a (116 ORFs, 62.7%), TnAV-6a (105 ORFs, 56.7%), and DpAV-4a (63 ORFs, 34.0%) (Supplementary Table S3).
Forty-four ascovirus ORFs were found to be conserved among HvAV-3h and the other 6 ascovirus and toursvirus genomes. Gene annotation indicated that these genes are involved in DNA/RNA replication/transcription/metabolism, viral packaging and assembly, sugar and lipid metabolisms, etc. (Table 1). All the ascovirus isolates except DpAV-4a shared 25 ORFs with HvAV-3h (Table 2). Some of them appear to play roles in DNA/RNA replication/transcription/metabolism, but the functions of most of the shared genes are still unknown (Table 2). A total of 159 genes including the above-mentioned genes were found to be shared between HvAV-3h and the three closely related isolates of the species HvAV-3a. In addition, 19 genes were found to be shared between HvAV-3h and any one but not all three previously reported isolates of the species HvAV-3a. Six ORFs including ORF49 (GIY-YIG-like endonuclease), ORF51, ORF86, ORF92, ORF96, and ORF159 were not found in any of the other three isolates. ORF49 showed significant similarity to GIY-YIG-like endonuclease with E-value of 6e-46. GIY-YIG-like endonuclease was found in SfAV-1a, TnAV-6a, and DpAV-4 as well as in viruses of the Baculoviridae and Iridoviridae families. Homologues of GIY-YIG-like endonuclease are known to be involved in viral DNA repair (Lindahl, 1982) and are required for efficient baculoviral virion production (Tang et al., 2013; Wu & Passarelli, 2012). ORF51 with an unknown function only showed significant similarity to ORF39 of SfAV-1a with E-value of 1e-44. The other four ORFs, including ORF86, ORF92, ORF96, and ORF159, showed no significant levels of similarity to genes in the GenBank database; hence, they were considered unique genes of HvAV-3h.
Classification Genes DNA/RNA/replication/transcription/ metabolism (16) DNA polymerase (orf1), RNA polymerase subunit (orf9), DEAD-like helicase (orf13), RNase III (orf24), poxvirus late transcription factor VLTF3 like protein (orf30), thymidine kinase (orf52), RNA polymerase subunit (orf61), DNA repair exonuclease (orf67), RNA polymerase subunit (orf80), ATPase involved in DNA metabolism (orf116), CDT phosphatase transcription factor (orf117), ATPase involved in DNA repair (orf125), ATPase involved in DNA replication (orf126), yabby-like transcription factor (orf135), RNA polymerase subunit (orf143), tyrosyl-DNA phosphodiesterase (orf157) Packaging and assembly (2) major capsid protein (orf53), sulfhydry1 oxidase Erv1 like protein (orf71) Sugar and lipid metabolism (4) Glycosyl transferase (orf83), lysophospholipid acetyl transferase (orf113), patatin-like phospholipase (orf133), fatty acid elongase (orf146) Others (12) zinc-dependent metalloprotease (orf17), iap-like protein-1 (orf20), iap-like protein-2 (orf25), cofactor of the virion S/T kinase encoded by thioredoxin (orf58), myristylated membrane protein-like protein (orf62), serine/threonine protein kinase (orf75), ABC-type transport system permease (orf84), cathepsinB (orf109), putative zinc-finger DNA binding protein (orf111), serine/threonine protein kinase (orf124), dynein-like β chain (orf152), bro* Unknown (10) orf48, orf50, orf54, orf63, orf64, orf65, orf76, orf118, orf134, orf158 Note: * HvAV-3h encodes 24bros:orf11, orf35, orf36, orf37, orf38, orf39, orf40, orf41, orf91, orf94, orf97, orf104, orf120, orf123, orf130, orf131, orf154, orf155, orf160, orf161, orf162, orf163, orf172, and orf173. Table 1. Classification of the 44 genes found in all Ascoviridae isolates
Classification Genes DNA/RNA/replication/ transcription/ metabolism (4) nuclease (orf78), putative S1/P1 nuclease (orf88), helicase (orf129), putative S1/P1 nuclease (orf139) Others (5) iap-like protein-3 (orf47), thioredoxin-like protein (orf107), BRCA1 like protein (orf127), lipopolysaccharide-modifying enzyme (orf136), caspase-like protein (orf171) Unknown (16) orf5, orf8, orf14, orf19, orf21, orf22, orf23, orf26, orf27, orf31, orf34, orf43, orf55, orf66, orf74, orf81 Table 2. Classification of the 25 genes found in all sequenced isolates of the genus Ascovirus
-
The baculovirus repeat ORF (bro) gene occurs as multiple copies per ascovirus genome (Asgari et al., 2007). For example, the genomes of SfAV-1a, TnAV-6a, HvAV-3e, HvAV-3f, HvAV-3g, and DpAV-4a show 7, 3, 23, 29, 25 and 12bro genes, respectively (Asgari et al., 2007; Bideshi et al., 2006; Bigot et al., 2009; Huang et al., 2012b; Wang et al., 2006; Wei et al., 2014). In our study, we found that HvAV-3h contains 24bro genes that encode from 99 to 521 residues (Figure 1, Supplementary Table S3). The bro genes in baculovirus have been believed to help the virus acquire a new functionality eventually (de Castro Oliveira et al., 2013), but their function in ascoviruses remains to be determined.
