Laboratory Detection and Diagnosis of Filoviruses
2011, 26(2): 73 doi: 10.1007/s12250-011-3186-9
Received: 28 January 2011
Accepted: 21 February 2011
Ebola virus (EBOV) and Marburg virus (MARV), belonging to the Filoviridae family, emerged four decades ago and caused severe viral hemorrhagic fever in human and other primates. As high as 50-90% mortality, filoviruses can cause significant threats to public health. However, so far no specific and efficient vaccine has been available, nor have other treatment methods proved to be effective. It is of great importance to detect these pathogens specific, rapidly and sensitively in order to control future filovirus outbreaks. Here, recent progresses in the development of detection and diagnosis methods for EBOV and MARV are summarized.

PIKA Provides an Adjuvant Effect to Induce Strong Mucosal and Systemic Humoral Immunity Against SARS-CoV*
2011, 26(2): 81 doi: 10.1007/s12250-011-3183-z
Received: 17 January 2011
Accepted: 18 February 2011
Severe Acute Respiratory Syndrome (SARS) is a deadly infectious disease caused by SARS Coronavirus (SARS-CoV). Inactivated SARS-CoV has been explored as a vaccine against SARS-CoV. However, safe and potent adjuvants, especially with more efficient and economical needle-free vaccination are always needed more urgently in a pandemic. The development of a safe and effective mucosal adjuvant and vaccine for prevention of emergent infectious diseases such as SARS will be an important advancement. PIKA, a stabilized derivative of Poly (I:C), was previously reported to be safe and potent as adjuvant in mouse models. In the present study, we demonstrated that the intraperitoneal and intranasal co-administration of inactivated SARS-CoV vaccine together with this improved Poly (I:C) derivative induced strong anti-SARS-CoV mucosal and systemic humoral immune responses with neutralizing activity against pseudotyped virus. Although intraperitoneal immunization of inactivated SARS-CoV vaccine alone could induce a certain level of neutralizing activity in serum as well as in mucosal sites, co-administration of inactivated SARS-CoV vaccine with PIKA as adjuvant could induce a much higher neutralizing activity. When intranasal immunization was used, PIKA was obligatorily for inducing neutralizing activity in serum as well as in mucosal sites and was correlated with both mucosal IgA and mucosal IgG response. Overall, PIKA could be a good mucosal adjuvant candidate for inactivated SARS-CoV vaccine for use in possible future pandemic.
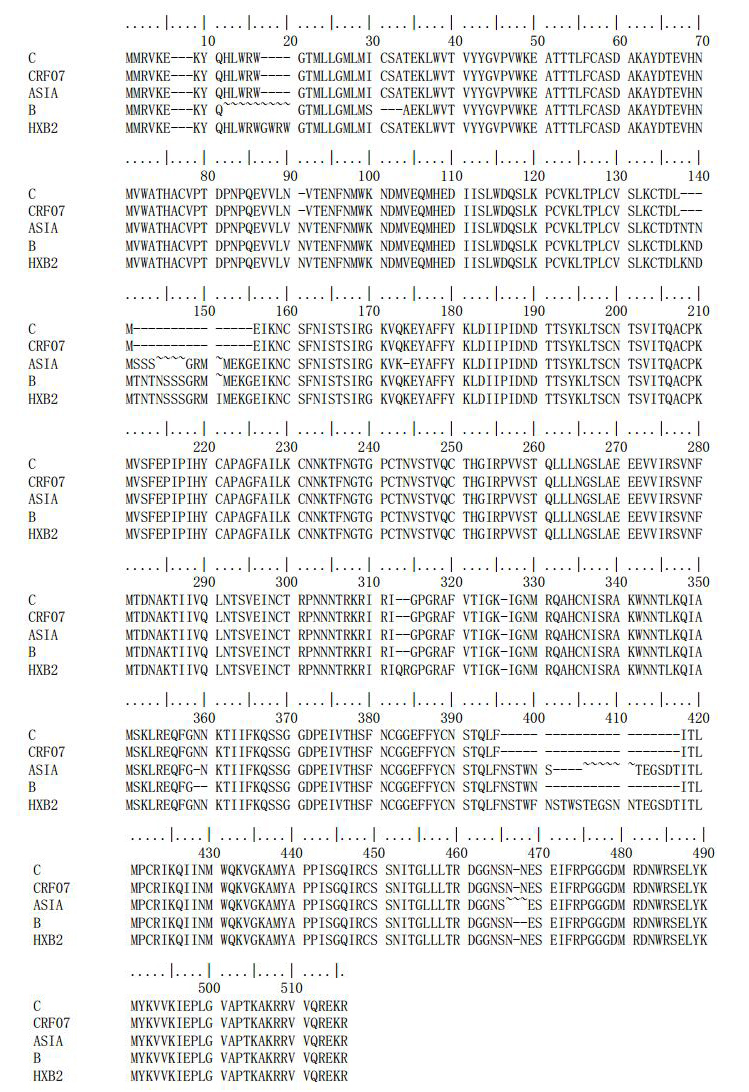

