Column
|
Mechanisms of HIV envelope-induced T lymphocyte apoptosis
2010, 25(5): 307 doi: 10.1007/s12250-010-3148-7
Received: 29 April 2010
Accepted: 13 June 2010
Infection by the human immunodeficiency virus (HIV) is characterized by a progressive depletion of CD4 T lymphocytes, which leads to dysfunction of the immune system. Although a variety of mechanisms may contribute to the gradual T cell decline that occurs in HIV-infected patients, abnormal apoptosis of infected or bystander T lymphocytes is an important event leading to immunodeficiency. The HIV envelope glycoprotein plays a crucial role in HIV associated apoptosis through both death receptor-mediated and mitochondria-dependent pathways. This review summarizes current knowledge of Env-mediated T lymphocyte apoptosis.
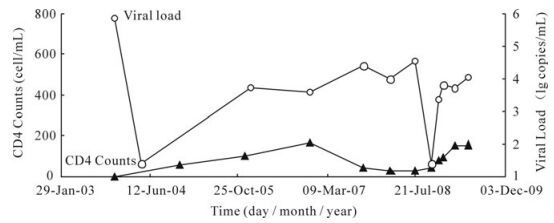
Indinavir Resistance Evolution in One Human Immunodeficiency Virus Type 1 Infected Patient Revealed by Single-Genome Amplification*
2010, 25(5): 316 doi: 10.1007/s12250-010-3122-4
Received: 31 December 2009
Accepted: 07 May 2010
1.???????? Alexander D C. 1990. A rapid method for determining clone frequency in complex populations using PCR and the Poisson distribution. Nucl Acids Res, 18(24):7453-7454. 2.???????? Bélec L, Piketty C, Si-Mohamed A, et al. 2000. High levels of drug-resistant Human Immunodeficiency Virus variants in patients exhibiting increasing CD4+ T cell counts despite virologic failure of protease inhibitor-containing antiretroviral combination therapy. J Infect Dis, 181(5):1808-1812. 3.???????? Chen Z, Li Y, Schock H B, et al. 1995. Three-dimensional structure of a mutant HIV-1 protease displaying cross-resistance to all protease inhibitors in clinical trials. J Biol Chem, 270(37):21433-21436. 4.???????? Clavel F, Hance A J. 2004. HIV drug resistance. N Engl J Med, 350(10):1023-1035. 5.???????? Coffin J M. 1995. HIV population dynamics in vivo: implications for genetic variation, pathogenesis, and therapy. Science, 267:483-489. 6.???????? Condra J H, Holder D J, Schleif W A, et al. 1996.Genetic ? 328 correlates of in vivo viral resistance to Indinavir, a Human Immunodeficiency Virus type 1 protease inhibitor. J Virol, 70(12):8270-8276. 7.???????? Descamps D, Joly V, Flandre P, et al. 2005. Genotypic resistance analyses in nucleoside-pretreated patients failing an indinavir containing regimen: results from a randomized comparative trial: (Novavir ANRS 073). J Clin Virol, 33:99-103. 8.???????? Dybul M, Daucher M, Jensen M A, et al. 2003. Genetic characterization of rebounding Human Immunodeficiency Virus type 1 in plasma during multiple interruptions of highly active antiretroviral therapy. J Virol, 77(5):3229-3237. 9.???????? Gunthard H F, Wong J K, Ignacio C C, et al. 1998. Comparative performance of high-density oligonucleotide sequencing and dideoxynucleotide sequencing of HIV type 1 pol from clinical samples. AIDS Res Hum Retroviruses, 14:869-876. 10.????? Kaplan A H, Michael S F, Wehbie R S, et al. 1994. Selection of multiple Human Immunodeficiency Virus type 1 variants that encode viral proteases with decreased sensitivity to an inhibitor of the viral protease. Proc. Natd. Acad. Sci, 91:5597-5601. 11.