-
HvAV-3h is the fourth sequenced isolate of the species HvAV-3a. HvAV-3h had a similar gene order and shared 159 ORFs with all the other isolates of the species HvAV-3a (Figure 2, Supplementary Table S3). However, there are still some gene reversions and drifts among the genomes. For example, 19 ORFs were found in any one but not all the four isolates, and 6 ORFs were only found in HvAV-3h (Figure 2, Supplementary Table S3). These findings show that although HvAV-3a isolates are closely related to each other, there are noticeable diversities in their genome organizations.
Comprehensive Geneparity Plots and phylogeny analyses of all the sequenced Ascoviridae isolates indicated that HvAV-3a isolates are distantly related to SfAV-1a and TnAV-6a and most distantly to DpAV-4a (Figure 2, Figure 3). All the Ascoviridae isolates shared 44 conserved genes (Table 1), which likely play important roles during the life cycle of the viruses. Among the 44 conserved ORFs, 16–DNA polymerase (orf1), RNA polymerase subunits (orf9, orf61, orf80, and orf143), DEAD-like helicase (orf13), RNase III (orf24), poxvirus late transcription factor VLTF3 like protein (orf30), thymidine kinase (orf52), DNA repair exonuclease (orf67), ATPase involved in DNA metabolism (orf116), CDT phosphatase transcription factor (orf117), ATPase involved in DNA repair (orf125), ATPase involved in DNA replication (orf126), yabby-like transcription factor (orf135), and tyrosyl-DNA phosphodiesterase (orf157)–are predicted to be involved in DNA/RNA replication/transcription and metabolism (Table 1). All the ascoviruses appear to encode many genes involved in nucleotide metabolism and transcription/replication. For example, TnAV-6a encodes 16 related ORFs (Wang et al., 2006) and SfAV-1a encodes 9 (Bideshi et al., 2006).Serine/threonine protein kinase and AV-like serine/ threonine protein kinase are also conserved in ascoviruses. These kinases likely participate in virus or virus-host regulation of signaling networks (Jacob et al., 2011); however, the roles of the kinases in ascovirus infection are not clear so far. Efforts also need to be made to reveal the functions of other unknown conserved ORFs (Table 1). Among the 44 core genes, only 2 were annotated to encode the structural proteins MCP (orf53) and sulfhydry1 oxidase Erv1-like protein (orf71) (Long et al., 2009; Wu and Passarelli, 2010). As ascoviruses have complicated morphologies, it is tempting to assume that more structural genes may be identified from the remaining unknown core genes.
Ascovirus genomes often contain large interspersed repeats of 1–3 kbp (Bigot et al., 2000), but no repeat regions (hrs) were found in the HvAV-3h genome (Figure 1). Similarly, no repeat regions have been found in the DpAV-4a genome (Bigot et al., 2009). In contrast, 5 copies of repeats have been found in the HvAV-3e genome (orf38 <hr1>, orf84 <hr2>, orf124 <hr3>, orf156 <hr4> and orf178 <hr5>) (Asgari et al., 2007) and the HvAV-3g genome (orf35, orf68, orf104, orf126, and orf173) (Huang et al., 2012b). Two repeat regions have been found in the HvAV-3f genome (orf95 and orf179) (Wei et al., 2014) and the SfAV-1a (orf34 and orf77) (Bideshi et al., 2006) and TnAV-6a (Hr1 and Hr2) genomes (Wang et al., 2006). Additionally, 24bros were found in the HvAV-3h genome in our study, and 12, 7, 3, 23, 26 and 29bros have been found previously in DpAV-4a (Bigot et al., 2009), SfAV-1a (Bideshi et al., 2006), TnAV-6a (Wang et al., 2006), HvAV-3e (Asgari et al., 2007), HvAV-3g (Huang et al., 2012b), and HvAV-3f (Wei et al., 2014), respectively. The function of repeat regions, bros, and hrs in the evolution and variety of ascovirus genomes needs to be investigated in future studies.