Use of Mutual Information Arrays to Predict Coevolving Sites in the Full Length HIV gp120 Protein for Subtypes B and C
2011, 26(2): 95 doi: 10.1007/s12250-011-3188-7
Received: 31 January 2011
Accepted: 22 February 2011
It is well established that different sites within a protein evolve at different rates according to their role within the protein; identification of these correlated mutations can aid in tasks such as ab initio protein structure, structure function analysis or sequence alignment. Mutual Information is a standard measure for coevolution between two sites but its application is limited by signal to noise ratio. In this work we report a preliminary study to investigate whether larger sequence sets could circumvent this problem by calculating mutual information arrays for two sets of drug na?ve sequences from the HIV gp120 protein for the B and C subtypes. Our results suggest that while the larger sequences sets can improve the signal to noise ratio, the gain is offset by the high mutation rate of the HIV virus which makes it more difficult to achieve consistent alignments. Nevertheless, we were able to predict a number of coevolving sites that were supported by previous experimental studies as well as a region close to the C terminal of the protein that was highly variable in the C subtype but highly conserved in the B subtype.
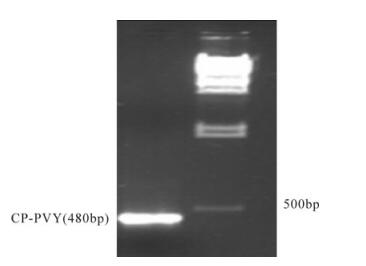
Potato Virus Y mRNA Expression Knockdown Mediated by siRNAs in Cultured Mammalian Cell Line
2011, 26(2): 105 doi: 10.1007/s12250-011-3161-x
Received: 28 July 2010
Accepted: 04 January 2011
RNA interference (RNAi) is a powerful tool for functional gene analysis which has been successfully used to downregulate the expression levels of target genes. The goal of this research was to provide a highly robust and concise methodology for in-vitro screening of efficient siRNAs from a bulk to be used as a tool to protect potato plants against PVY invasion. In our study, a 480bp fragment of the capsid protein gene of potato virus Y (CP-PVY) was used as a target to downregulate PVY mRNA expression in-vitro, as the CP gene interferes with viral uncoating, translation and replication. A total of six siRNAs were designed and screened through transient transfection assay and knockdown in expression of CP-PVY mRNA was calculated in CHO-k cells. CP-PVY mRNA knockdown efficiency was analyzed by RT-PCR and real-time PCR of CHO-k cells co-transfected with a CP gene construct and siRNAs. Six biological replicates were performed in this study. In our findings, one CP gene specific siRNA out of a total of six was found to be the most effective for knockdown of CP-PVY mRNA in transfected CHO-k cells by up to 80%-90%.

