-
Porcine reproductive and respiratory syndrome (PRRS) was first observed in America in 1987, and subsequently spread worldwide, becoming one of the main infectious diseases in the swine industry and the cause of substantial economical loss. PPRS is characterized by pregnant sow reproductive failure, dyspenea, high mortality in piglets and respiratory symptoms in growing pigs [5, 7]. The PRRS virus (PRRSV), is a small enveloped virus with a single-stranded RNA genome containing eight ORFs. These are the viral replicase encoded by ORF1, which contains ORF1a and ORF1b; the virus envelope glycoproteins GP2, GP3, GP4 and GP5 encoded by ORFs 2-5 respectively [6]; the viral unglycosylated membrane protein M encoded by ORF6; and the highly conserved nucleocapsid protein N encoded by ORF7[4, 3]. GP5 protein is a major, multifunctional glycoprotein of 25 kDa. It has six antigenic determinants that can induce the production of antibodies that neutralize viral infection in vitro[1, 8]. GP5 shows the greatest variability of the PRRSV proteins, and stimulates cellular and humoral immunity. Therefore, establishing ELISA detection methods using GP5 protein as the antigen is essential. In vivo, antibodies against GP5 protein appears in the late stage of PRRSV infection.
A rapid and accurate diagnostic method for PRRS is essential for avoiding both detection failure, and unnecessary and expensive implementation of control measures. The current diagnostic techniques for PRRS antibodies are the American IDEXX PRRSV-Ab ELISA kit and the French LSI PRRSV-Ab ELISA kit. Although effective and sensitive, these methods are expensive, which is an important limitation. To explore additional detection options, we cloned and expressed the genes for the PRRSV major glycoprotein gene GP5 from the recent PRRSV isolate LZh/07, in Escherichia coli. We used the recombinant proteins to develop a method to detect PRRS antibody in swine.
HTML
-
The PRRSV LZh/07 strain used in this study was isolated and maintained in the State Key Laboratory of Veterinary Etiological Biology, Lanzhou Veterinary Research Institute of China. 50 Positive and 30 negative PRRSV sera were kindly provided by Dr. Licheng Zhang of the QingHai Disease Control and Prevention Center of China. A total of 300 clinical samples were collected from five provincial veterinary stations of China from 2006 to 2008. Foot-and-mouth disease (FMD), Classical swine fever (CSF), Swine vesicular disease (SVD) and Pseudorabies serum were obtained from the State Key Laboratory of Veterinary Etiological Biology, Lanzhou Veterinary Research Institute.
-
The complete ORF5 coding region from PRRSV LZh/07 was inserted into a pGEX 6P-1 plasmid (Novagen) using Xho I and BamH I restriction sites. Recombinant plasmids were transformed into E. coli (BL21) (Novagen) and protein expression induced with 1 mM IPTG. GP5 was expressed as fusion proteins with 26-kDa N-terminal GST tags. Expression was stimulated according to the pTriExTM SystemManual (Novagen).
The GP5 fusion protein was purified by resuspending E. coli cell pellets in a 0.1× culture volume of inclusion body (IB) wash buffer (20 mmol/L Tris-HCl, pH 7.5; 10 mmol/L EDTA; 1% Triton X-100) supplemented with lysozyme to a final concentration of 100 mg/mL. The mixture was incubated at room temperature for 15 min and sonicated on ice until no longer viscous. Inclusion bodies were collected by centrifugation at 10 000 × g for 10 min, washed twice with IB washing buffer, and once with a second wash buffer (20 mmol/L Tris-HCl, pH 7.5; 2 mol/L urea; 1% Triton X-100). Inclusion bodies were solubilized by the addition of IB solubilization buffer (50 mmol/L CAPS, 0.3% N-lauroylsarcosine, 1 mmol/L DTT, pH 11). Fusion protein was purified with a GST•BindTM Purification Kit (Novagen, Cat. No. 70541), according to the manufacturer's instructions. The purified fusion proteins were used as antigens for Western blotting and ELISA for detecting PRRSV antibodies.
Purified recombinant GP5 (0.5 μg/well) was separated on 10% SDS polyacrylamide gels and transferred to nitrocellulose membranes (Bio-Rad), incubated for 1 h in blocking solution (5% non-fat dry milk, 0.1% Tween-20 in phosphate buffered saline, PBS) and probed with a PRRSV-positive serum diluted 1:800 in blocking solution. Specific anti-PRRSV antibodies were detected with protein A-horseradish peroxidase conjugate (Sigma) and the reaction developed with luminol. Affinity chro-matography-purified GP5 proteins was stored at -70℃ for use as ELISA antigens.
-
A checkerboard titration was performed to determine the optimal working dilution of the coating antigen, serum and horseradish peroxidase-labelled rabbit-anti-swine IgG (HRP-IgG) (Sigma) using a 96-well ELISA plate. Antigen coating concentrations were 0.8 μg, 0.4 μg, 0.2 μg, 0.1 μg and 0.05 μg per well, and serum dilutions were 1:10, 1:20, 1:40 and 1:80. The dilutions that gave the maximum difference between positive and negative serum (P/N) by absorbance at 450 nm were selected for large-scale testing of serum samples. Test sera included positive, negative and blank sample controls. The reaction temperature, time and other conditions were optimised by P/N value[2].