In summary, we sequenced the genome of a HvAV-3h isolate from China and compared the sequence to those of 6 other previously published ascovirus genome sequences to establish the evolutionary relationship between different ascovirus species. Our findings indicate a close relationship between different HvAV-3a isolates and distant relationship between HvAV-3h and some other species of the Ascoviridae family. Future studies need to be focused on elucidating the diversities in the genome organizations of different HvAV-3h isolates and on understanding the role of different genes in the ascoviral genomes as well as the conservation of these genes.
-
The authors would like to thank Jue Hu, Xing-Shi Gu, Wen-Fei Xian, Hai-Zhou Liu, Lei Zhang, and Shun-Ji Li for their help in virus amplification and data analysis. The authors would also like to acknowledge the technical assistance received from the Core Facility and Technical Support of Wuhan Institute of Virology. This study was supported partly by the National Natural Science Foundation of China (No. 31371995 and 31621061), the Strategic Priority Research Program of the Chinese Academy of Sciences (grant XDB11030400), STS Project of the Chinese Academy of Sciences (grant KFJ-SW-STS-143-3), and the Hunan Provincial Natural Science Foundation for Distinguished Young Scholar of China (14JJ1023).
-
The authors declare that they have no conflict of interest. This article does not contain any studies with human or animal subjects performed by any of the authors.
-
Conceived and designed the experiments: HGH, DHH, MW, XWC, ZH. Performed the experiments: HGH, DHH. Analyzed the data: HGH, DHH, ZH. Contributed reagents/materials/analysis tools: HGH, DHH. Wrote the paper: HGH, DHH, ZH.
Supplementary figure/tables are available on the websites of Virologica Sinica: www.virosin.org;link.springer.com/journal/12250.
-
Primer name Sequences (5′ to 3′) HvAV-3h_1 forward GGGCCCAGTTTCCCGCACAA reverse ACCGAGTCAGTGAACGCGCC HvAV-3h_2 forward TCAACCGCGCCGGACTGTTC reverse ACGCCGTCGGTGTTGACGAC HvAV-3h_11 forward CGCAACGTTATCGCCGCACG reverse GACATCGTCGGCGCTCTCGG HvAV-3h_12 forward ACATGGACCCCGATGCAATTC reverse CCGGTGGTGTCATTCAAGGA HvAV-3h_13 forward CTCCCTCTAGAAACGGTTAGTG reverse ACCACCGTATGACACATTGC Table S1. Primers for polymerase chain reaction
BamH I Hind III Nde I Pst I Sac I Xho I A 35,071 33,369 25,756 19,178 27,735 37,139 B 25,390 16,998 21,689 15,785 16,831 23,423 C 15,809 8,440 19,446 13,999 15,089 16,842 D 15,429 8,396 17,508 13,962 14,768 11,159 E 13,896 7,345 12,964 11,199 14,400 8,362 F 12,982 6,610 9,950 11,019 13,311 8,351 G 8,511 6,577 8,118 10,411 12,051 8,344 H 8,137 6,503 7,798 9,864 7,972 8,257 I 7,143 6,292 7,703 8,643 6,923 7,546 J 6,657 6,111 7,240 7,597 6,718 6,303 K 5,497 5,622 6,836 7,589 5,828 4,833 L 5,041 5,363 6,684 7,060 5,335 4,367 M 4,948 5,150 5,073 6,185 5,223 4,306 N 4,720 5,053 4,650 5,866 4,509 4,071 O 4,261 4,894 4,240 4,457 3,998 3,984 P 3,938 4,817 3,983 4,426 3,836 3,945 Q 3,572 4,441 3,154 4,325 3,588 3,887 R 2,437 4,253 2,613 4,261 3,072 3,170 S 2,410 3,981 2,232 4,179 3,063 2,980 T 1,855 3,888 2,213 4,121 2,826 2,554 U 1,133 3,679 1,846 3,924 2,559 2,500 V 1,109 3,412 1,715 1,859 2,218 2,474 W 387 3,400 1,102 1,841 2,100 2,326 X 186 3,385 1,075 1,817 1,805 1,721 Y 3,132 1,051 1,716 1,369 1,597 Z 2,649 891 1,666 1,081 1,495 AA 2,598 834 1,185 821 1,431 BB 2401 799 936 727 973 CC 2050 635 877 615 582 DD 1820 452 379 148 473 EE 1682 269 121 452 FF 1505 72 317 GG 1419 237 HH 1174 97 II 747 11 JJ 546 10 KK 397 LL 231 MM 189 Total 190,519 190,519 190,519 190,519 190,519 190,519 Table S2. Restriction enzyme digestion results of the HvAV-3h genome