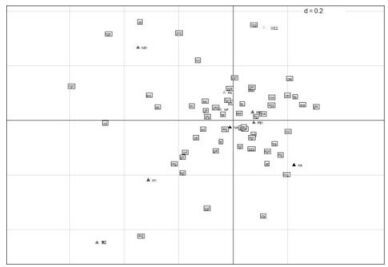
Analysis of Synonymous Codon Usage Bias in 09H1N1
2010, 25(5): 329 doi: 10.1007/s12250-010-3123-3
Received: 05 January 2010
Accepted: 30 April 2010
A novel subtype of influenza A virus 09H1N1 has rapidly spread across the world. Evolutionary analyses of this virus have revealed that 09H1N1 is a triple reassortant of segments from swine, avian and human influenza viruses. In this study, we investigated factors shaping the codon usage bias of 09H1N1 and carried out cluster analysis of 60 strains of influenza A virus from different subtypes based on their codon usage bias. We discovered that more preferentially used codons of 09H1N1 are A-ended or U-ended, and the intra-genomic codon usage bias of 09H1N1 is quite low. Base composition constraint, dinucleotide biases and translational selection are the main factors influencing the codon usage bias of 09H1N1. At the genome level, we find that the codon usage bias of 09H1N1 is similar to H1N1 (A/swine/Kansas/77778/2007H1N1), H9N2 from Asia, H1N2 from Asia and North America and H3N2 from North America. Our results provide insight for understanding the processes governing evolution, regulation of gene expression, and revealing the evolution of 09H1N1.

In Vitro Anti-influenza Virus Activities of Sulfated Polysaccharide Fractions from Gracilaria lemaneiformis*
2010, 25(5): 341 doi: 10.1007/s12250-010-3137-x
Received: 22 March 2010
Accepted: 07 July 2010
In this paper, in vitro anti-influenza virus activities of sulfated polysaccharide fractions from Gracilaria lemaneiformis were investigated. Cytotoxicities and antiviral activities of Gracilaria lemaneiformis polysaccharides (PGL), Gracilaria lemaneiformis polysaccharide fraction-1 (GL-1), Gracilaria lemaneiformis polysaccharide fraction-2 (GL-2) and Gracilaria lemaneiformis polysaccharide fraction-3 (GL-3) were studied by the Methyl thiazolyl tetrazolium (MTT) method, and the inhibitory effect against Human influenza virus H1-364 induced cytopathic effect (CPE) on MDCK cells were observed by the CPE method. In addition, the antiviral mechanism of PGL was explored by Plaque forming unit (PFU), MTT and CPE methods. The results showed: i) Cytotoxicities were not significantly revealed, and H1-364 induced CPE was also reduced treated with sulfated polysaccharide fractions from Gracilaria lemaneiformis; ii) Antiviral activities were associated with the mass percentage content of sulfate groups in polysaccharide fractions, which was about 13%, in polysaccharides (PGL and GL-2) both of which exhibited higher antiviral activity; iii) A potential antiviral mechanism to explain these observations is that viral adsorption and replication on host cells were inhibited by sulfated polysaccharides from Gracilaria lemaneiformis. In conclusion, Anti-influenza virus activities of sulfated polysaccharide fractions from Gracilaria lemaneiformis were revealed, and the antiviral activities were associated with content of sulfate groups in polysaccharide fractions.
Development of a Polyclonal Antibody-based AC-ELISA and Its Comparison with PCR for Diagnosis of Canine Parvovirus Infection
2010, 25(5): 352 doi: 10.1007/s12250-010-3132-x
Received: 15 March 2010
Accepted: 09 June 2010
A polyclonal antibody-based antigen-capture ELISA (AC-ELISA) has been developed for detection of Canine parvovirus (CPV) antigens in faecal samples of dogs. The assay uses rabbit anti-CPV polyclonal antibody as the capture antibody, guinea pig anti-CPV polyclonal antibody as tracing antibody and anti-guinea pig HRPO conjugate as the detection system. The optimum dilution of the capture antibody and the tracing antibody capable of detecting the CPV-2 antigens was found to be 1:1 600 and 1:400, respectively, in the check-board titration. In this study, a total of 152 samples (129 faecal samples and 23 cell culture supernatant) were tested both by AC-ELISA and by polymerase chain reaction (PCR). Of the samples tested, 69 and 78 samples were found positive by AC-ELISA and PCR, respectively. The AC-ELISA had relative sensitivity, relative specificity and accuracy of 88.4%, 100.0% and 91.4% respectively. The analytical sensitivity of AC-ELISA was estimated to be 102.8 TCID50/mL whereas PCR sensitivity was 100.8 TCID50/mL. The AC-ELISA is a simple, quick and reliable method for screening large numbers of faecal samples of dogs suspected of CPV infection.