After optimisation, indirect ELISA was performed using the following procedure. Ninety-six well ELISA plates (Costar) were coated with 0.2 μg total purified fusion protein per well, diluted in carbonate buffer, and incubated overnight at 4℃. After three washes with phosphate buffered saline (PBS) containing 0.05% Tween 20 (PBST), plates were sealed with 10 mg/mL gelatin (Sigma), and incubated for 45 min at 37℃. After three washes with PBST, serum samples were diluted 1:40 in dilution buffer (PBS containing 5% skimmed milk, 10% horse serum and 3% E. coli lysate), in a 100 µL volume per well, and incubated for 45 min at 37℃. After three washes, horseradish peroxidase-conjugated rabbit anti-swine serum (Sigma) was added in the same dilution buffer at an appropriate working concentration, 100 µL per well, and incubated at 37 C for 30 min. After three washes, color was developed with 3, 3′, 5, 5′-tetramethyl-benzidine (TMB, Sigma), and the reaction was stopped after 15 min with 2.0 mol/L H2SO4. The OD450 was read with a microplate reader (Model 680. Bio-Rad).
-
Indirect immunofluorescence was performed on Marc-145 cells infected with PRRSV. Preconfluent Marc-145 cells were infected with PRRSV LZh/07 isolate for 20 h, washed with PBS, fixed at -20℃ in methanol for 5 min, washed again and incubated for 1 h at room temperature with 1:40 dilutions of serum samples in PBS plus 0.1% bovine serum albumin. After PBS washing, slides were incubated with protein A-FITC conjugate (Sigma) diluted 1:50 in PBS for 30 min at room temperature, further washed, mounted and examined by fluorescence microscopy.
-
To set negative/positive cutoff values, 50 positive and 30 negative samples were tested in duplicate by ELISA. The P/N value (OD450 of test serum / negative serum) of the 80 serum samples was compared with the indirect immunofluorescence assay (IIFA) results. The end-point cut-off was determined by analysis of a receiver operating characteristic (ROC) curve based on 80 P/N values. Cut-off values were determined as the mean + 2 standard deviations (SDs) and mean + 3 SDs derived from the P/N values from the 30 negative samples. Estimates of diagnostic sensitivity and specificity were calculated using the three cut-off values.
For the test system to be valid, we determined that the ODpos should be higher than 0.5 and at least three times higher than the ODneg, so the ODneg should be lower than 0.2. For values outside these limits, the test was repeated.
-
To validate the test and evaluate repeatability, the co-efficient of variation (CV) was calculated between plates (inter-assay variation) and within the same plate (intra-assay variation) for 20 sera samples. For inter-assay CV, each sample was tested on four different plates on different occasions, and for intra-assay CV, four replicates within each plate were assayed. A total of 300 sera samples were used to determine agreement of the three tests (including two ELISA kit and IIFA).
-
Four samples positive serologically for related swine viruses were tested by the indirect ELISA to assess the degree of assay cross-reactivity.
Virus strain and serum sample
Recombinant protein expression, purification andWestern blot analysis
Development of indirect ELISA
Indirect immunofluorescence
Determination of cut-off values
Validation, repeatability and comparison of the GP5-Ⅰ-ELISA
Assessment of cross-reactivity
-
Western blotting for recombinant GP5 protein showed a specific protein band with a molecular weight of 42 kDa for the GP5 protein, after induction at 28℃ (Fig. 1). This was consistent with the expected sizes, since GST added 26 kDa to the molecular mass of 16 kDa for GP5. The expression level peaked at 12 h after induction. Chromatography scanning suggested that recombinant GP5 was 30% of the total protein content. The purity of the target protein after affinity chromatography reached 90%. Bands corresponding to the target proteins were observed by Western blot analysis (Fig. 1), indicating reaction with anti-PRRSV serum.
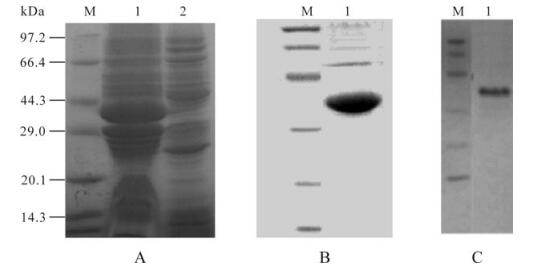
Figure 1. SDS-PAGE, Western blotting analysis of the expressed GP5 protein. A: SDS-PAGE detection the cell lysate of E. coli for expressing GP5 protein. Lane 1, The cell lysate of Escherichia coli expressed GP5 protein; 2, The uninduced bacteria harboring 6P-1-5/BL21; M, The protein marker. B: SDS–PAGE analysis. C: Western blot analysis of the purified recombinant protein. Lane 1, GP5 protein; M, Protein marker.
-
By checkerboard ELISA, the optimal concentration of coating antigen was determined to be 0.2 μg/well GP5 protein. The optimal antibody dilutions were 1:40 for serum and 1:40 000 for HRP-IgG. Optimal exposure time was 45 min for serum samples and 15 min for the conjugate at 37℃.
-
Receiver-operating characteristic curve analysis was used to set a cut-off value of 0.2, determined from a mean P/N of 1.516 with a SD of 0.056 for the 30 PRRSV negative sera. The specificity and sensitivity of the assay at three different cut-off values, 0.10, 0.2 and 0.3, are shown in Table 1. At the cut-off value of 0.2, the sensitivity and specificity were relatively high for three animal species. Thus, the cut-off values were established to be: positive≥ 0.3; suspicious, 0.2 -0.3; negative≤0.2. Serum samples classified as suspicious were re-tested, with the sample judged to be positive if the result was confirmed by the repeat test.