HvAV–3h
ORFsPosition aa# Product Description BLAST best match orthologous orfs number (Id%$) HvAV–3e HvAV–3f HvAV–3g SfAV–1a TnAV–6a DpAV–4a 1* 1 > 3261 1086 DNA polymerase 1(98) 1(98) 1(98) 1(66) 1(44) 1(30) 2 3456 < 4235 259 hypothetical protein 2(97) 2(95) 2(97) – 84(23) – 3 4232 < 4846 204 hypothetical protein 3(97) 3(71) 3(96) 2(30) – – 4 4892 < 5602 236 hypothetical protein 4(95) 4(84) 4(95) 3(38) – – 5 5636 > 6409 257 hypothetical protein 5(99) 5(99) 5(99) 4(72) 164(40) – 6 6466 > 7029 187 hypothetical protein 6(95) 6(94) 6(95) – – – 7 7310 > 8545 411 hypothetical protein 7(95) 7(92) 7(94) 5(36) – – 8 8548 > 9855 435 hypothetical protein 8(96) 8(94) 8(96) 7(39) 21(23) – 9* 10136 < 12715 859 RNA polymerase subunit 11(99) 10(99) 11(99) 8(79) 42(61) 89(35) 10 13173 > 13835 220 hypothetical protein 12(94) 11(91) 12(93) – – – 11 14052 > 14927 291 bro1 13(96) 12(96) 13(96) 30(39) – – 12 14930 > 15226 98 hypothetical protein 14(99) 13(98) 14(99) – 41(40) – 13* 15291 > 18125 944 DEAD–like helicase 15(98) 14(98) 15(97) 9(65) 161(43) 20(36) 14 18359 < 19489 376 hypothetical protein 17(95) 15(95) 17(96) 11(37) 160(28) – 15 19683 > 20576 297 hypothetical protein 18(97) 16(71) 18(97) 12(31) – – 16 20699 < 22666 655 lipase like protein 19(97) 17(95) 19(97) 13(48) – – 17* 23086 < 24510 474 zinc–dependent metalloprotease 20(96) 19(94) 20(96) 14(59) 158(29) 29(30) 18 24739 > 24900 53 hypothetical protein 21(93) – 21(92) – – – 19 24976 > 26916 646 hypothetical protein 22(96) 20(88) 22(96) 15(47) 6(31) – 20* 27150 > 27458 102 iap–like protein–1 23(98) 21(95) 23(98) 16(44) 3(28) 7(33) 21 27539 > 28225 228 hypothetical protein 24(97) 22(98) 24(97) 17(61) 4(30) – 22 28194 > 28784 196 hypothetical protein 25(98) 23(97) 25(98) 18(59) 5(38) – 23 29013 > 31337 774 hypothetical protein 26(97) 24(95) 26(97) 20(51) 7(24) – 24* 31648 > 33834 728 RNaseIII 27(98) 25(95) 27(98) 22(55) 8(41) 3(35) 25* 33838 > 34578 246 iap–like protein–2 28(99) 26(91) 28(99) 25(40) 9(29) 7(31) 26 34644 < 35048 134 hypothetical protein 29(99) 27(95) 29(99) 26(48) 10(36) – 27 35064 < 37343 759 hypothetical protein 30(98) 28(94) 30(98) 27(44) 11(30) – 28 37443 > 38507 354 hypothetical protein 31(99) 29(97) 31(98) 28(36) – – 29 38550 > 38846 98 hypothetical protein 32(99) 30(98) 32(99) – 13(40) – 30* 38941 > 40116 391 poxvirus late transcription factor VLTF3 like protein 33(97) 31(97) 33(97) 29(70) 17(53) 103(25) 31 40201 < 41181 326 hypothetical protein 34(96) 32(91) 34(92) 32(33) 26(34) – 32 41257 > 41814 185 hypothetical protein 36(93) 33(92) 36(96) – – – 33 41899 > 42321 140 hypothetical protein 37(96) 34(98) 37(96) – – – 34 42403 < 43311 302 hypothetical protein 39(91) 35(90) 38(92) 32(34) 26(38) – 35 43846 > 44565 239 bro2 42(29) 36(94) 40(95) 72(26) 137(26) – 36 44635 > 45399 254 bro3 42(33) 37(88) 42(93) – 27(27) – 37 45418 > 46284 288 bro4 42(78) 38(68) 43(77) 80(54) 137(39) 4(23) 38 46341 > 47420 359 bro5 40(78) 41(81) 44(74) 79(62) 165(65) 110(24) 39 47510 > 48529 339 bro6 41(86) 40(73) 47(97) 79(64) 165(70) 98(22) 40 48619 > 49665 348 bro7 43(88) 41(79) 44(86) 