Establishment of the Eukaryotic Cell Lines for Inducible Control of SARS-CoV Nucleocapsid Gene Expression*
2010, 25(5): 361 doi: 10.1007/s12250-010-3124-2
Received: 11 January 2010
Accepted: 30 April 2010
In order to establish the eukaryotic cell lines for inducible control of SARS-CoV nucleocapsid gene expression.The recombinant plasmid of pTRE-Tight-SARS-N was constructed by using the plasmid p8S as the PCR template which contains a cDNA clone covering the nucleocapsid gene of SARS-CoV HKU-39449. Restriction enzymes digestion and sequence analysis indicated the recombinant plasmid of pTRE-Tight-SARS-N contained the nucleocapsid gene with the optimized nucleotide sequence which will improve the translation efficiency. Positive cell clones were selected by cotransfecting pTRE-Tight-SARS-N with the linear marker pPUR to BHK-21 Tet-on cells in the presence of puromycin. A set of double-stable eukaryotic cell lines (BHK-Tet-SARS-N) with inducible control of the SARS-CoV neucleocapsid gene expression was identified by using SDS-PAGE and Western-blot analysis. The expression of SARS-CoV nucleocapsid protein was tightly regulated by the varying concentration of doxcycline in the constructed double-stable cell line. The constructed BHK-Tet-SARS-N cell strains will facilitate the rescue of SARS-CoV in vitro and the further reverse genetic research of SARS-CoV.

Cxcl16 Interact With SARS-CoV N Protein In and Out Cell
2010, 25(5): 369 doi: 10.1007/s12250-010-3129-x
Received: 22 January 2010
Accepted: 30 June 2010
Our study investigated the host cell protein which can interact with SARS-CoV N protein, and explored the functional connections. The eukaryotic expression vectors pEGFP-N1/SARS-CoVN and pdsRed2-N1/ CXCL16 were constructed and used to co-transfect HEK293FT cells by the calcium phosphate method. The HIS-tagged fusion protein SARS-CoVN-GFP was then built and purified for the binding assay in vitro. The co-localization of SARS-CoVN and CXCL16 in the cytoplasm of HEK293FT cells was also shown using confocal laser scanning microscopy. It is suggested that their interaction might be through direct combination. Under a fluorescence microscope, it was observed that the purified fusion protein SARS-CoVN-GFP was attached to the cell membrane of CXCL16-transfected cells, indicating that SARS-CoVN and CXCL16 can be mutually combined.
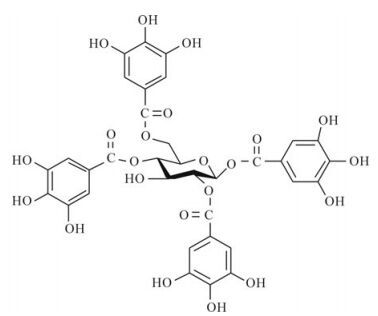
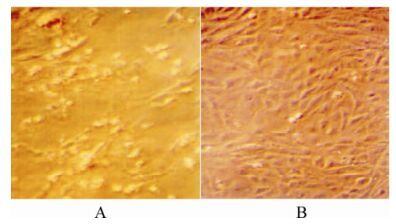
Effects of 1, 2, 4, 6-tetra-O-galloyl-β-D-glucose from P. emblica on HBsAg and HBeAg Secretion in HepG2.2.15 Cell Culture*
2010, 25(5): 375 doi: 10.1007/s12250-010-3144-y
Received: 14 April 2010
Accepted: 07 June 2010
A polyphenolic compound, 1,2,4,6-tetra-O-galloyl-β-D-glucose (1246TGG), was isolated from the traditional Chinese medicine Phyllanthus emblica L. (Euphorbiaceae) and assayed for its potential as an anti-hepatitis B virus (HBV) agent. The cytotoxicity of 1246TGG on HepG2.2.15 as well as HepG2 cells was determined by observing cytopathic effects, and the effects of 1246TGG on secretion of HBsAg and HBeAg in HepG2.2.15 cells were assayed by enzyme immunoassay. Results indicates that treatment with 1246TGG (6.25 μg/mL, 3.13 μg/mL), reduced both HBsAg and HBeAg levels in culture supernatant, yet the inhibitory effects tend to decline with the assay time. This study provides a basis for further investigation of the anti-HBV activity and possible mechanism of action of 1246TGG.