Down-regulation of HIV-1 Infection by Inhibition of the MAPK Signaling Pathway*
2011, 26(2): 114 doi: 10.1007/s12250-011-3184-y
Received: 20 January 2011
Accepted: 15 February 2011
The human immunodeficiency virus type 1 (HIV-1) can interact with and exploit the host cellular machinery to replicate and propagate itself. Numerous studies have shown that the Mitogen-activated protein kinase (MAPK) signal pathway can positively regulate the replication of HIV-1, but exactly how each MAPK pathway affects HIV-1 infection and replication is not understood. In this study, we used the Extracellular signal-regulated kinase (ERK) pathway inhibitor, PD98059, the Jun N-terminal kinase (JNK) pathway inhibitor, SP600125, and the p38 pathway inhibitor, SB203580, to investigate the roles of these pathways in HIV-1 replication. We found that application of PD98059 results in a strong VSV-G pseudotyped HIV-1NL4-3 luciferase reporter virus and HIV-1NL4-3 virus inhibition activity. In addition, SB203580 and SP600125 also elicited marked VSV-G pseudotyped HIV-1NL4-3 luciferase reporter virus inhibition activity but no HIV-1NL4-3 virus inhibition activity. We also found that SB203580 and SP600125 can enhance the HIV-1 inhibition activity of PD98059 when cells were treated with all three MAPK pathway inhibitors in combination. Finally, we show that HIV-1 virus inhibition activity of the MAPK pathway inhibitors was the result of the negative regulation of HIV-1 LTR promoter activity.
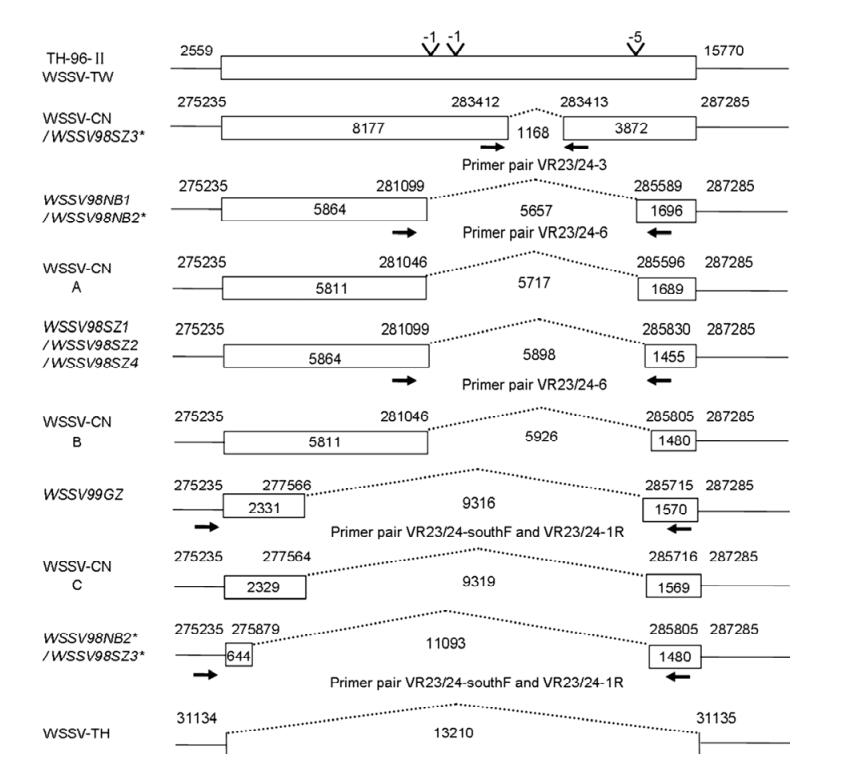
Genotyping of White Spot Syndrome Virus in Chinese Cultured Shrimp during 1998-1999*
2011, 26(2): 123 doi: 10.1007/s12250-011-3181-1
Received: 11 January 2011
Accepted: 18 February 2011
Recent studies showed that white spot syndrome virus (WSSV) isolates from different geographic locations share a high genetic similarity except the variable regions in ORF23/24 and ORF14/15, and variable number of tandem repeats (VNTR) within ORF94. In this study, genotyping was performed according to these three variable regions among WSSV isolates collected during 1998/1999 from Southern China. These WSSV isolates contain a deletion of 1168, 5657, 5898, 9316 and 11093 bp, respectively in the variable region ORF23/24 compared with WSSV-TW, and a deletion of 4749 or 5622 bp in the variable region ORF14/15 relative to TH-96-II. Four types of repeat units (RUs) (6, 8, 9 and 13 RUs) in ORF94 were detected in these isolates, with the shortest 6 RUs as the most prevalent type. Our results provide important information for a better understanding of the spatio-temporal transmission mode and the WSSV genetic evolution lineage.

Novel Evidence Suggests Hepatitis B Virus Surface Proteins Participate in Regulation of HBV Genome Replication*
2011, 26(2): 131 doi: 10.1007/s12250-011-3190-0
Received: 14 February 2011
Accepted: 02 March 2011
Naturally occurring mutations in surface proteins of Hepatitis B virus (HBV) usually result in altered hepatitis B surface antigen (HBsAg) secretion efficiency. In the present study, we reported two conserved residues, M75 and M103 with respect to HBsAg, mutations of which not only attenuated HBsAg secretion (M75 only), but also suppressed HBV genome replication without compromising the overlapping p-gene product. We also found M75 and M103 can initiate truncated surface protein (TSPs) synthesis upon over-expression of full-length surface proteins, which may possibly contribute to HBV genome replication. However, attempts to rescue replication- defective HBV mutant by co-expression of TSPs initiated from M75 or M103 were unsuccessful, which indicated surface proteins rather than the putative TSPs were involved in regulation of HBV genome replication.

Canine Distemper Virus Utilizes Different Receptors to Infect Chicken Embryo Fibroblasts and Vero cells*
2011, 26(2): 139 doi: 10.1007/s12250-011-3176-3
Received: 05 December 2010
Accepted: 15 February 2011
Inducing animal viruses to adapt to chicken embryos or chicken embryo fibroblasts (CEF) is a common method to develop attenuated live vaccines with full security. Canine distemper virus (CDV) also does this, but the mechanisms and particular receptors remain unclear. Virus overlay protein blot assays were carried out on CEF membrane proteins, which were extracted respectively with a Mem-PERTM kit, a radioimmunoprecipitation assay buffer or a modified co-immunoprecipitation method, and revealed a common 57 kDa positive band that differed from the 42-kDa positive band in Vero cells and also from those receptors reported in lymphocytes and 293 cells, indicating a receptor diversity of CDV and the possibility of the 57-kDa protein acting as a receptor that is involved in adaptive infection of CDV Kunming strain to CEF.