79(63) 165(62) 17(24) 41 49782 > 50243 153 bro8 44(93) 42(84) 48(94) 79(34) 165(37) – 42 50305 > 51009 234 hypothetical protein 45(94) 43(92) 49(96) 78(53) – – 43 51179 > 51880 233 hypothetical protein 46(98) 44(93) 50(98) 33(54) 18(27) – 44 51889 > 52194 101 hypothetical protein 47(96) 45(98) 51(97) – – – 45 52227 > 52469 80 hypothetical protein 48(98) 46(98) 52(98) – – – 46 52522 < 53295 257 hypothetical protein 49(96) 47(96) 53(95) – 34(32) – 47 53416 > 54231 271 iap–like protein–3 51(96) 49(95) 55(96) 74(42) 32(44) – 48* 54330 > 55790 486 hypothetical protein 52(96) 50(91) 56(96) 36(50) 157(29) 40(19)% 49 55699 < 56031 110 GIY–YIG–like endonuclease – – – 37(65) 87(46) 77(49) 50* 56102 < 56671 189 hypothetical protein 53(97) 52(83) 58(97) 38(56) 156(70) 63(33) 51 56685 > 57308 207 hypothetical protein – – – 39(41) – – 52* 57207 < 57860 217 thymidine kinase 55(98) 54(96) 60(98) 40(55) 154(33) 55(29) 53* 57905 > 59272 455 major capsid protein 56(99) 55(97) 61(99) 41(68) 153(34) 19(23) 54* 59343 < 60077 244 hypothetical protein 57(92) 56(94) 62(92) 42(46) 150(37) 60(35)% 55 60091 < 60768 225 hypothetical protein 58(99) 57(99) 63(99) 43(73) 149(32 – 56 60809 < 61537 242 hypothetical protein 59(94) 58(94) 64(95) – 148(43) 23(28) 57 61572 < 65825 1417 chromosome segregation
SMC like protein60(95) 59(94) 65(96) 46(35) – – 58* 66015 < 67856 613 cofactor of the virion S/T kinase encoded by thioredoxin 61(96) 60(94) 66(95) 48(68) 141(31) 8(28) 59 67889 < 68407 172 putative nucleic acid–
binding protein62(97) 61(97) 67(98) 49(45) – – 60 68502 < 69194 230 iap–like protein–4 63(97) 62(92) 69(98) 51(52) – – 61* 69296 > 72763 1155 RNA polymerase subunit 64(99) 63(98) 70(99) 52(81) 138(58) 73(41) 62* 72811 > 73689 292 myristylated membrane protein–like protein 65(26)% 64(98) 71(98) 54(60) 129(34) 65(28) 63* 73703 > 74608 301 hypothetical protein 66(98) 65(97) 72(97) 55(67) 126(40) 35(28) 64* 74724 > 76007 427 hypothetical protein 68(96) 66(95) 74(96) 56(44) 125(42) 64(32) 65* 75981 < 76742 253 hypothetical protein 69(96) 67(97) 75(96) 57(54) 124(40) 9(22)% 66 76820 < 77932 370 hypothetical protein 70(97) 68(97) 76(98) 58(47) 122(34) – 67* 77933 < 79132 399 DNA repair exonuclease 71(98) 69(98) 77(99) 59(61) 121(50) 26(37) 68 79169 > 79429 86 hypothetical protein – 70(95) 78(99) – 120(39) – 69 79437 > 79673 78 hypothetical protein 72(97) 71(90) 79(97) 60(63) – – 70 79816 < 80070 84 hypothetical protein 73(99) 72(100) 80(100) – – 71* 80142 > 80768 208 sulfhydry1 oxidase Erv1
like protein74(96) 73(95) 81(96) 61(49) 118(33) 41(30) 72 80785 > 81252 155 phenylalanyl–tRNA
synthetase α subunit75(96) 74(94) 82(96) – 117(31) – 73 81249 < 81482 77 hypothetical protein – – 83(94) 62(76) – – 74 81465 > 81989 174 hypothetical protein 76(95) 75(98) 84(95) 63(46) 116(36) – 75* 82035 > 84473 812 serine/threonine protein kinase 77(98) 76(98) 85(98) 64(53) 115(27) 46(24) 76* 84583 > 85797 404 hypothetical protein 78(94) 77(96) 86(94) 65(49) 113(33) 43(25) 77 85852 > 86103 83 hypothetical protein 79(99) 78(96) 87(99) – – – 78 86157 < 87149 330 nuclease 80(97) 79(96) 88(97) 66(54) 112(38) – 79 87189 < 87566 125 hypothetical protein 81(97) 80(98) 89(96) – – 80* 87588 > 88937 449 RNA polymerase subunit 82(98) 81(97) 90(98) 67(61) 110(52) 70(30) 81 89208 > 89942 244 hypothetical protein 83(99) 82(98) 91(99) 68(43) 159(24) – 82 90087 > 91070 327 