Table 1. Specificity and sensitivity according to three different cut-off values
-
The inter-assay CV ranged from 1.9% to 5.8%, and the intra-assay CV ranged from 1.4% to 5.0%, for 20 sera selected for validation testing (Table 2). When comparing the GP5-Ⅰ-ELISA with the LSI PRRSV Ab-ELISA, the same results were obtained in 266 of 300 samples for an the agreement rate of 88.7% (Table 3).

Table 2. Intra-and inter-assay coefficient of variation (CV) obtained from assessment of 20 sera

Table 3. Comparison of PRRSV GP5-Ⅰ-ELISA with LSIR PRRSV Ab-ELISA
-
No evidence of cross-reactivity with known positive sera to swine viruses was observed. The results show that four samples positive serologically for related swine viruses have no cross-reactivity. All tested sera gave values below the defined cut-off point (Table 4).

Table 4. Results of cross-reactivity test
Purification and Western blot of recombinant GP5
Development of the GP5 indirect ELISA (Ⅰ-ELISA) assay
Determination of the cut-off value
Validation of the GP5 -Ⅰ-ELISA
Assessment of cross-reactivity
-
The glycosylated envelope protein GP5 is one of three major structure proteins in PRRSV and has good immunogenicity, inducing the secretion of both neutralizing and non-neutralizing antibodies. Thus, GP5 is important in pathogenicity, diagnosis, prevention and control of PRRSV. Therefore, establishing ELISA detection methods with GP5 protein as the antigen is essential, since in PRRSV-infected pigs, late stage antibodies are mainly produced against GP5 protein. This study is report the use of the GP5 protein to develop an ELISA method for detecting PRRSV antibodies in pigs.
In this study, we analyzed the fusion protein expressed by the pGEX plasmid for use as a diagnostic method to detect PRRSV antibodies. Although the pGEX expression system has been successfully used to obtain a number of recombinant proteins in soluble form, the recombinant PRRSV GP5 protein expressed in E.coli in non-denaturing conditions was insoluble after mechanical cell lysis. However, GP5-GST could be solubilised with 8 mol/L urea, and using this procedure we recovered approximately 330 μg of GP5-GST per mL of extract, which corresponds to a yield of more than 60 mg of GP5-GST per liter of bacterial culture. The polypeptides obtained gave a single homogeneous band of 42 kD by SDS-PAGE analysis, and purified GP5 was recognized by PRRSV-positive serum in Western blots.
Serological studies have proven that GP5 protein has important roles in PRRSV diagnosis. Therefore, we developed an indirect ELISA by coating polystyrene plates with a combination of purified GP5 protein. Preliminary optimization of the assay showed that the best results were obtained with 0.2 μg purified recombinant GP5 in each well, followed by blocking with 10 mg/mL gelatin. These conditions allowed maximum OD450 absorbance. A crucial factor for establishing an indirect ELISA is eliminating potential false-negatives and false-positives that result from the variable antibody titres of different sera [2, 9]. To address this issue, we performed ELISA on serial dilutions to determine an optimal cut-off using a ROC curve based on 80 P/N values. The obtained cut-off value gave an assay with a high degree of specificity and sensitivity, with an inter-assay CV of 69.28% and an intra-assay of 66.0%. The assay also had good repeatability and promises to be useful in clinical contexts.
In summary, the GP5-Ⅰ-ELISA established here was sensitive and specific for PRRSV antibody detection, and was easier to produce and perform, and less expensive, than existing serological methods for PRRSV antibody detection. The results suggest that the GP5-Ⅰ-ELISA could be used to develop a reliable tool for large scale detection of PRRSV antibodies in herd tests.














 DownLoad:
DownLoad: