Structural Virology
Editor(s): Peng Gong, PhD, Professor Wuhan Institute of Virology, China
A special topic section on Structural Virology is included in this issue. The section contains three review articles, which cover the recent research advancement on the structures of reovirus particle, flavivirus NS2B-NS3 protease, and enterophage T7 RNA polymerase. The issue cover presents: i) Crystal structure of a late initiation complex of T7 RNA Polymerase; ii) Crystal structure of the NS2B-NS3 protease with a peptide-based inhibitor; iii) Cryo-EM structure of the turreted reovirus core.
A special topic section on Structural Virology is included in this issue. The section contains three review articles, which cover the recent research advancement on the structures of reovirus particle, flavivirus NS2B-NS3 protease, and enterophage T7 RNA polymerase. The issue cover presents: i) Crystal structure of a late initiation complex of T7 RNA Polymerase; ii) Crystal structure of the NS2B-NS3 protease with a peptide-based inhibitor; iii) Cryo-EM structure of the turreted reovirus core.
-
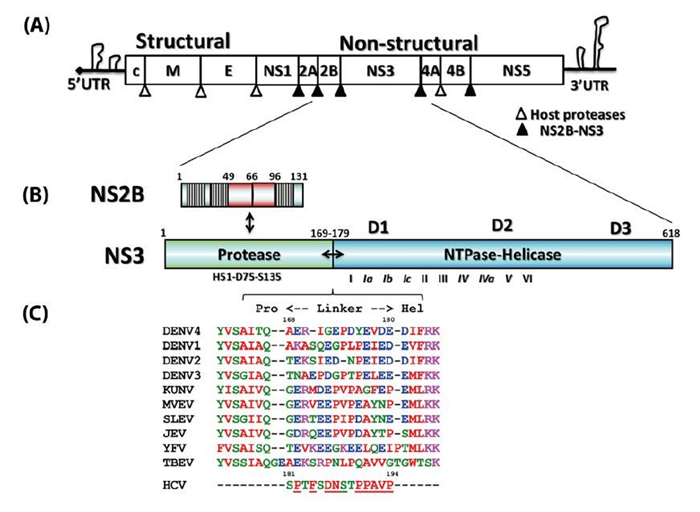
Functional interplay among the flavivirus NS3 protease, helicase, and cofactors
2014, 29(2): 74 doi: 10.1007/s12250-014-3438-6
Received: 21 January 2014 Accepted: 19 March 2014 Published: 26 March 2014Flaviviruses are positive-sense RNA viruses, and many are important human pathogens. Nonstructural protein 2B and 3 of the flaviviruses (NS2BNS3) form an endoplasmic reticulum (ER) membraneassociated hetero-dimeric complex through the NS2B transmembrane region. The NS2BNS3 complex is multifunctional. The N-terminal region of NS3, and its cofactor NS2B fold into a protease that is responsible for viral polyprotein processing, and the C-terminal domain of NS3 possesses NTPase/RNA helicase activities and is involved in viral RNA replication and virus particle formation. In addition, NS2BNS3 complex has also been shown to modulate viral pathogenesis and the host immune response. Because of the essential functions that the NS2BNS3 complex plays in the flavivirus life cycle, it is an attractive target for antiviral development. This review focuses on the recent biochemical and structural advances of NS2BNS3 and provides a brief update on the current status of drug development targeting this viral protein complex. -

Snapshots of a Viral RNA Polymerase Switching Gears from Transcription Initiation to Elongation
2013, 28(6): 337 doi: 10.1007/s12250-013-3397-3
Received: 23 October 2013 Accepted: 28 November 2013 Published: 02 December 2013During transcription initiation, RNA polymerase binds tightly to the promoter DNA defining the start of transcription, transcribes comparatively slowly, and frequently releases short transcripts (3-8 nucleotides) in a process called abortive cycling. Transitioning to elongation, the second phase of transcription, the polymerase dissociates from the promoter while RNA synthesis continues. Elongation is characterized by higher rates of transcription and tight binding to the RNA transcript. The RNA polymerase from enterophage T7 (T7 RNAP) has been used as a model to understand the mechanism of transcription in general, and the transition from initiation to elongation specifically. This single-subunit enzyme undergoes dramatic conformational changes during this transition to support the changing requirements of nucleic acid interactions while continuously maintaining polymerase function. Crystal structures, available of multiple stages of the initiation complex and of the elongation complex, combined with biochemical and biophysical data, offer molecular detail of the transition. Some of the crystal structures contain a variant of T7 RNAP where proline 266 is substituted by leucine. This variant shows less abortive products and altered timing of transition, and is a valuable tool to study these processes. The structural transitions from early to late initiation are well understood and are consistent with solution data. The timing of events and the structural intermediates in the transition from late initiation to elongation are less well understood, but the available data allows one to formulate testable models of the transition to guide further research. -

The Flavivirus Protease As a Target for Drug Discovery
2013, 28(6): 326 doi: 10.1007/s12250-013-3390-x
Received: 08 October 2013 Accepted: 01 November 2013 Published: 14 November 2013Many flaviviruses are significant human pathogens causing considerable disease burdens, including encephalitis and hemorrhagic fever, in the regions in which they are endemic. A paucity of treatments for flaviviral infections has driven interest in drug development targeting proteins essential to flavivirus replication, such as the viral protease. During viral replication, the flavivirus genome is translated as a single polyprotein precursor, which must be cleaved into individual proteins by a complex of the viral protease, NS3, and its cofactor, NS2B. Because this cleavage is an obligate step of the viral life-cycle, the flavivirus protease is an attractive target for antiviral drug development. In this review, we will survey recent drug development studies targeting the NS3 active site, as well as studies targeting an NS2B/NS3 interaction site determined from flavivirus protease crystal structures. -
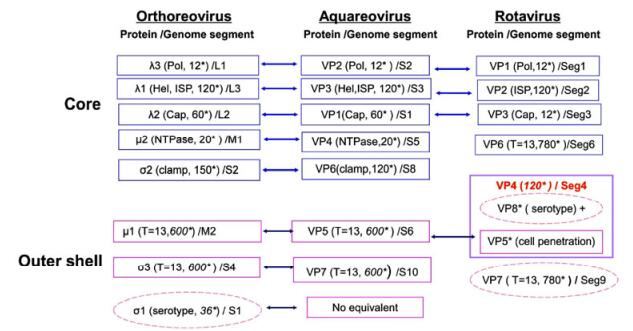
High-resolution 3D Structures Reveal the Biological Functions of Reoviruses
2013, 28(6): 318 doi: 10.1007/s12250-013-3341-6
Received: 05 May 2013 Accepted: 30 September 2013 Published: 06 November 2013Viruses in the family Reoviridae are non-enveloped particles comprising a segmented double-stranded RNA genome surrounded by a two-layered or multi-layered icosahedral protein capsid. These viruses are classified into two sub-families based on their particle structural organization. Recent studies have focused on high-resolution three-dimensional structures of reovirus particles by using cryo-electron microscopy (cryo-EM) to approach the resolutions seen in X-ray crystallographic structures. The results of cryo-EM image reconstructions allow tracing of most of the protein side chains, and thus permit integration of structural and functional information into a coherent mechanism for reovirus assembly and entry.