hypothetical protein 85(98) 83(95) 92(99) – – – 83* 91150 < 91608 152 glycosyltransferase 86(100) 84(95) 93(100) 119(35) 107(45) 52(34) 84* 91614 < 92309 231 ABC–type transport system permease 87(94) 85(92) 94(94) 119(44) 107(44) 52(25) 85 92863 < 93132 89 aegerolysin – 87(83) – – 29(69) – 86 93184 < 93510 108 hypothetical protein – – – – – – 87 93765 > 94262 165 crystal superfamily – – 96(94) – – 88 94285 > 95067 260 putative S1/P1 nuclease 134(37) 89(91) 148(37) 75(41) 135(37) – 89 95073 > 95849 258 hypothetical protein – 91(73) – – – – 90 96020 < 96838 272 iap–like protein–5 89(85) 93(56) 97(74) – – – 91 96847 < 98334 495 bro9 90(76) 94(84) 98(94) 79(46) 165(46) 98(29) 92 99124 > 101097 657 hypothetical protein – – – – – – 93 101371 > 102309 312 gamma_glutamyl hydrolase–like protein – 162(52) 99(95) – – – 94 102460 < 104010 516 bro10 92(86) 96(81) 100(99) 79(48) 165(41) 34(33) 95 104416 > 104799 127 hypothetical protein 96(52) 92(92) 101(100) 78(29) – – 96 104926 < 105156 76 hypothetical protein – – – – – – 97 105319 < 106827 502 bro11 93(98) 97(81) 102(97) 70(86) 165(29) 75(29) 98 107101 > 107325 74 hypothetical protein – 98(93) – – – – 99 107359 < 107625 88 hypothetical protein – 100(91) 105(98) – – – 100 107626 > 108231 201 nicotinate–nucleotide pyrophosphorylase 94(90) 99(55) 106(93) – 136(55) – 101 108420 > 108965 181 NAD–glutamate dehydrogenase 95(99) 102(77) 107(99) 76(28) – – 102 109065 < 109457 130 hypothetical protein 96(99) 92(51) 108(99) 78(26) – – 103 109528 < 109740 70 hypothetical protein 97(96) 103(96) 109(96) – – – 104 110148 > 110618 156 bro12 98(98) 104(94) 110(97) – – – 105 110656 > 110901 81 hypothetical protein 99(94) – 111(95) – – – 106 110942 < 111634 230 hypothetical protein 101(97) 105(90) 112(97) – – – 107 111721 < 112212 163 thioredoxin–like protein 102(99) 106(99) 113(99) 116(51) 103(34) – 108 112251 > 112895 214 hypothetical protein – – 114(96) – – – 109* 112798 < 114033 411 cathepsin B 103(95) 107(92) 115(95) 114(52) 102(38) 48(31) 110 114056 < 114697 213 hypothetical protein – 108(88) 116(98) – – – 111* 114753 > 115304 183 putative zinc–finger DNA binding protein 105(97) 109(94) 117(97) 113(42) 100(36) 108(33) 112 115368 < 115601 77 transcription elongation
factor S–II– 110(100) 118(100) – 99(59) 82(44) 113* 115612 > 116649 345 lysophospholipid acetyltransferase 106(97) 111(95) 119(97) 112(48) 98(41) 81(26) 114 116650 > 116895 81 lectin–like protein 107(99) 112(85) 120(99) 115 117019 > 118821 600 hypothetical protein 108(97) 113(94) 121(97) – 96(34) – 116* 118944 > 120014 356 ATPase involved in DNA metabolism 109(98) 114(97) 122(98) 110(73) 95(61) 86(47) 117* 120111 > 120680 189 CDT phosphatase
transcription factor110(99) 115(98) 123(99) 109(69) 93(52) 117(34) 118* 120730 > 121236 168 hypothetical protein 111(98) 116(99) 124(98) 108(63) 91(41) 97(29) 119 121334 < 122080 248 hypothetical protein 112(97) 117(96) 125(97) – 84(38) – 120 122496 > 122975 159 bro13 113(93) 118(94) 127(93) – – – 121 123169 > 123732 187 hypothetical protein 114(98) 119(92) 128(98) – – – 122 124002 > 124427 141 hypothetical protein 115(99) 120(91) 129(99) 107(42) – 123 124507 > 125022 171 bro14 116(99) 121(90) 130(98) – – 124* 125522 < 127492 656 serine/threonine protein kinase 117(98) 122(98) 131(98) 104(63) 88(42) 36(30) 125* 127637 > 130294 885 ATPase involved in DNA repair 118(97) 123(97) 132(97) 103(41) 77(33) 33(26) 126* 130549 > 133707 1052 ATPase involved in DNA replication 119(99) 124(98) 133(98) 99(78) 78(45) 93(29) 127 133919 < 134410 163 BRCA1 like protein 120(98) 125(95) 134(98) 97(62) 80(29) – 128 134438 < 135244 268 hypothetical protein 121(97) 126(95) 135(97) 96(32) – – 129 135347 > 136873 508 helicase 122(95) 127(88) 136(95) 95(67) 82(39) – 130 136935 < 138371 478 bro15 123(98) 128(70) 137(76) 70(27) 34(29) 131 138520 < 139107 195 bro16 150(40) 97(50) 139(83) 79(32) 165(29) 34(29) 132 139265 < 140152 295 hypothetical protein 127(95) 131(85) 141(94) 94(25) – – 133* 140305 > 141231 308 patatin–like phospholipase 128(98) 132(98) 142(96) 93(72) 67(49) 87(27) 134* 141291 < 141950 219 hypothetical protein 129(100) 133(100) 143(99) 92(76) 60(54) 116(24) 135* 142269 > 142646 125 yabby–like transcription factor 130(100) 134(100) 144(100) 91(84) 59(61) 22(29) 136 142737 < 145694 985 lipopolysaccharide modifying enzyme 131(98) 135(97) 145(98) 90(54) 58(35) – 137 145736 > 146218 160 hemolysin–like protein 132(98) 136(99) 146(98) – 57(30) – 138 146215 < 146679 154 hypothetical protein 133(98) 137(96) 147(99) – – – 139 146711 < 147544 277 putative S1/P1 nuclease 134(99) 138(95) 148(99) 75(42) 135(68) – 140 147676 < 148314 212 hypothetical protein 135(99) 139(99) 149(99) – 54(42) – 141 148373 > 148816 147 hypothetical protein 136(91) 140(93) 150(91) – 142 148902 < 149483 193 hypothetical protein 137(97) 141(94) 151(97) – 52(35) – 143* 149517 < 150155 212 RNA polymerase subunit 138(98) 142(98) 152(98) 89(51) 51(29) 10(26)% 144 150188 < 150883 231 hypothetical protein 139(96) 143(95) 153(96) 88(41) – – 145 150870 < 151187 105 hypothetical protein – 144(88) – – – – 146* 151252 > 152064 270 fatty acid elongase 140(99) 145(98) 154(99) 87(70) 46(48) 15(34) 147 152099 > 152794 231 DEAD–like helicase 141(91) 146(90) 155(92) – 45(29) 148 153097 > 153723 208 hypothetical protein 142(95) 147(89) 156(94) – – 149 153887 < 154303 138 hypothetical protein 143(89) 148(90) 157(88) – – – 150 154409 > 155110 233 Uyr/REP helicase 144(98) 149(98) 158(98) 86(65) – 28(42) 151 155175 < 155786 203 hypothetical protein 145(95) 150(97) 159(96) 85(50) – – 152* 155844 > 159617 1257 Dynein–like β chain 146(99) 151(98) 160(99) 84(64) 43(34) 85(23) 153 159843 > 160766 307 hypothetical protein 148(95) 152(87) 161(93) – 68(60) – 154 160883 < 162403 506 bro17 149(77) 153(69) 162(88) 70(27) 165(31) 98(30) 155 162640 < 164091 483 bro18 149(75) 154(79) 162(69) 70(27) 165(27) 67(30) 156 164415 < 165569 384 hypothetical protein – 101(92) – – – – 157* 165772 < 166293 173 tyrosyl–DNA phosphodiesterase 151(90) 160(87) 164(96) 119(61) 162(38) 52(29) 158* 166315 < 166827 170 hypothetical protein 152(92) 161(89) 165(93) 119(32) 162(35) 52(26) 159 166957 > 167190 77 hypothetical protein – – – – – – 160 167208 < 168773 521 bro19 153(96) 163(89) 166(93) 79(45) 137(45) 34(28) 161 169273 > 169788 171 bro20 154(90) 164(77) 167(90) 79(45) 137(43) 98(28) 162 169835 > 170134 99 bro21 – 166(86) 43(31) – – – 163 170259 > 171353 364 bro22 157(98) 167(91) 169(90) 79(67) 165(83) 32(26) 164 171656 < 172540 294 hypothetical protein 158(94) 168(95) 170(96) 117(38) – – 165 172736 < 173089 117 hypothetical protein 159(83) 169(88) 171(83) – – – 166 173155 > 173700 181 hypothetical protein 160(95) 170(85) 172(96) – – – 167 174138 > 174620 160 RING–finger–containing E3 ubiquitin ligase 161(91) 172(98) 175(92) – – – 168 174608 > 175123 171 hypothetical protein 162(91) 173(91) 176(91) – – – 169 175201 > 176178 325 hypothetical protein 163(92) 174(91) 177(92) 30(38) – – 170 176262 > 176540 92 hypothetical protein 164(85) 175(89) 178(90) – – – 171 177151 < 178095 314 caspase–like protein 165(91) 176(94) 179(93) 73(37) 72(22) – 172 178429 > 179121 230 bro23 166(94) 177(95) 180(95) 72(32) 137(28) 75(24) 173 179156 > 179908 250 bro24 167(89) 178(94) 181(95) 30(32) – – 174 179966 < 180838 290 hypothetical protein 168(95) 181(95) 182(93) – 35(36) – 175 181107 > 181631 174 RING–finger–containing E3 ubiquitin ligase 169(92) – 183(97) 81(41) – – 176 181508 < 182572 354 DUF2661 superfamily – 182(93) 185(91) – – 78(27) 177 182672 > 184042 456 S–TKc serine/threonine
protein kinase170(96) 183(96) 186(96) 82(33) – – 178 184076 > 185143 355 hypothetical protein 171(94) 184(92) 187(93) 83(29) – – 179 185278 < 185715 145 hypothetical protein 172(89) 185(92) 188(92) – – – 180 185773 < 186219 148 hypothetical protein 173(95) 186(97) 189(95) – 35(26) – 181 186437 > 186772 111 hypothetical protein 174(97) 187(98) 190(97) – – – 182 186974 < 188161 395 hypothetical protein 175(98) 188(94) 191(97) – – – 183 188363 < 188578 71 hypothetical protein – – 192(96) – – – 184 188784 < 189602 272 hypothetical protein 177(92) 189(90) 193(92) 123(32) – – 185 189703 < 190365 220 hypothetical protein – 190(88) 194(88) – – – Note: # The length of amino acid; $ Amino acid identity with the homologue of HvAV-3h; * Conserved Ascoviridae proteins except bro genes; % indicates the low query cover and high E value. hrs, homologous regions; ORF, open reading frame; bro, baculovirus repeat ORFs; iap, inhibitor of apoptosis protein; SfAV, Spodoptera frugiperda ascovirus; TnAV, Trichoplusia ni ascovirus; HvAV, Heliothis virescens ascovirus; DpAV, Diadromus pulchellus ascovirus; kbp, kilo base pairs. Table S3. Annotation of the HvAV-3h genome
Genome analysis of Heliothis virescens ascovirus 3h isolated from China
- Received Date: 08 December 2016
- Accepted Date: 15 February 2017
- Published Date: 30 March 2017
Abstract: No ascovirus isolated from China has been sequenced so far.Therefore,in this study,we aimed to sequence the genome of Heliothis virescens ascovirus 3h (HvAV-3h) using the 454 pyrosequencing technology.The genome was found to be 190,519-bp long with a G+C content of 45.5%.We also found that it encodes 185 hypothetical open reading frames (ORFs) along with at least 50 amino acids,including 181 ORFs found in other ascoviruses and 4 unique ORFs.Gene-parity plots and phylogenetic analysis revealed a close relationship between HvAV-3h and three other HvAV-3a strains and a distant relationship with Spodoptera frugiperda ascovirus 1a (SfAV-1a),Trichoplusia ni ascovirus 6a (TnAV-6a),and Diadromus pulchellus ascovirus 4a (DpAV-4a).Among the 185 potential genes encoded by the genome,44 core genes were found in all the sequenced ascoviruses.In addition,25 genes were found to be conserved in all ascoviruses except DpAV-4a. In the HvAV-3h genome,24 baculovirus repeat ORFs (bros) were present,and the typical homologous repeat regions (hrs) were absent.This study supplies information important for understanding the conservation and functions of ascovirus genes as well as the variety of ascoviral genomes.
















 DownLoad:
DownLoad: